Clear Sky Science · en
The first mitochondrial genome for Sterictiphorinae (Hymenoptera: Argidae) and insights into argid phylogeny
Why tiny leaf‑eaters matter
Sawflies in the family Argidae may look unassuming, but their caterpillar‑like young can strip leaves from crops and forests, making them important agricultural pests. Yet, at the level of their DNA, especially in the energy‑producing mitochondria inside their cells, these insects have remained poorly studied. This paper reports the first complete mitochondrial genome for a Korean sawfly, Sterictiphora koreana, and uses it to explore how this pest‑ridden family evolved over more than 160 million years.
Peeking inside a sawfly’s power plants
The authors decoded all of the mitochondrial DNA from a single female S. koreana. Like most animals, this tiny “power plant” genome is a loop of DNA carrying 37 genes that help the cell harvest energy. In S. koreana, the loop is unusually long—about 17,900 DNA letters compared with roughly 15,600 in three related Argidae species—mainly because one non‑coding stretch rich in the letters A and T has expanded. Despite this size difference, the overall layout of genes closely matches the typical pattern seen in other sawflies, with most genes on one DNA strand and a smaller set on the opposite strand, and transfer‑RNA genes folding into classic cloverleaf shapes with a few quirks.

Letter patterns that hint at family ties
Mitochondrial DNA is heavily biased toward the letters A and T in many insects, and these sawflies are no exception. All four studied Argidae species have genomes that are roughly 80 percent A or T, but the balance is not identical. The three previously known species, all from the subfamily Arginae, are even more A+T‑rich than S. koreana, which belongs to the subfamily Sterictiphorinae. Subtle imbalances in how often A appears compared with T, and G compared with C, differ between species and between parts of the genome. One species, Arge aurora, even reverses some of these skews, standing out from its relatives. The researchers also show that certain amino acids, such as leucine, isoleucine, methionine and phenylalanine, are encoded more often than others, reflecting how this biased alphabet shapes the genetic “words” used by sawflies.
Shuffled genes and slow‑changing code
Although the overall mitochondrial blueprint is conservative, small genes called tRNAs are prone to being moved around, and their new positions can mark major branches in the sawfly family tree. In Arginae species, one tRNA gene (trnW) has hopped to a new spot near the A+T‑rich region. In S. koreana, a different set of tRNAs (trnK, trnD, trnI and trnM) has been reshuffled in a consistent pattern. These distinct rearrangements likely flag the two subfamilies as separate lineages. When the team examined how quickly the protein‑coding genes accumulate changes, they found most changes are weeded out by natural selection, a pattern called purifying selection. One gene in particular, cox1, changes especially slowly, reinforcing its usefulness as a DNA “barcode” for telling species apart.
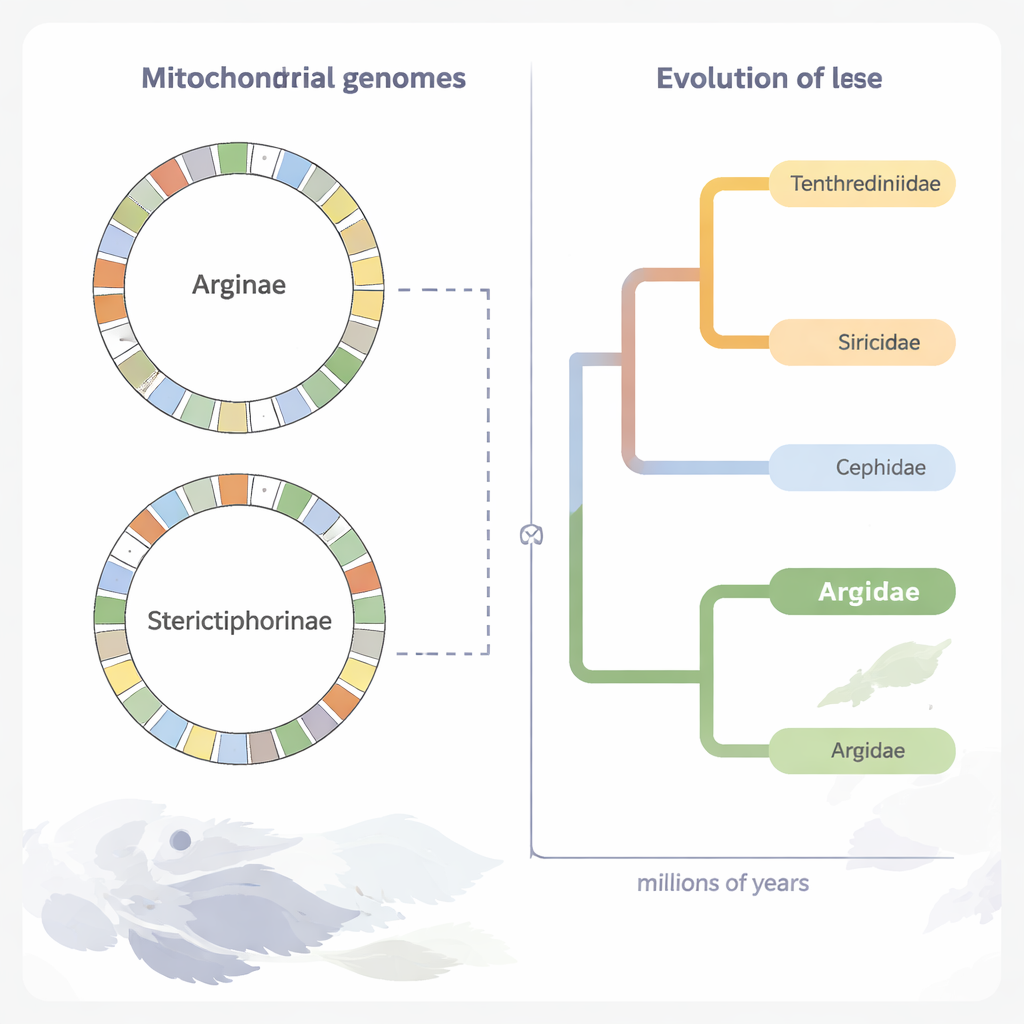
Rebuilding an ancient sawfly family tree
Using the protein‑coding portions of mitochondrial DNA from 70 sawfly species, the researchers reconstructed a detailed evolutionary tree. Their analyses confirm that Argidae forms a natural group and is closely allied with another family, Pergidae, within the larger sawfly superfamily Tenthredinoidea. By layering in fossil evidence, they estimate that the Argidae–Pergidae line split from other sawfly families around 206 million years ago, and that Argidae itself arose roughly 166 million years ago, in the Middle Jurassic. The two major argid subfamilies, Arginae and Sterictiphorinae, appear to have parted ways in the Early Cretaceous, about 126 million years ago, a time when flowering plants were also diversifying.
What this means for science and pest control
This first mitochondrial genome from Sterictiphorinae plugs a major gap in our genetic picture of Argidae. It shows that fine‑scale features—such as genome length, gene order, and letter biases—can reliably separate major groups within this pest family and help anchor them in the broader sawfly tree of life. For non‑specialists, the take‑home message is that by reading and comparing the tiny loops of DNA in insect cell power plants, scientists can trace how destructive leaf‑eating sawflies are related, estimate when their lineages emerged, and ultimately build a stronger framework for identifying species and understanding their spread in crops and forests.
Citation: Park, B., Hwang, U.W. The first mitochondrial genome for Sterictiphorinae (Hymenoptera: Argidae) and insights into argid phylogeny. Sci Rep 16, 7154 (2026). https://doi.org/10.1038/s41598-026-38021-9
Keywords: sawfly evolution, mitochondrial genome, insect phylogeny, forest and crop pests, DNA barcoding