Clear Sky Science · en
Computational identification and mechanistic characterization of natural product binders targeting the PDE6D prenyl binding tunnel
Turning Cancer’s Favorite Switch Against Itself
Many of the deadliest cancers—from pancreatic to colorectal and lung tumors—depend on a single molecular “on switch” called RAS. When this switch gets jammed in the ON position, cells grow uncontrollably. Drug developers have struggled for decades to turn RAS off directly. This study explores a different tactic: hunting for natural compounds that can quietly block the helper protein that RAS needs to reach its place of action, potentially dimming its signal without having to hit RAS head-on. 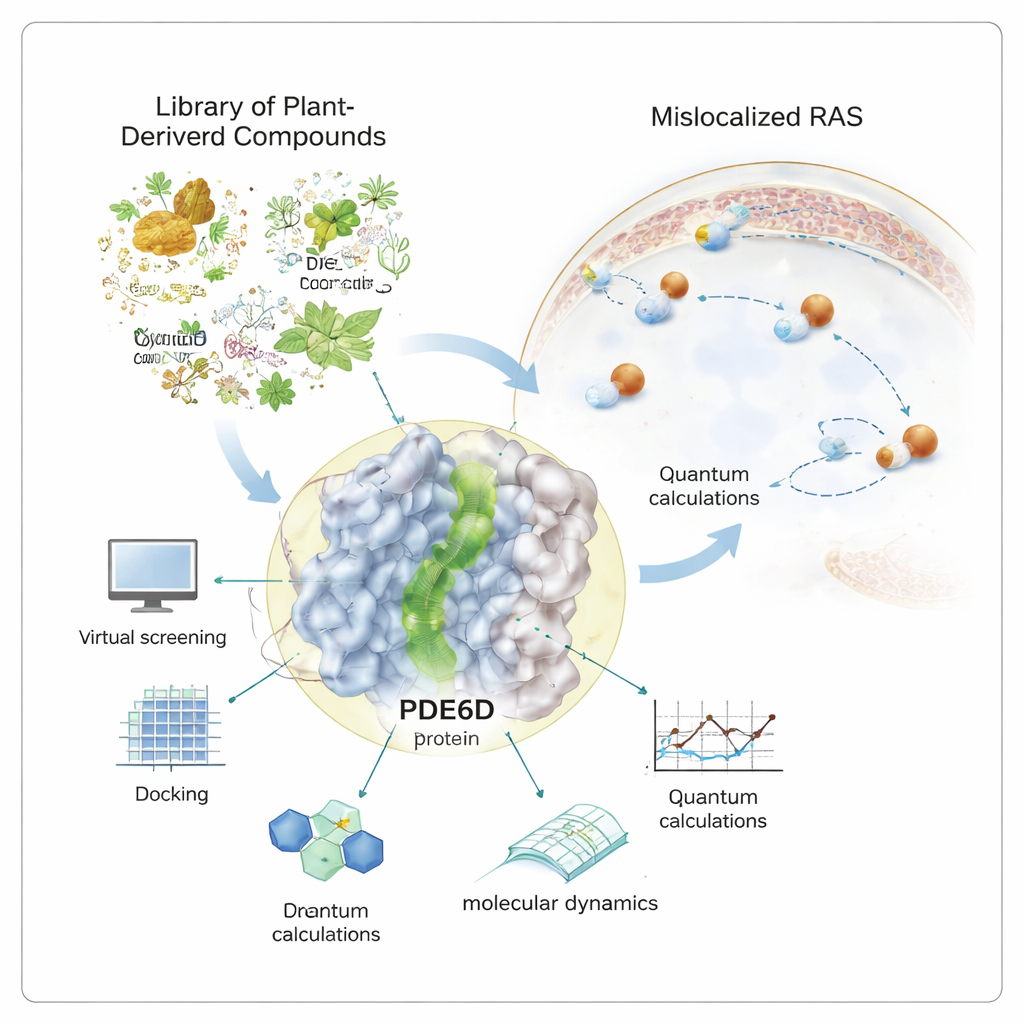
A Hidden Helper in Cancer Cells
RAS proteins must sit on the inner surface of the cell membrane to send growth signals. To get there, they carry a greasy tail that allows them to anchor in the membrane. A chaperone protein called PDE6D acts like a taxi, shielding this tail as RAS travels through the watery interior of the cell. PDE6D has a narrow, oil-lined tunnel that grips the RAS tail. If that tunnel is plugged, RAS is more likely to end up in the wrong place, weakening its ability to drive cancer. Because many existing RAS drugs only work for a small subset of mutations, researchers are keen to explore PDE6D as an indirect but more broadly relevant target.
Using Computers to Mine Nature’s Chemical Library
Rather than relying on synthetic chemicals alone, the author turned to a curated library of over a thousand purchasable natural products—molecules originally derived from plants, microbes, and other living sources. Powerful computer tools were used to screen this library against the 3D structure of PDE6D. In this virtual test, each molecule was “docked” into the tunnel to see how snugly and favorably it might bind. The best-scoring candidates were then examined more closely with quantum-level calculations, which probe how electrons are arranged in each molecule and how easily they can participate in stabilizing interactions inside the protein pocket.
Stress-Testing Promising Candidates in Motion
Proteins and small molecules do not sit still inside cells, so the study went beyond static snapshots. Using long molecular dynamics simulations that follow atomic motions over half a microsecond, the researcher watched how each promising natural compound behaved inside the PDE6D tunnel over time. Two candidates, labeled MolPort-039-052-621 and MolPort-002-507-186, stayed lodged in the tunnel with stable contact networks, while a third molecule drifted toward the tunnel exit and became more weakly attached. Additional analyses mapped which parts of the PDE6D structure flexed or quieted down upon binding, and how the complex explored its “energy landscape”—essentially, which shapes were most comfortable and long-lived. 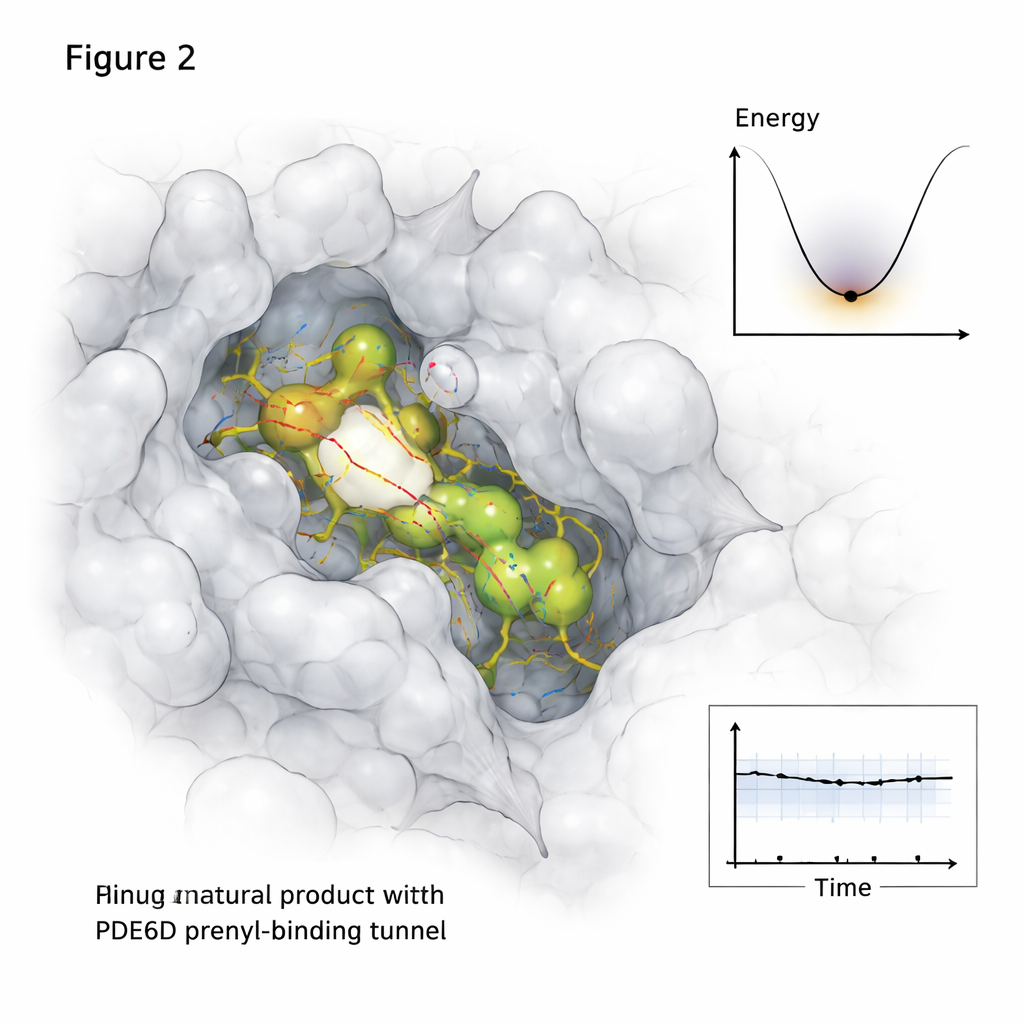
Balancing Binding Strength and Drug-Like Behavior
The study also used online prediction tools to ask whether these natural compounds look like realistic starting points for medicines. All three top hits had chemical features consistent with basic “drug-likeness,” but each carried red flags. Some were predicted to have poor absorption or solubility; others showed signs of possible toxicity, such as effects on heart rhythm or DNA. These issues mean the molecules, as they are, are unlikely to be ready-made drugs. Instead, they are best viewed as structural templates that chemists might modify, keeping the tunnel-fitting features while trimming problematic parts.
What This Means for Future Cancer Treatments
In simple terms, this work does not deliver a new cancer drug, but it does chart a promising route. By combining several layers of computation—from fast docking to detailed simulations and quantum calculations—the study narrows a vast set of natural molecules down to a small group that appear well suited to plugging the PDE6D tunnel. If future lab experiments confirm that these compounds truly misdirect RAS inside cells and slow the growth of RAS-driven tumors, they could inspire a new class of treatments that target cancer’s “logistics” rather than its main driver directly.
Citation: Alshahrani, M.M. Computational identification and mechanistic characterization of natural product binders targeting the PDE6D prenyl binding tunnel. Sci Rep 16, 6571 (2026). https://doi.org/10.1038/s41598-026-37939-4
Keywords: RAS signaling, PDE6D inhibitors, natural products, computational drug discovery, cancer therapy