Clear Sky Science · en
Whole-cell modeling predicts alternative proteome allocation strategies in the archaeon Methanococcus maripaludis
Why tiny methane-makers matter
Methanococcus maripaludis is a microscopic archaeon that helps turn simple chemicals like carbon dioxide and formate into methane, a major component of natural gas. Understanding how this microbe budgets its internal resources—especially its proteins—can reveal general rules of life under energy limitation and guide efforts to harness it as a biological methane factory or a platform for green chemistry.
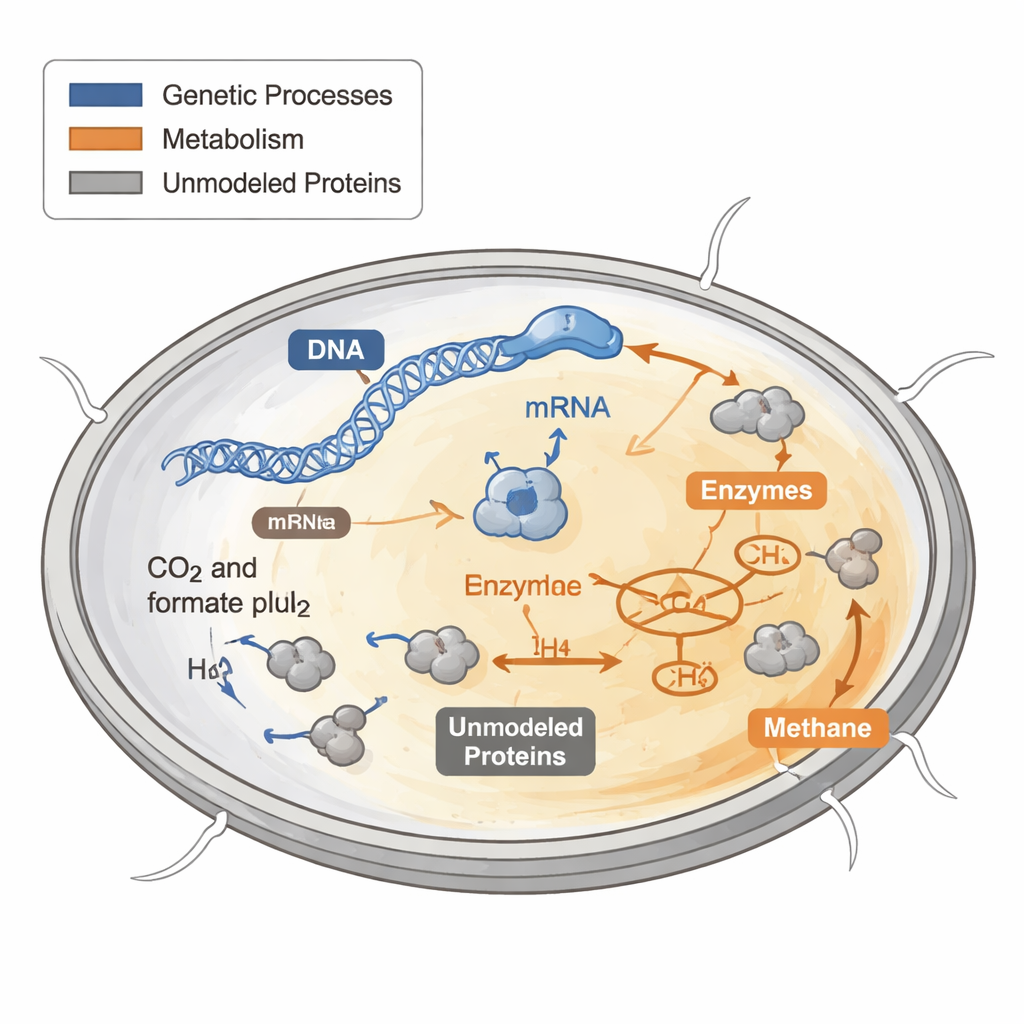
How cells spend their protein budget
Every cell has to decide how to spend its limited protein “budget.” Some proteins build new cell parts, some harvest energy, and others perform housekeeping tasks. Earlier computer models of metabolism treated these proteins as if they were free and unlimited, which works poorly when trying to predict growth under harsh or nutrient-poor conditions. In bacteria and yeast, experiments have shown that fast-growing cells devote a larger slice of their protein budget to ribosomes, the molecular machines that build new proteins, while slow-growing cells shift resources elsewhere.
An archaeon that breaks the rules
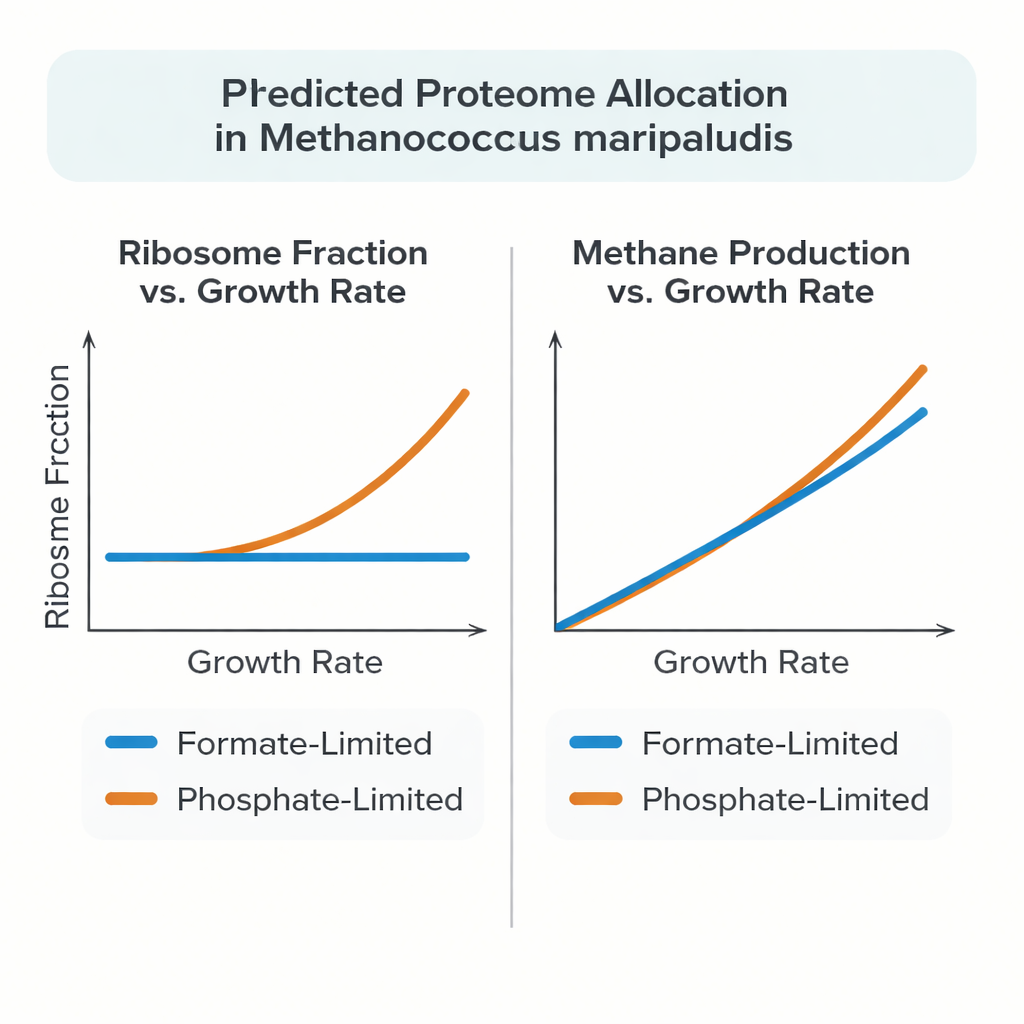
Recent experiments revealed that M. maripaludis does not always follow this pattern. When it grows using formate as both food and energy source, the fraction of its protein budget invested in ribosomes stays almost constant as growth slows down. Under phosphate limitation, however, the archaeon looks more conventional: the ribosome fraction rises with growth rate, similar to Escherichia coli and yeast. To explain this unusual behavior, the authors built a detailed whole-cell style model that links metabolism to the costly process of making and assembling proteins and RNA, rather than treating proteins as abstract reaction rates.
Building a whole-cell style model
The researchers started from an existing metabolic map of M. maripaludis and layered on thousands of additional reactions that describe gene expression: copying DNA into RNA, charging transfer RNAs, assembling ribosomes and enzyme complexes, and diluting these components during cell division. They gathered enzyme speed limits from biochemical databases and literature, and estimated how much of the protein mass is made of “unmodeled” proteins that are not directly tied to the reactions in the map. They then imposed global constraints that limit the total mass of protein and RNA, link reaction rates to the amount of enzyme available, and tie protein production to ribosome capacity. With these constraints, the model has to choose how to allocate a fixed protein budget to support growth.
What the simulations revealed
When tested against high-quality chemostat experiments, the model accurately reproduced measured protein and RNA masses, methane production rates, and growth yields over a range of slow growth rates. Under formate limitation, the simulated cell naturally settled on a nearly constant ribosomal protein fraction as growth changed, mirroring the experimental data. Under phosphate limitation, the model matched the observed increase in ribosome fraction only when ribosomes were allowed to work faster at low growth rates, consistent with measurements showing more ribosomes actively engaged in protein synthesis. The model also suggested that a substantial share of protein in M. maripaludis is not strictly required for minimal growth, and that excess carbon and energy can be stored or diverted into other large molecules such as glycogen or free amino acids, depending on which nutrient is limiting.
Probing mutants and growth limits
Because the model explicitly accounts for the protein cost of each pathway, it can predict not just whether genetic mutants grow, but how well they grow relative to the wild type. The authors simulated strains lacking different hydrogen-processing enzymes and compared predicted fitness with laboratory measurements across several growth conditions, including the presence or absence of hydrogen and carbon monoxide. In many cases, the model captured whether mutants could grow and produced reasonable estimates of their relative growth rates, improving on earlier models that could only give yes-or-no answers. The same framework was used to estimate maximum possible growth rates under batch conditions for different substrate combinations.
What this means for science and technology
In everyday terms, this study shows that M. maripaludis uses an alternative strategy for budgeting its protein workforce, keeping its protein factories running at a steady share of the budget under some conditions rather than throttling them back as growth slows. By embedding that behavior into a detailed, data-driven model, the work provides a test bed for exploring how energy-limited microbes balance growth, maintenance, and storage. For applied research, the model offers a roadmap for engineering this archaeon to make more methane or other products by identifying which enzymes and pathways are most costly in terms of protein investment. More broadly, it extends powerful whole-cell style modeling beyond bacteria and yeast into the archaeal domain, helping bridge the gap between molecular details and the large-scale behavior of microbial communities involved in climate, energy, and industrial biotechnology.
Citation: Kasem, G.S., Soliman, T.H.A., Mousa, M.A.A. et al. Whole-cell modeling predicts alternative proteome allocation strategies in the archaeon Methanococcus maripaludis. Sci Rep 16, 7386 (2026). https://doi.org/10.1038/s41598-026-37887-z
Keywords: Methanococcus maripaludis, proteome allocation, whole-cell modeling, methanogenesis, archaea metabolism