Clear Sky Science · en
Modeling and application of alzheimer’s disease complex trait prediction based on multi-task learning
Why this research matters for families and patients
Alzheimer’s disease is one of the most feared diagnoses of our time, yet doctors still struggle to predict who will decline quickly, who will stay stable for years, and which early signs really matter. This study asks a simple but powerful question: if we look at several Alzheimer’s-related test results and brain scans together, and combine them with a person’s genetic information, can modern artificial intelligence learn patterns that help us forecast the course of the disease more accurately?
Many faces of the same disease
Alzheimer’s is not just memory loss. Patients differ in how they perform on thinking tests, how well they manage daily tasks, and what their brain scans look like. These different measurements—such as common memory and thinking scales, questionnaires about daily functioning, and PET scans of brain metabolism or amyloid buildup—are known to be partly influenced by genes. Importantly, they also share some of the same genetic roots. Traditional prediction methods usually focus on one measure at a time, throwing away the useful fact that these traits are related. The authors argue that, like a doctor who sees the full picture instead of a single test, models should learn from several traits together.
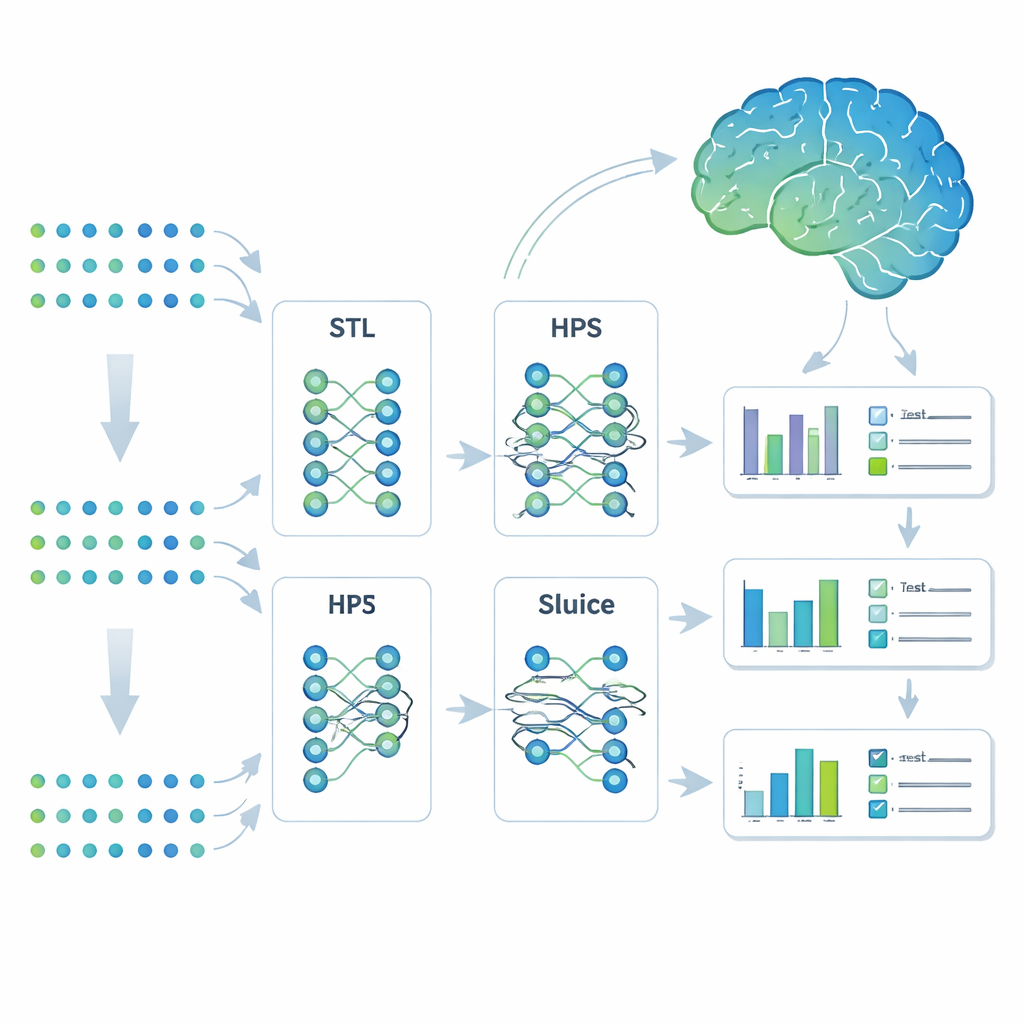
Teaching one model to learn many related tasks
The researchers turned to a machine-learning strategy called multi-task learning. Instead of building separate models for each outcome, they trained a single system to predict seven Alzheimer’s-related traits at once. They compared four approaches: completely separate models (single-task learning), a simple shared model that splits only at the end (hard parameter sharing), a more flexible branching design that can split tasks into subgroups, and a highly adaptable design called the Sluice Network that can fine-tune how much information is shared at each layer of the network. All four models saw the same genetic inputs; the difference lay in how they shared what they learned across traits.
Testing ideas in simulated genomes
Before trusting any model on real patients, the team built detailed simulations using real genetic patterns taken from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) but with outcomes they could fully control. They created scenarios where all traits shared the same genetic causes, where traits formed overlapping groups, and where each trait had distinct causes. They also varied how strong the genetic signals were and how much noise they added, mimicking the messy reality of human data. Across nearly all conditions, the Sluice Network delivered the most accurate predictions and remained stable even when traits were only weakly related. Simpler shared models worked well when traits had many genetic factors in common but faltered when that sharing was low, while completely separate models were steady but less accurate overall.
Real-world data and the power of grouping genes
The authors then applied these models to actual ADNI data from 463 individuals, using nearly 3,800 genetic markers drawn from 56 genes previously linked to Alzheimer’s. Here they added a biologically inspired twist: instead of feeding in thousands of individual genetic markers, they first grouped markers by gene and let the network learn a compact “summary” signal for each gene before predicting the seven outcomes. This gene-level aggregation boosted performance for most models and especially for the Sluice Network, which roughly doubled its average correlation with the real outcomes. Gains were clearest for PET imaging measures and certain cognitive and functional scores, suggesting that subtle genetic effects become more detectable when combined at the gene level rather than treated as isolated markers.
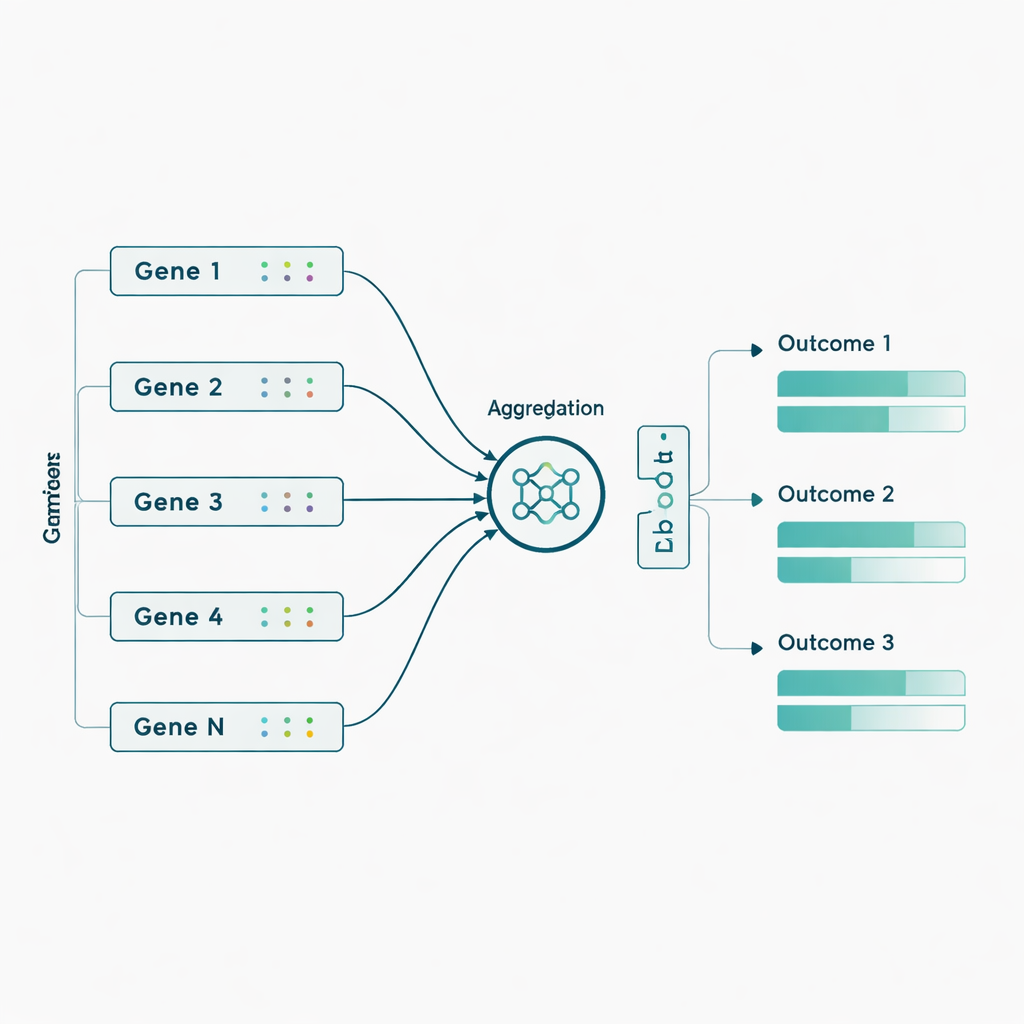
What this means for future prediction and care
To a non-specialist, the message is that smarter, more flexible AI models can squeeze more insight out of the same genetic and clinical data by learning from several related outcomes at once and by respecting how biology is organized into genes. While the current gains are modest and far from a clinical test, the approach points toward more reliable tools for estimating a person’s risk profile, tracking likely progression, and perhaps tailoring monitoring or interventions. In complex diseases like Alzheimer’s, where many small genetic effects interact, methods that share information across traits and aggregate weak signals may offer a clearer, more informative picture than traditional one-trait-at-a-time scores.
Citation: Zhou, W., Xue, Z., Liang, J. et al. Modeling and application of alzheimer’s disease complex trait prediction based on multi-task learning. Sci Rep 16, 7749 (2026). https://doi.org/10.1038/s41598-026-37820-4
Keywords: Alzheimer's disease genetics, multi-task learning, deep learning prediction, neuroimaging biomarkers, gene-level aggregation