Clear Sky Science · en
Cancer classification with radiomics in controlled preclinical models
Reading cancer from scans and blood
Modern cancer care increasingly relies on computers to sift through medical images and lab tests for patterns that humans can’t easily see. This study asks a simple but important question: if we want to tell what kind of cancer a patient has, is it better to read hidden information from their scans, or from their blood? Using carefully controlled experiments in mice, the researchers directly compared these two approaches to see which one offers more reliable answers.
What it means to take a “virtual biopsy”
Radiomics is a fast-growing technique that treats each medical scan as a rich data source rather than just a picture. Special software combs through three-dimensional CT images of a tumor and converts them into hundreds of numerical features that describe its shape, brightness, and fine-grained texture. In principle, these patterns might reflect the tumor’s biology in a way similar to a biopsy, but without needles or surgery—a so-called “virtual biopsy.” Advocates hope radiomics will help classify cancers, gauge how aggressive they are, and guide treatment choices. But there are concerns: results can be hard to reproduce, easy to confound by technical quirks, and difficult for doctors to interpret.
A fair head‑to‑head test in mice
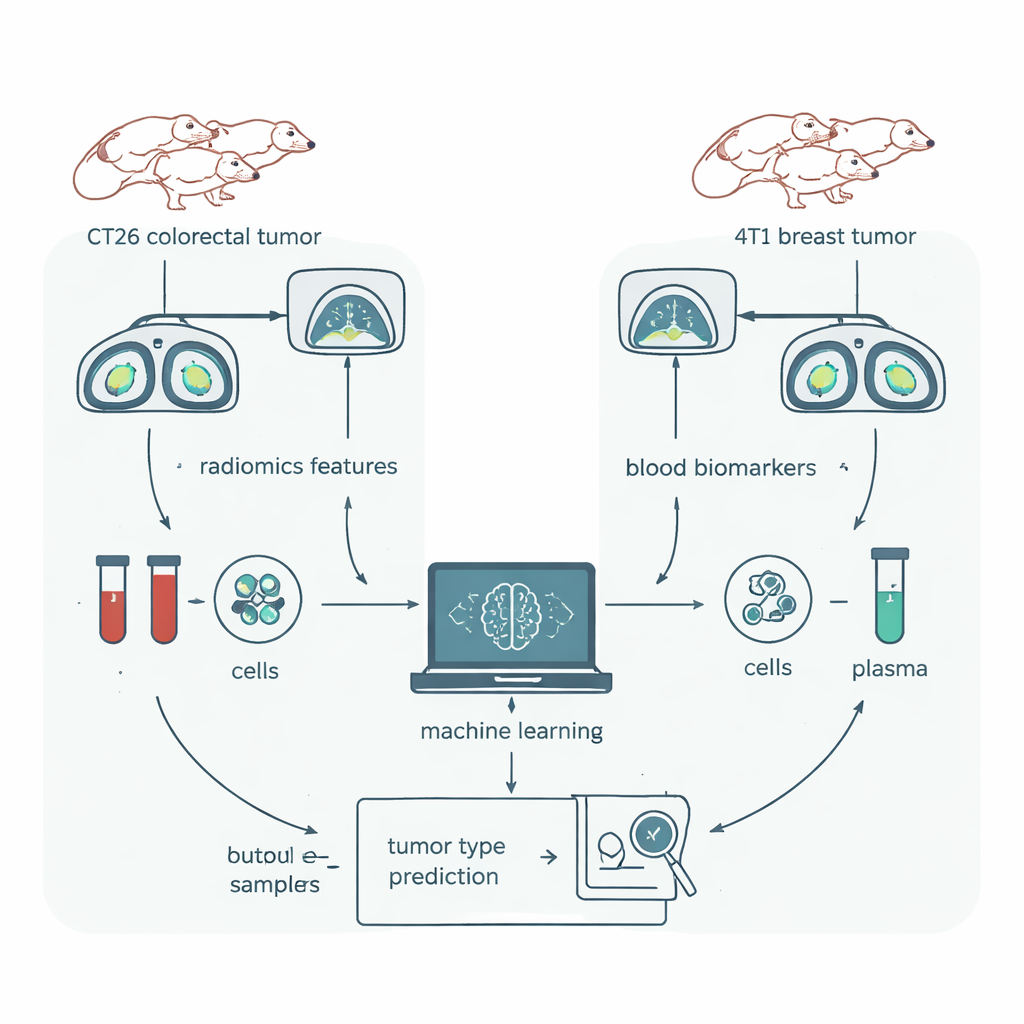
To give radiomics a rigorous test, the team turned to a mouse model where nearly everything could be controlled. Groups of genetically identical mice were implanted with one of two tumor types: CT26, a model of colorectal cancer, and 4T1, a model of breast cancer. All animals were the same strain, sex, and similar age, kept in the same environment, and scanned on the same CT machine. Tumors were carefully outlined in 3D software, and a popular radiomics package extracted 1,409 numerical features from each scan. In parallel, the researchers drew blood from the same animals and measured immune cell types and dozens of proteins—biomarkers that their earlier work had already shown could distinguish these cancer models almost perfectly.
Compressing thousands of image details into a useful signal
Most of the raw image features turned out to be unhelpful: some barely varied from mouse to mouse, and many were almost duplicates of each other. After several rounds of statistical filtering, only 18 non‑redundant radiomic features remained, largely describing subtle texture patterns rather than simple size or shape. The team then used a standard machine‑learning method, Random Forest, to see how well these refined image features could tell the two tumor types apart. They also applied visualization tools to see whether the data naturally formed separate clusters for each cancer type without knowing the labels in advance.
Blood signals beat image signals
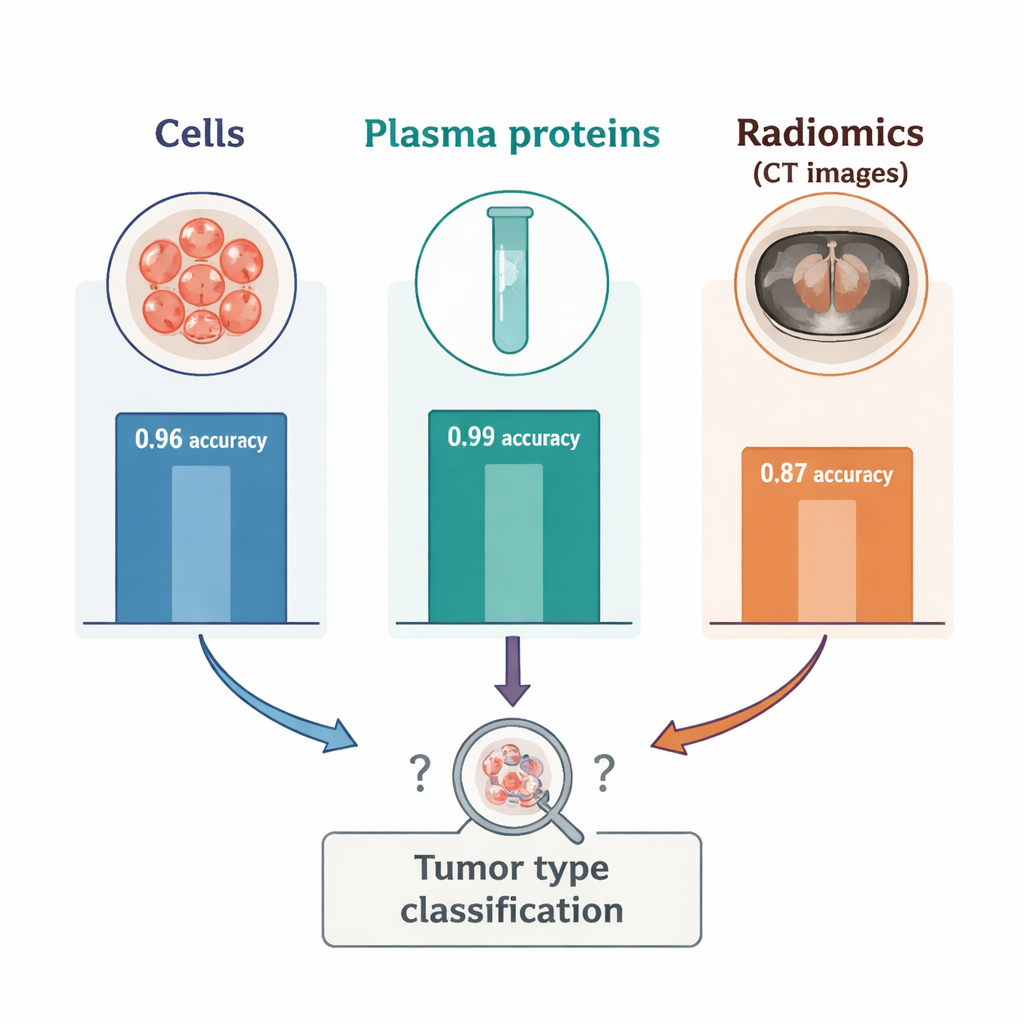
The contrast between blood and imaging was striking. When the researchers reduced the blood cell and plasma protein data to two dimensions, the two tumor types formed cleanly separated clusters, confirming that blood captured strong, tumor‑specific signals. In the radiomics data, however, three mixed clusters appeared, each containing a blend of the two cancers and suggesting that other, unknown factors were shaping the images. In supervised tests, radiomics alone classified tumor type with an accuracy of about 87 percent—good, but noticeably worse than the 96 percent accuracy from immune cell counts and the 99 percent accuracy from plasma proteins. Adding radiomics to the blood markers did not improve performance; in some combinations, it slightly reduced accuracy. A further experiment showed that using only a small spherical region inside the tumor, instead of outlining the whole mass, made radiomics perform even worse, underscoring how sensitive these features are to how the tumor is drawn on the scan.
What this means for future cancer tests
For a lay reader, the takeaway is clear: while advanced image analysis can provide some useful clues, in this study it was outperformed by relatively simple blood tests when it came to telling two cancer types apart. Even in a tightly controlled laboratory setting with identical mice and standardized imaging, small technical differences and the complexity of image processing appeared to blur the radiomic signal. The authors conclude that radiomics is not yet ready to serve as a standalone, highly dependable cancer classifier. Instead, they argue that stronger standardization of imaging, better contouring tools, and clearer links between image patterns and underlying biology will be needed before virtual biopsies can reliably guide clinical decisions alongside, or in place of, well-established blood biomarkers.
Citation: Drover, K., Davis, D.A.S., Gosling, K. et al. Cancer classification with radiomics in controlled preclinical models. Sci Rep 16, 6647 (2026). https://doi.org/10.1038/s41598-026-37757-8
Keywords: radiomics, cancer biomarkers, medical imaging, machine learning in oncology, blood tests for cancer