Clear Sky Science · en
Multidimensional strategy enables scalable metabolome diversity in microbial fermentations
Why tiny factory tweaks matter for new medicines
Many of today’s medicines come from natural chemicals made by bacteria and fungi. But turning a promising lab hit into a real drug lead often fails at a surprisingly down‑to‑earth step: growing the same microbe in a different vessel or at a larger scale. This study asks a simple but crucial question—how can we keep a microbe’s chemical output stable and diverse when we move it from small test plates to larger flasks and bioreactors? The answer could speed the search for the next generation of antibiotics and other therapies.
Three ways to grow the same microbe
The researchers focused on a soil bacterium, Streptomyces griseochromogenes, known for making many “secondary metabolites” – small molecules with potential as drugs. They compared three common cultivation systems used in early discovery: baffled shake flasks, 48‑well “flower” microtiter plates, and a stirred tank bioreactor. In each system they measured growth, cell shape and structure, and, crucially, the pattern of chemical signals detected by mass spectrometry, used here as a fingerprint of the bacterium’s secreted metabolome. They first applied a classical engineering rule: keep oxygen availability the same across all systems and expect similar behavior. Instead, they found that growth curves, cell forms and chemical fingerprints differed sharply from one vessel to the next.
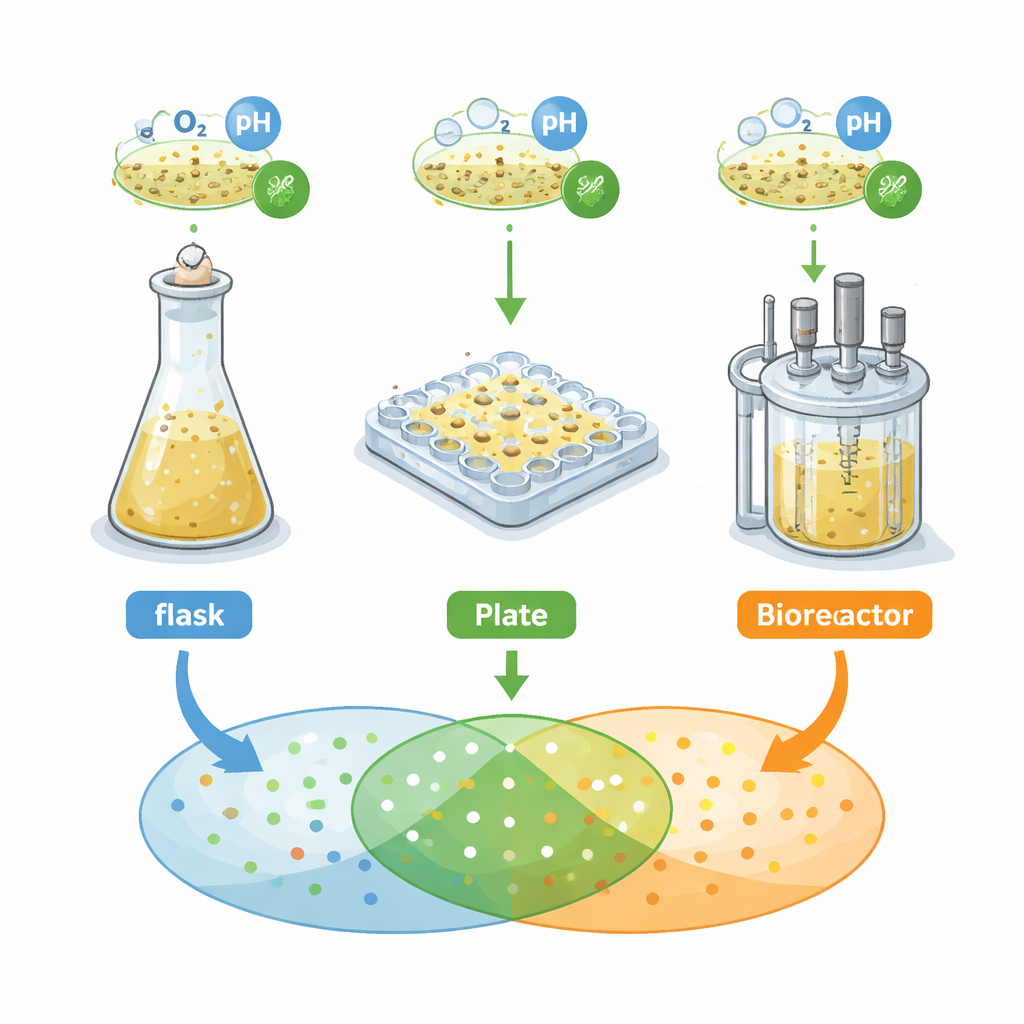
When one rule is not enough
Matching oxygen alone produced very different “metabolic footprints” – the sets of mass features corresponding to individual molecules. Only about 18 percent of these features were shared across the three systems. The microtiter plates generated far more distinct signals than flasks or the bioreactor. The team then tried another single‑factor approach, changing either the amount of ethanol in the medium (a small molecule that can nudge bacteria to make more metabolites) or the agitation speed, which alters how much oxygen gets into the culture. These one‑dimensional tweaks modestly improved the overlap in chemical footprints, but only by another 18 percent or so. In other words, simple recipes like “add more oxygen” or “add more ethanol” could not guarantee that a metabolite seen in a tiny well would reliably appear in a larger fermenter.
How cell shape steers chemical diversity
To untangle what really matters, the authors pooled data from 80 different cultivations and used a statistical method that looks for hidden factors shaping the results. This analysis highlighted two main themes: the cultivation system itself and the cells’ morphology – whether the bacteria grew as compact pellets, loose mycelial mats or finely dispersed filaments. Conditions that produced similar morphologies tended to produce more similar metabolic footprints and made it easier to “scale up” molecules from small systems into the stirred tank. By deliberately selecting sets of conditions that gave comparable cell shapes in all three systems, the researchers increased the overlap in chemical features by about 50 percent compared with classical oxygen‑matching alone.
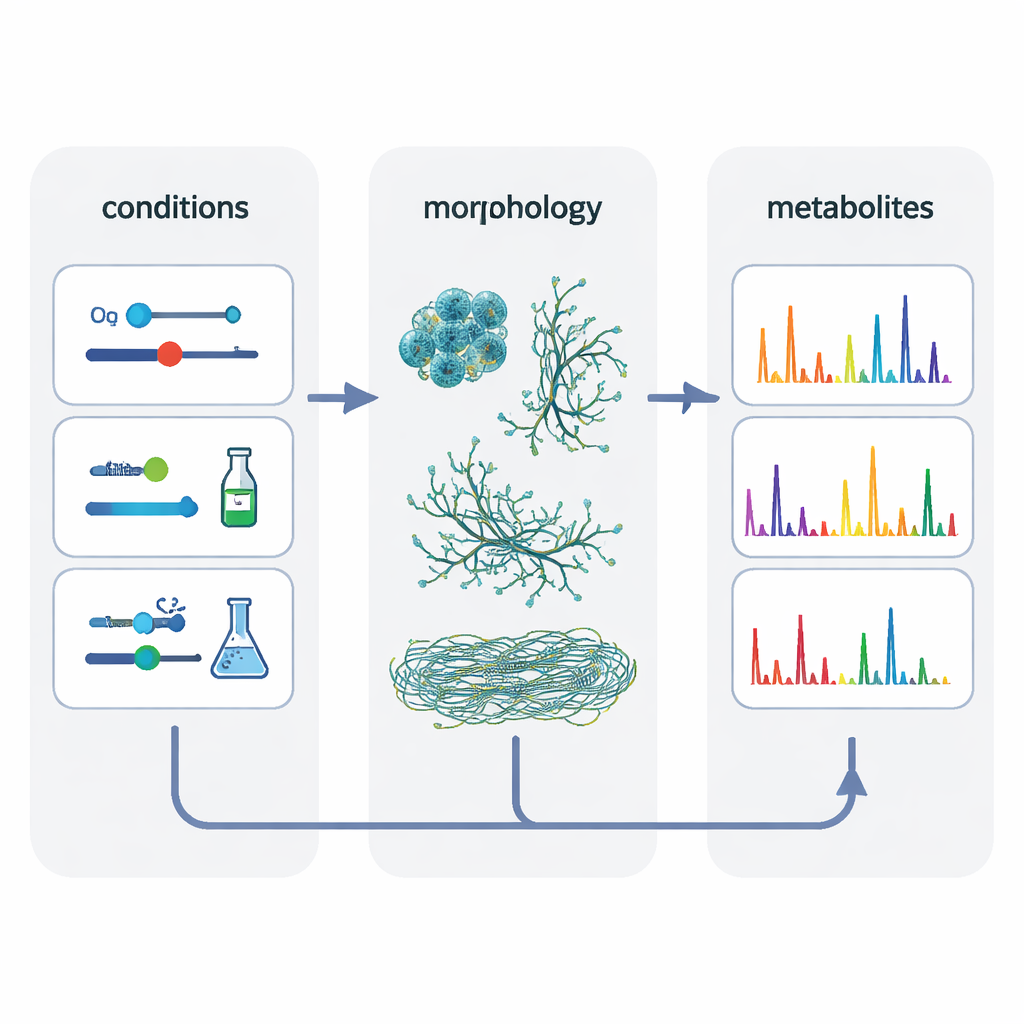
Hidden chemical families and system‑specific molecules
Beyond simple counts of signals, the team built molecular networks that grouped related mass features into families of structurally similar metabolites. Large families often contained at least one member found in every cultivation system, suggesting some core chemistry that is robust to scale changes. But many smaller families – and some individual molecules – appeared only in one type of vessel. The polystyrene microtiter plates in particular produced the richest variety of compounds, including many variants of iron‑binding siderophores called desferrioxamines and several non‑ribosomal peptides. The authors suggest that subtle stresses in these plates, such as higher levels of reactive oxygen species, may push the bacteria to diversify their chemistry, whereas the well‑mixed bioreactor favors a smaller set of “core” structures.
What this means for finding new natural products
For drug discovery teams, the message is clear: you cannot rely on a single engineering rule or one “optimal” condition if you want both rich chemical diversity and reliable scale‑up. Instead, you need a multidimensional strategy that takes vessel type, oxygen transfer, additives like ethanol and, importantly, real‑time monitoring of cell morphology into account. Using microtiter plates that mimic key features of bioreactors, and tuning conditions to produce similar growth forms, can make it far more likely that promising molecules discovered at microliter scale will reappear when grown in liters. This approach helps turn fragile early hits into robust, reproducible candidates, increasing the chances that novel natural products will survive the journey from bench to medicine cabinet.
Citation: Lindig, A., Fataeri, M., Hubmann, G. et al. Multidimensional strategy enables scalable metabolome diversity in microbial fermentations. Sci Rep 16, 4084 (2026). https://doi.org/10.1038/s41598-026-37748-9
Keywords: natural product discovery, Streptomyces fermentation, metabolomics, bioreactor scale-up, secondary metabolites