Clear Sky Science · en
Mechanism and evolutionary divergence of a novel oxidized polyvinyl alcohol hydrolase in Stenotrophomonas rhizophila QL-P4
Why a common “green” plastic still lingers in nature
Polyvinyl alcohol, or PVA, shows up in everything from laundry detergent pods to textile coatings and paper. It is often marketed as environmentally friendly because microbes can, in principle, break it down. Yet in real rivers, soils, and oceans this plastic-like material decomposes painfully slowly and can persist for decades as micro- and nanoplastic fragments. This study takes a close look at one of nature’s tools for tackling PVA: a newly discovered enzyme from a soil bacterium that can slice this stubborn polymer into smaller, safer pieces.
A tiny soil helper with a big job
Researchers previously isolated a bacterium called Stenotrophomonas rhizophila QL-P4 from mountain soil in China and found that it can grow using PVA as food. In the new work, they focused on a single suspect gene, named BAY15_0160, that looked like it might code for an enzyme able to cut oxidized PVA. By carefully deleting this gene, adding it back, and forcing the bacterium to overproduce it, they showed that BAY15_0160 is essential for efficient PVA breakdown. When the gene was removed, the microbe lost about 40% of its ability to consume PVA; restoring the gene brought performance back, confirming that its product is a key player in the degradation pathway.
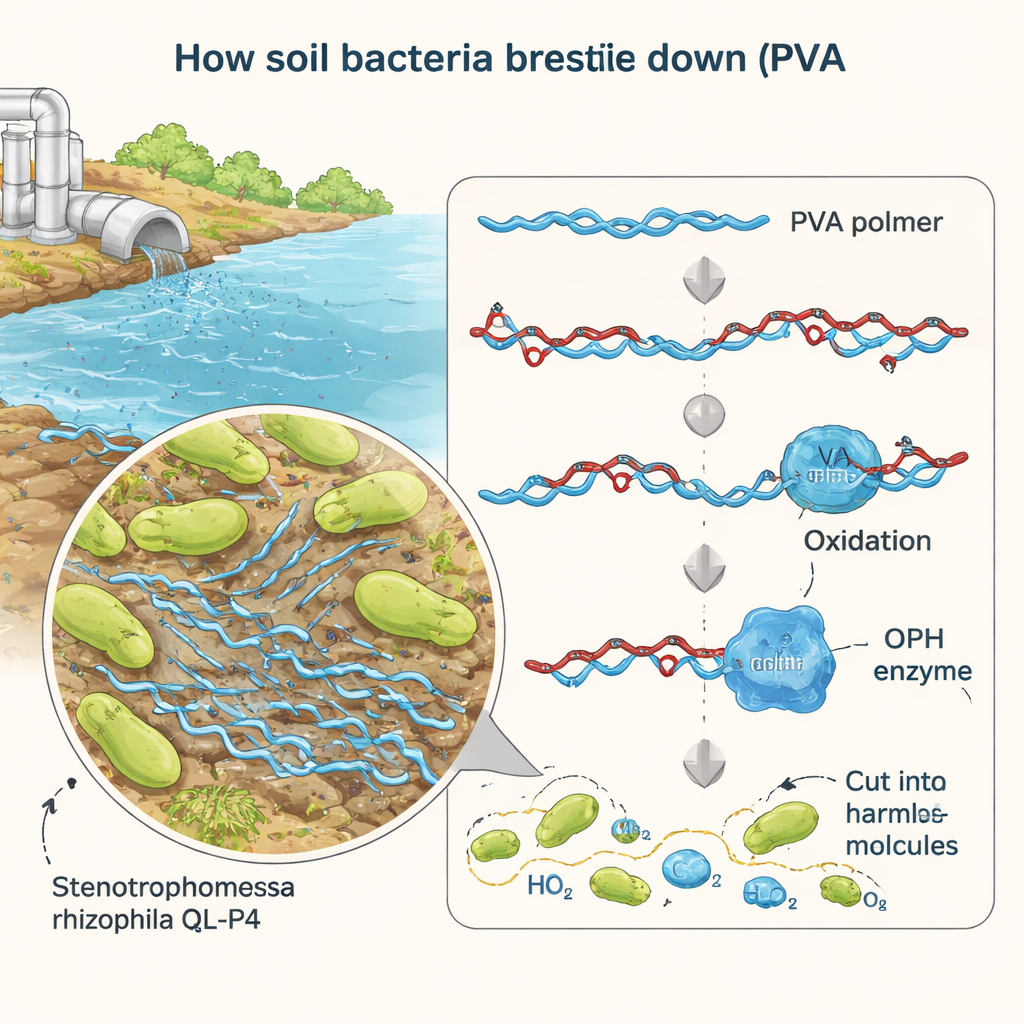
Measuring how the enzyme does its work
To study the enzyme in detail, the team produced it in large amounts using laboratory strains of Escherichia coli. They purified the protein, which turned out to be about 35 kilodaltons in size, and tested its activity under different conditions. Using a simple test molecule that releases a yellow dye when cut, they tracked how fast the enzyme worked at various temperatures and acidity levels. The enzyme was most active at around room temperature (30 °C) and at a neutral pH similar to that of many natural waters. Under these mild conditions, it showed high catalytic efficiency, meaning each enzyme molecule can process many substrate molecules every second—encouraging news for potential real-world cleanup applications.
Zooming in on the molecular scissors
Enzymes are nature’s miniature machines, and their shape determines what they can do. The researchers used state-of-the-art prediction programs, RoseTTAFold and AlphaFold, to build three-dimensional models of the PVA-cutting enzyme, now recognized as a novel oxidized PVA hydrolase, or OPH. Both tools agreed that the enzyme has a classic “alpha/beta hydrolase” fold found in many biological scissors. At its heart lies a short motif—often written as Gly–X–Ser–X–Gly—that helps position a trio of amino acids (serine, aspartate, and histidine) to attack chemical bonds. Computer simulations showed oxidized PVA fragments nestling into this groove, with the key serine likely making the first cut in the chain.
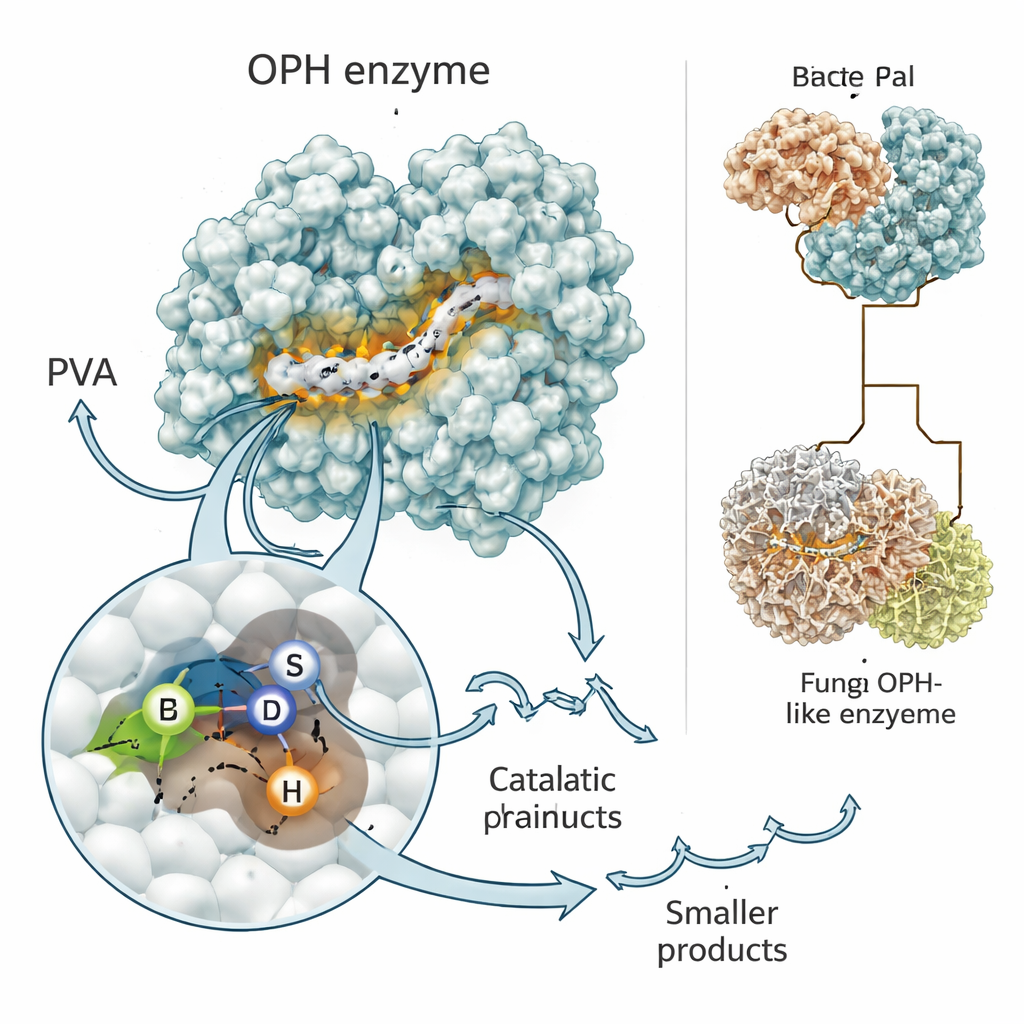
Signals, switches, and evolutionary twists
The team also explored which parts of the enzyme are absolutely required for it to function in the bacterium. One region near the front of the protein acts like a postal code, directing the enzyme outside the cell where the PVA polymer actually is. Another region contains the active motif that grips and cleaves the plastic’s carbon–carbon bonds. When the scientists engineered versions of the gene lacking either the signal segment or the active-site motif, the bacterium still made RNA from the gene but could no longer efficiently digest PVA. Looking across many bacteria and fungi, they found relatives of this enzyme that share the same core “cutting” region but differ in add-on parts, including a striking fungal version that fuses the cutting domain to a built-in transporter, hinting at an even more efficient import‑and‑digest strategy in extreme environments like Antarctic rocks.
What this means for cleaning up plastic pollution
For non-specialists, the main message is that scientists are learning how some microbes naturally chew through a supposedly biodegradable plastic that otherwise sticks around far too long. This study pinpoints a single enzyme, OPH from S. rhizophila QL-P4, as a powerful molecular scissor that operates under gentle conditions and targets a crucial step in the breakdown of PVA. By understanding its structure, working conditions, and evolutionary cousins, researchers can begin to design better microbial strains or enzyme mixtures tailored for wastewater treatment plants, industrial effluents, or contaminated soils. In the long run, such insights bring us closer to practical, biology-based solutions for managing plastic waste rather than leaving it to accumulate in the environment.
Citation: Zhou, Y., Bold, N., Feng, J. et al. Mechanism and evolutionary divergence of a novel oxidized polyvinyl alcohol hydrolase in Stenotrophomonas rhizophila QL-P4. Sci Rep 16, 6411 (2026). https://doi.org/10.1038/s41598-026-37715-4
Keywords: polyvinyl alcohol, biodegradation, plastic pollution, microbial enzymes, bioremediation