Clear Sky Science · en
Lipid metabolism classification of gliomas
Why fats in brain tumors matter
Brain tumors called gliomas are among the most dangerous cancers, yet patients with what looks like the same diagnosis can have very different outcomes. This study asks a deceptively simple question with big implications: how do the ways tumors use fats – the body’s lipids – shape how aggressive they are, how they respond to treatment, and whether we can see these differences on standard brain scans? By tracking lipid use in hundreds of tumors, the authors uncover hidden subtypes of glioma that could change how doctors predict prognosis and design therapies.
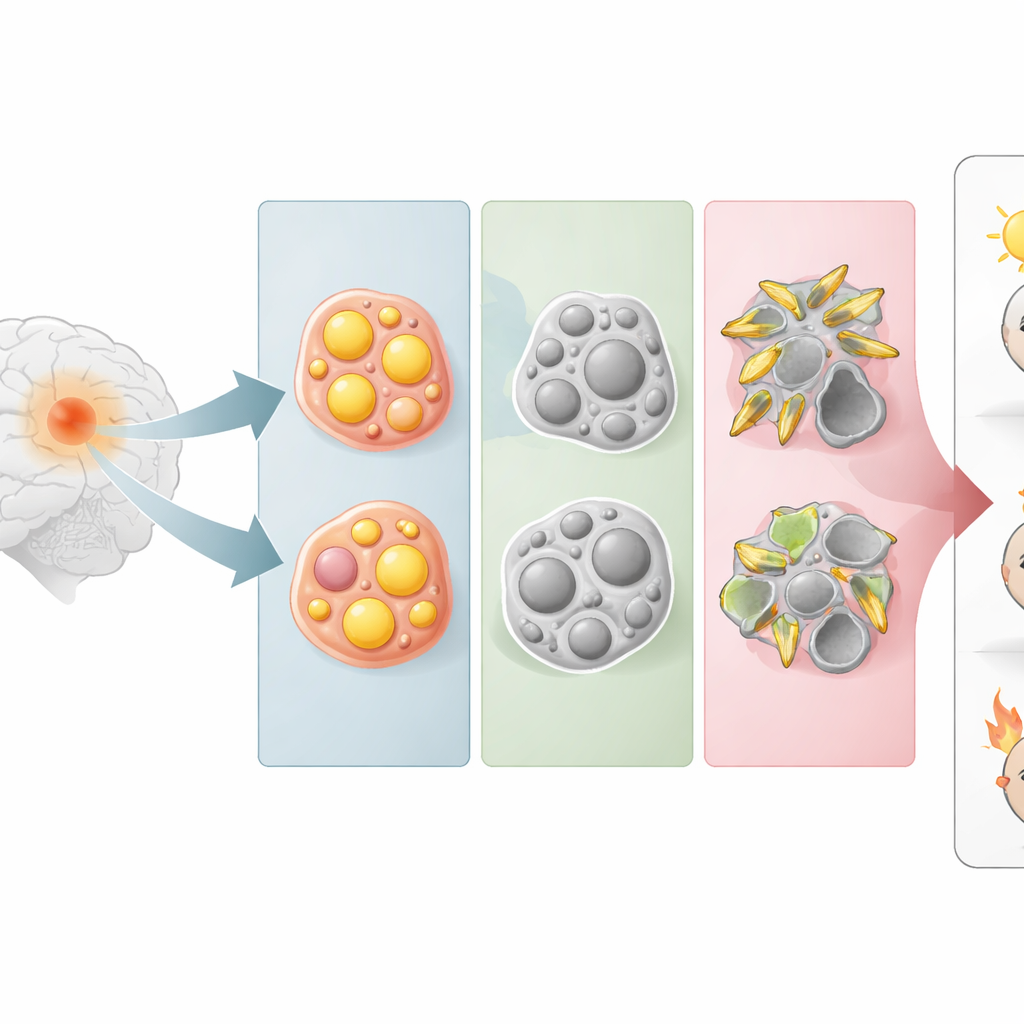
Three hidden faces of the same brain tumor
Instead of starting from how tumor cells look under a microscope, the researchers grouped gliomas by how strongly they activate five major lipid pathways, including those that handle steroid fats, triglycerides, and sphingolipids (key building blocks of cell membranes). Using gene activity profiles from large public tumor databases, they found that gliomas naturally fell into three groups. One group leaned heavily on steroid metabolism (ST-type), another on triglyceride metabolism (TC-type), and a third on sphingolipid metabolism (SP-type). These metabolic groupings cut across classic tumor categories, revealing that cells with similar lipid habits can be found in otherwise different-looking gliomas.
From metabolism to patient outlook
The team then asked how these three fat-usage styles relate to real-world outcomes. Patients whose tumors were in the ST-type group generally had the longest survival, and their tumors were more often lower-grade and carried well-known favorable genetic changes. At the other extreme, SP-type tumors were usually high-grade glioblastomas lacking protective mutations and occurring more often in older patients. Even after statistically adjusting for tumor grade and key genetic markers, belonging to the SP-type still predicted a much poorer outlook, suggesting that how the tumor handles sphingolipids captures an independent dimension of danger that standard tests miss.
A hostile neighborhood inside the brain
Digging deeper, the authors examined the tumor microenvironment – the mix of immune cells, blood vessels, and supporting tissue around the cancer. SP-type tumors showed a crowded and conflicted immune landscape, with both attack-oriented and suppressive immune cells present, plus strong signals that dampen effective anti-tumor responses. Pathways linked to rapid cell growth, invasion, new blood vessel formation, scarring, and inflammation were all more active in this subtype. Measures that estimate how tumors respond to radiation suggested that SP-type gliomas are the most resistant to radiotherapy, dovetailing with their worse survival. In contrast, ST-type tumors looked more “calm,” with lower levels of these aggressive features.
Reading tumor metabolism from MRI scans
Because surgically removing tumor tissue is invasive and not always possible, the researchers explored whether standard magnetic resonance imaging (MRI) could hint at a tumor’s lipid behavior. They extracted more than two thousand subtle texture and shape features from two common MRI sequences and trained a machine-learning model to distinguish SP-type from all other tumors. The model performed well in both a hospital-based training set and an independent public validation set, correctly separating SP-type tumors much more often than chance. This suggests that the metabolic fingerprint of an especially aggressive glioma subtype leaves a detectable imprint on routine brain scans.
A key gene at the center of an aggressive network
To move from broad pathways to concrete targets, the team searched for genes that were central in lipid-related networks, strongly overactive in SP-type tumors, linked to worse survival, and accurate at telling SP-type tumors apart from others. Three genes – GLA, GLB1, and HSD3B7 – met all criteria. All were more active in SP-type gliomas and together formed a powerful diagnostic signature. The authors focused on HSD3B7, whose role in brain tumors had been largely unexplored. Tissue staining from 100 glioma patients showed that the HSD3B7 protein was higher in more advanced and more malignant tumors, and patients whose tumors had high levels of this protein lived significantly shorter lives.
How one lipid gene reshapes the tumor ecosystem
Single-cell analyses, which profile individual cells within tumors, revealed that HSD3B7 is active not only in cancer cells but also in several types of immune and support cells. High levels of this gene were associated with a web of signals that promote blood vessel growth, chronic inflammation, and immune evasion. Communication among certain protective cell types appeared weakened, while self-reinforcing loops within tumor-supporting cells were strengthened. Together, these patterns suggest that heightened HSD3B7 activity helps create and maintain a hostile microenvironment that favors tumor growth and resistance to therapy.
What this means for patients and future care
In practical terms, this work shows that gliomas can be meaningfully divided into three lipid-based subtypes, with the sphingolipid-heavy SP-type standing out as particularly dangerous and therapy-resistant. These differences are not just academic: they can be read from routine MRI scans using advanced image analysis and traced down to specific genes like HSD3B7 that may become future drug targets. While experimental work is still needed to test how blocking these lipid pathways might slow tumors or improve radiotherapy, the study offers a new metabolic lens through which to view brain cancer and moves the field closer to more personalized, biology-informed treatment decisions.
Citation: Tu, S., Zhang, P., Chi, X. et al. Lipid metabolism classification of gliomas. Sci Rep 16, 8219 (2026). https://doi.org/10.1038/s41598-026-37697-3
Keywords: glioma, lipid metabolism, brain tumor imaging, radiomics, tumor microenvironment