Clear Sky Science · en
Whole-transcriptome identification of deleterious variants in candidate genes linked to bovine paratuberculosis
Hidden Genes Behind a Costly Cattle Disease
Bovine paratuberculosis, also known as Johne’s disease, quietly saps the health and productivity of dairy herds around the world, costing farmers hundreds of millions of dollars each year and raising concerns about links to human intestinal disorders. This study peeks under the genetic hood of Holstein cows to ask a simple but crucial question: what differences in their genes might help some animals resist this infection while others succumb? By reading the RNA—the working copies of genes—in blood and gut tissue, the researchers traced how subtle DNA changes in key immune genes may steer the course of disease.
Why This Cow Disease Matters
Paratuberculosis is caused by the bacterium Mycobacterium avium subspecies paratuberculosis (MAP). Cows typically become infected early in life, but signs often appear years later. In the silent and subclinical stages, animals look healthy yet may shed low levels of bacteria and produce less milk. In clinical stages, they develop chronic diarrhea, severe weight loss, and major drops in milk yield. Herd-level infection rates can exceed 50% in many regions, including parts of Europe and North America, creating an economic and animal-welfare challenge. Because the bacteria may also act as an environmental trigger for human inflammatory diseases such as Crohn’s disease, there is growing pressure to improve control strategies in cattle.
Reading the Working Genome
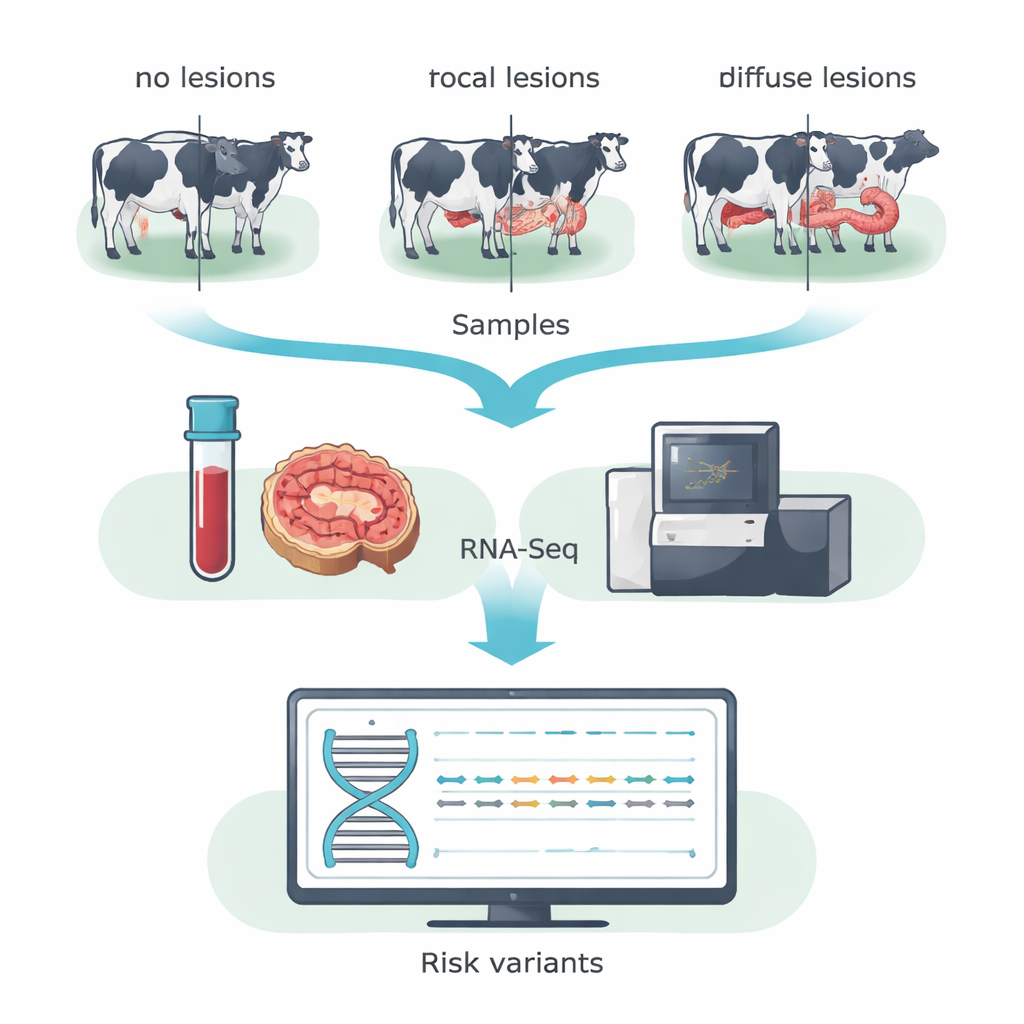
Instead of scanning the entire DNA sequence, the team used RNA sequencing (RNA-Seq) to focus on genes that are actually active in the animal. They collected blood and samples from the ileocecal valve—an intestinal region heavily affected by MAP—from 14 Holstein cows in a Spanish dairy herd. Based on microscopic examination of gut tissues, cows were grouped into three categories: animals without detectable lesions, animals with small, localized (focal) lesions typical of long subclinical infection, and animals with severe, widespread (diffuse) lesions associated with clinical disease. By merging data from blood and gut within each group, the researchers boosted their ability to detect single-letter DNA changes in expressed genes, known as coding SNPs.
Finding Harmful Changes in Key Immune Genes
From hundreds of thousands of variants, the team focused on those that change the protein sequence and are predicted to damage protein function—so‑called deleterious variants. They applied stringent filters to ensure high confidence, then used established prediction tools to flag risky changes. This winnowing process revealed 31 such variants unique to cows without lesions, 15 unique to cows with focal lesions, and 31 unique to cows with diffuse lesions. Many of these fell in genes that guide how immune cells recognize and clear infections, regulate cell death, and manage metabolism. A standout was the BOLA gene family, the bovine version of the major histocompatibility complex class II, which helps immune cells present bacterial fragments to T cells. Different potentially harmful variants in BOLA were found in all three cow groups, suggesting that specific BOLA versions may tilt animals toward resistance, controlled infection, or damaging inflammation.
From DNA Variants to Disease Pathways
To understand what these gene changes might mean in practice, the researchers examined which biological pathways were enriched in each group. Cows without lesions showed altered variants in genes linked to antigen processing, vesicle trafficking, and intestinal immune balance, including BOLA, AP3B1, and CHGA. These changes may promote efficient digestion of bacteria within immune cells and a stable gut environment that limits damage. In cows with focal lesions, harmful variants clustered in genes (ORMDL3 and KANK2) that dampen programmed cell death and tune cellular metabolism, potentially helping the host keep bacterial numbers low during a long, subclinical phase. In cows with diffuse lesions, the affected genes pointed to overactive immune pathways such as Th1/Th2 cell differentiation and antigen presentation, along with bile transport and drug-response routes. Here, altered BOLA family genes may drive a strong, sometimes self‑damaging inflammatory reaction, echoing patterns seen in many human autoimmune and inflammatory diseases.
Clues for Breeding More Resilient Herds
Beyond illuminating how different genetic variants shape the immune response to MAP, the study also linked these risky changes to known regions of the bovine genome associated with health traits, including susceptibility to paratuberculosis and other infections. While these results still require validation in larger herds and cannot yet be used as stand‑alone diagnostic markers, they offer a promising catalogue of candidate variants and genes. In plain terms, the work suggests that some cows carry gene versions that help them quietly control infection, while others carry versions that favor runaway inflammation and severe disease. In the future, such information could support selective breeding and genetic tests that shift herds toward greater natural resistance, reducing both economic losses and the need for intensive disease control measures.
Citation: Badia-Bringué, G., Lam, S., Cánovas, Á. et al. Whole-transcriptome identification of deleterious variants in candidate genes linked to bovine paratuberculosis. Sci Rep 16, 6243 (2026). https://doi.org/10.1038/s41598-026-37675-9
Keywords: bovine paratuberculosis, Johne’s disease genetics, dairy cattle immunity, RNA sequencing, disease resistance breeding