Clear Sky Science · en
Leveraging medical imaging and deep learning for diagnosis of breast cancer using histopathological images
Why early detection matters
Breast cancer is one of the leading causes of cancer death in women worldwide, but outcomes improve dramatically when the disease is caught early. Doctors usually diagnose breast cancer by examining tiny slices of tissue under a microscope, a process called histopathology. These images contain rich detail about whether cells are harmless or dangerous, yet reading them is time‑consuming and can vary from one specialist to another. This study explores how modern artificial intelligence can help pathologists spot breast cancer more quickly and consistently, potentially giving patients faster answers and more effective treatment options.
A closer look at tissue images
Under the microscope, breast tissue does not neatly separate into “healthy” and “cancerous.” Cells overlap, colors vary from lab to lab, and subtle changes in shape or texture can carry life‑or‑death meaning. Traditional computer‑aided systems struggled with this complexity because engineers had to hand‑design the features the computer should look for, and small changes in staining or image quality could throw them off. Deep learning, a branch of artificial intelligence that learns patterns directly from data, has recently transformed how computers interpret images, including medical scans. The authors build on this progress to design a system tailored to the messy reality of breast tissue slides.
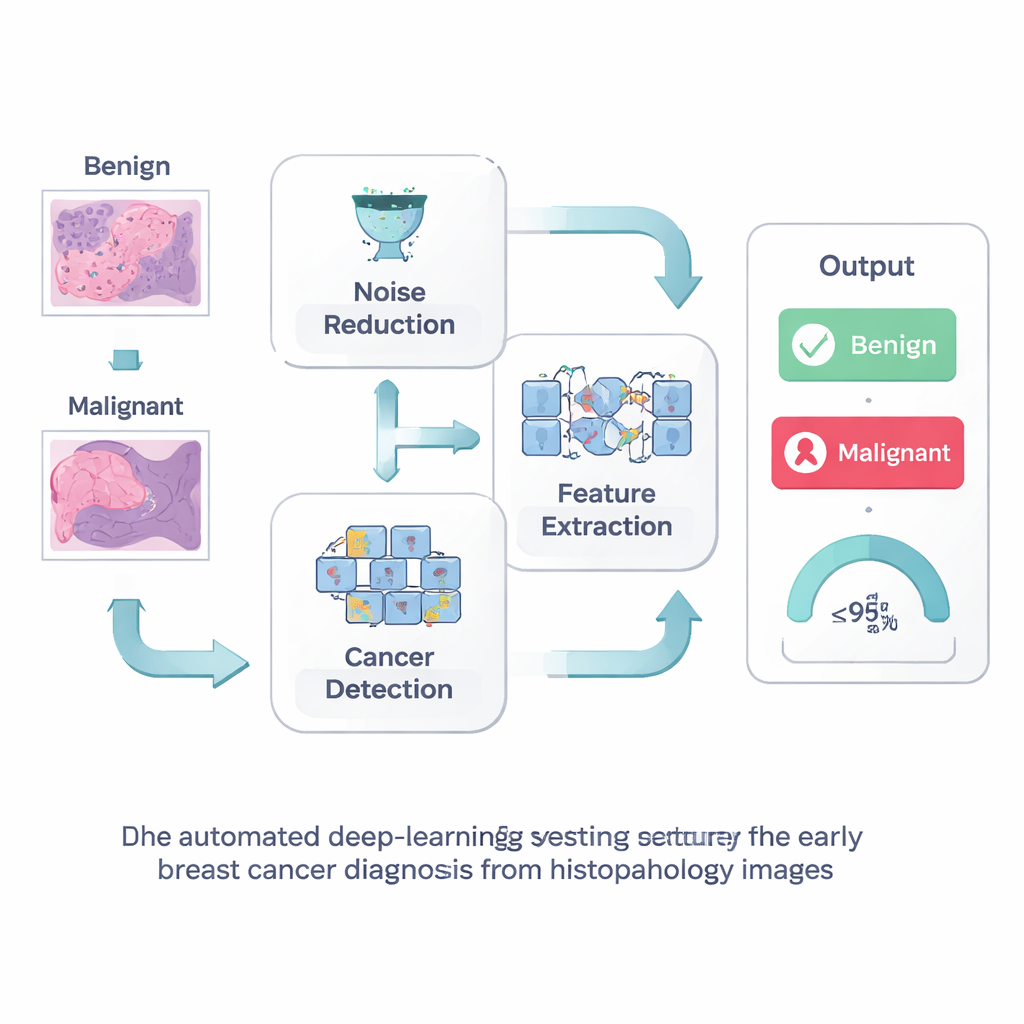
Cleaning the picture before reading it
The first step in their approach is simple but powerful: clean up the image before asking a computer to interpret it. Histopathology slides often contain visual “noise” from the staining and imaging process, which can obscure the fine structures that signal early cancer. The researchers use a technique called Wiener filtering, which smooths away random speckles while preserving sharp edges and tiny details such as cell borders and small clusters. By presenting a clearer picture to the computer, this step helps avoid both missed cancers and false alarms that could send patients for unnecessary tests.
Teaching the computer what to pay attention to
Next, the team turns to a sophisticated deep‑learning model known as SE‑ResNet to study the cleaned images. In simple terms, this model scans the slide patch by patch, gradually building up an internal “vocabulary” of visual patterns: how normal ducts look, how tumor cells group together, and how textures change as cancer becomes more aggressive. A built‑in attention mechanism helps the network emphasize the most informative image channels and downplay irrelevant background. This makes the model more sensitive to subtle, disease‑related patterns while keeping computation efficient enough that it could run on real‑world hospital hardware.
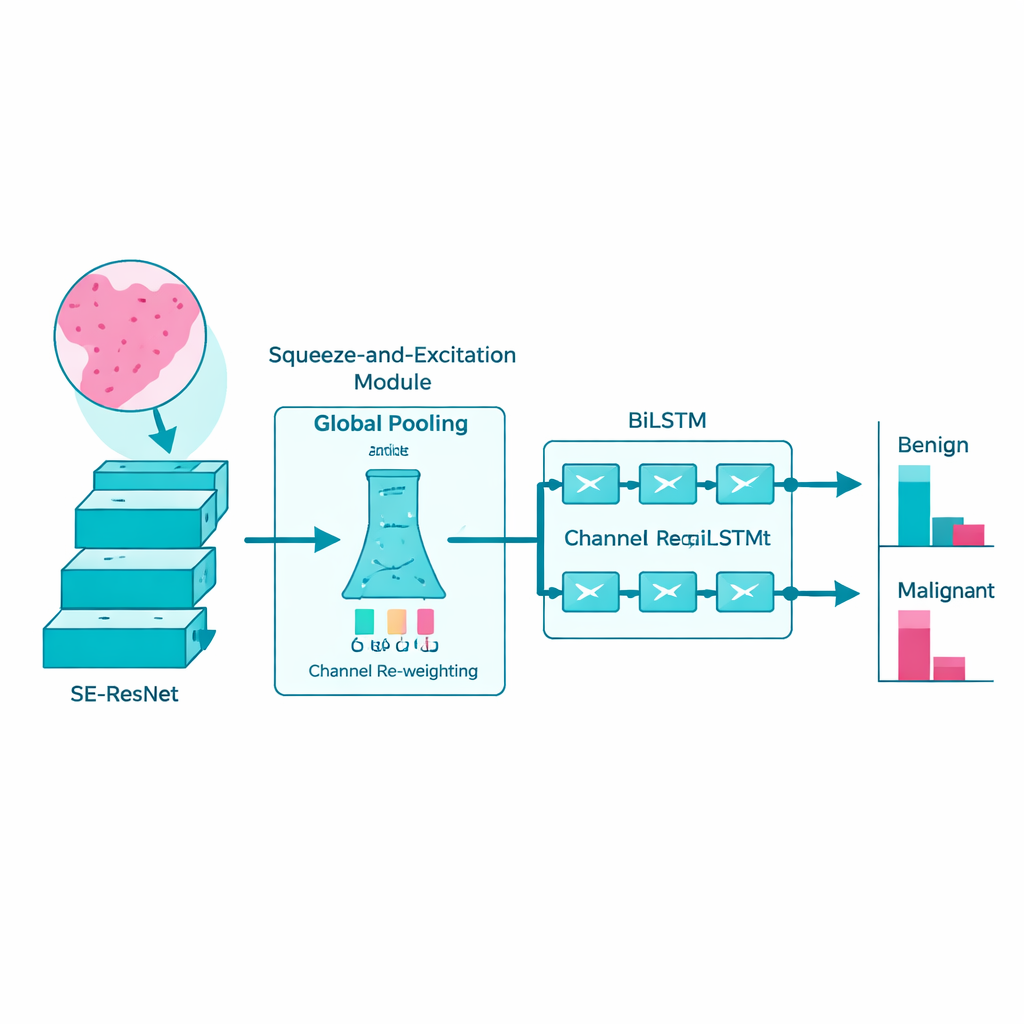
Following patterns across space like a story
Rather than treating each patch of tissue as an isolated snapshot, the researchers recognize that disease signs often unfold like a story across the slide. To capture this, they feed the features extracted by SE‑ResNet into a bi‑directional long short‑term memory network, or BiLSTM. This type of model is designed to understand sequences: it looks at how patterns change from one region to the next, both forward and backward, much like reading a sentence in both directions to grasp its full meaning. By learning these spatial relationships, the BiLSTM becomes better at distinguishing benign changes from truly malignant ones.
How well the system works in practice
The authors tested their full pipeline—noise reduction, feature learning, and sequence modeling—on large public collections of breast tissue images, including the widely used BreakHis dataset. They split the data into training and testing groups at different ratios and compared their method against many established deep‑learning models. Across these experiments, their system correctly classified benign versus malignant samples nearly 99% of the time, outperforming competing methods while also running faster. The model remained strong across different magnifications of the tissue, suggesting it can adapt to slides prepared under varying conditions. However, the study also notes limitations: the datasets are still modest in size, the model focuses on a simple two‑class decision rather than detailed tumor subtypes, and it has not yet been proven in real‑world clinical workflows.
What this means for patients and doctors
To a lay person, the take‑home message is that computers are getting much better at reading microscope images of breast tissue and flagging suspicious areas. The proposed system does not replace the pathologist; instead, it acts like a highly attentive assistant that highlights regions likely to be cancerous and provides a second opinion with very high accuracy. If validated in larger and more diverse patient groups, such tools could shorten the time to diagnosis, reduce the chance that a small cancer is missed, and help overstretched hospitals manage growing caseloads. Future work will need to test the method on more varied slides and integrate it into everyday lab routines, but this study shows that carefully designed deep‑learning systems can be a powerful ally in the fight against breast cancer.
Citation: Nagalakshmi, V., Ahammad, S.H. Leveraging medical imaging and deep learning for diagnosis of breast cancer using histopathological images. Sci Rep 16, 6236 (2026). https://doi.org/10.1038/s41598-026-37663-z
Keywords: breast cancer diagnosis, histopathology images, deep learning, medical imaging, computer-aided detection