Clear Sky Science · en
Revisiting bioluminescence and sucrose utilization in aquatic pathogens Vibrio harveyi and V. campbellii using genome-wide in silico mapping and phenotyping
Why glowing germs matter to seafood
For shrimp and fish farmers, mysterious nighttime glows in hatchery tanks can signal disaster. Two nearly twin bacteria, Vibrio harveyi and Vibrio campbellii, are behind many deadly outbreaks, yet they are so similar that even specialists often mix them up. This study revisits two simple traits long used to tell them apart—whether they glow in the dark and whether they can digest common table sugar—and combines field tests with modern genome analysis to clarify who is who and how that affects disease control in aquaculture.
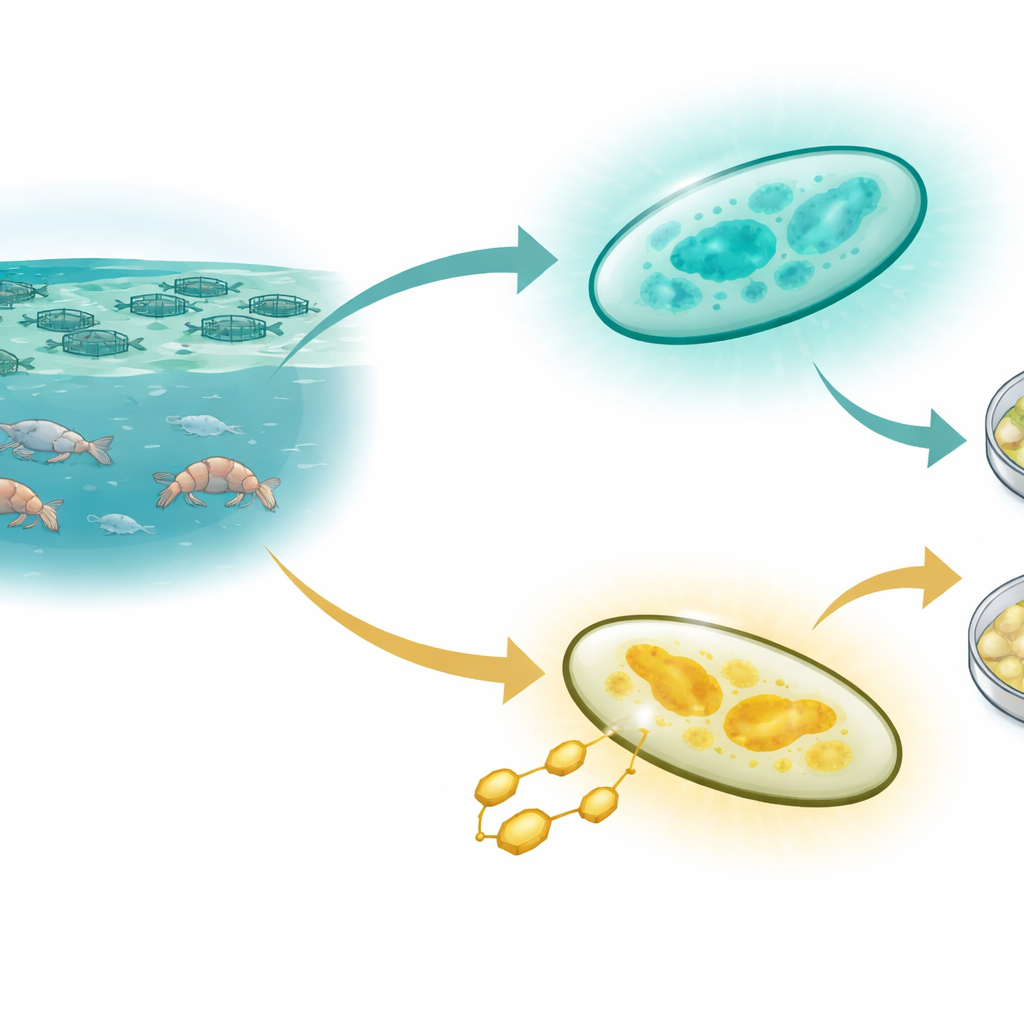
Two look‑alike troublemakers
Both Vibrio harveyi and Vibrio campbellii live in seawater and can cause mass deaths of shrimp and marine fish in farms around the world. Under the microscope and in basic DNA tests, they look almost identical. For decades, workers in hatcheries have relied on two quick clues: glowing colonies that suggest a dangerous "luminescent" infection, and the color of colonies on a standard lab plate containing the sugar sucrose. In theory, these traits should separate species neatly, but real‑world observations have been confused and sometimes contradictory, leading to misdiagnosed outbreaks and uncertainty about which species is actually present.
Reading entire genomes for a clearer picture
The researchers assembled high‑quality, near‑complete genomes for one Vibrio harveyi strain and six Vibrio campbellii strains, then compared them to more than 300 publicly available genomes from the broader group to which these bacteria belong. By measuring overall DNA similarity and building evolutionary family trees from thousands of shared genes, they corrected many past mislabels and firmly separated 204 strains as V. harveyi and 78 as V. campbellii. These analyses also showed that V. harveyi represents an older, genetically tight lineage, while V. campbellii is a younger, more variable offshoot that is still splitting into sub‑groups.
Who really glows and who eats sugar
With species identities clarified, the team searched each genome for the clusters of genes that power light production and sugar use. The genes for bioluminescence were almost completely missing from V. harveyi: only about 3% of its strains still carried a full set. In contrast, every V. campbellii strain had either a complete, working light‑making pathway or a damaged version of it. The picture was flipped for sucrose: nearly 90% of V. harveyi strains carried a full set of sucrose‑use genes, while essentially all V. campbellii strains lacked them, apart from a single apparent recent acquisition. In lab tests on 49 live isolates, these genetic patterns matched real behavior: almost all V. campbellii strains glowed strongly but could not grow on sucrose, whereas all tested V. harveyi strains formed yellow, sucrose‑fermenting colonies and did not glow.
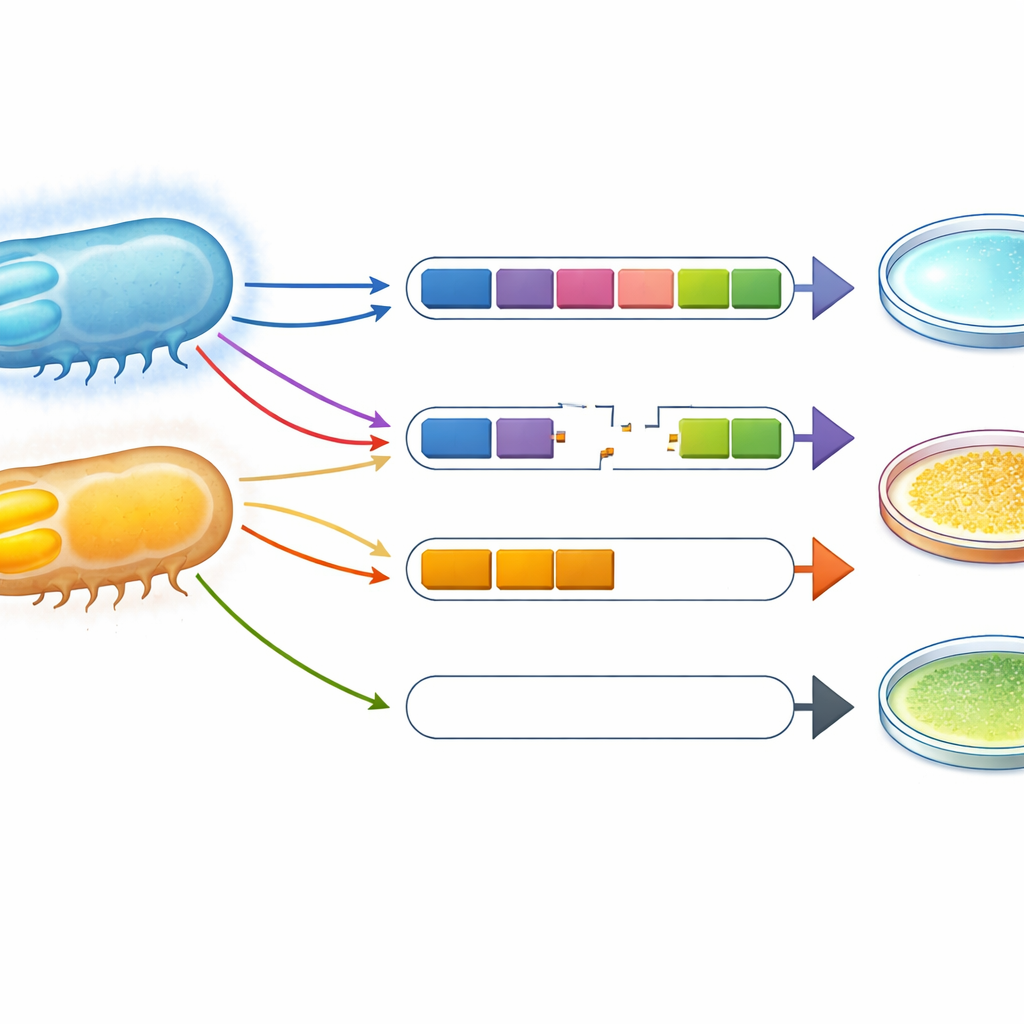
How these traits evolved in the sea
By examining the order of genes around the light and sugar clusters, and nearby mobile DNA elements such as insertion sequences, the authors traced how these traits likely spread and changed. In V. campbellii, light‑making genes sit in a stretch of DNA that, in some sub‑groups, is flanked by mobile elements, suggesting it has been reshaped over time and occasionally broken, creating non‑glowing lineages. In a few V. harveyi and V. campbellii strains, sucrose‑use genes appear to have jumped in from distant relatives, again via mobile DNA. Taken together with where strains were isolated—from host animals versus open ocean—these patterns suggest that V. harveyi has specialized as a host‑attached, sugar‑using, mostly dark bacterium, while V. campbellii has remained a more free‑living, light‑producing generalist.
What this means for farms and diagnostics
For farmers and field laboratories, the study offers a practical and reassuring message: simple plate tests remain powerful when interpreted correctly. The evidence shows that, in most cases, yellow, sucrose‑fermenting colonies on standard Vibrio plates are V. harveyi, while glowing, sucrose‑negative, green colonies are V. campbellii. Because mislabeling in the past often reversed these assumptions, this work helps straighten out the record. It also highlights how combining modern genome mapping with straightforward visual traits can sharpen disease diagnosis and guide better responses to outbreaks in the fast‑growing aquaculture industry.
Citation: Kumar, S., Nishanthini, B., Robinson, A. et al. Revisiting bioluminescence and sucrose utilization in aquatic pathogens Vibrio harveyi and V. campbellii using genome-wide in silico mapping and phenotyping. Sci Rep 16, 8678 (2026). https://doi.org/10.1038/s41598-026-37651-3
Keywords: aquaculture disease, bioluminescent bacteria, Vibrio pathogens, bacterial genomics, shrimp hatcheries