Clear Sky Science · en
A genome-structure adaptive framework for ROH-based inbreeding estimation in Penaeus vannamei
Why shrimp family trees matter for your dinner plate
Modern shrimp farms supply much of the world’s seafood, but breeding the same family lines over and over can quietly erode their health. When close relatives mate, harmful hidden genes can pair up, reducing growth, survival and disease resistance. This study asks a deceptively simple question with big implications for global aquaculture: how can we measure inbreeding in farmed shrimp accurately enough to keep stocks healthy and productive?
Hidden genetic footprints of inbreeding
Inbreeding leaves telltale fingerprints in DNA. Instead of carrying two slightly different versions of many genes, inbred animals tend to have long stretches where both copies are identical. Geneticists call these stretches “runs of homozygosity,” or ROH. By adding up how much of an animal’s genome falls into these runs, researchers can estimate its inbreeding level more precisely than by tracking paper pedigrees, which are often incomplete or contain errors. This ROH-based measure, known as FROH, has become standard in cattle, pigs and other livestock, but shrimp genomes pose special challenges that make off‑the‑shelf methods unreliable.
Why shrimp genomes are especially tricky
Whiteleg shrimp (Penaeus vannamei), the world’s most widely farmed shrimp, have highly fragmented and complex genomes. Instead of long, continuous chromosome sequences, many available genome maps are broken into thousands of smaller pieces, separated by gaps and repetitive regions. Genetic markers are unevenly scattered across this patchwork, and the species has very high genetic diversity. Methods and software settings originally tuned for tidy, well‑mapped mammal genomes can therefore mistake technical gaps for real breaks in ROH, or confuse short, background similarities with true signs of inbreeding. The result is a high risk of mis‑estimating how inbred a shrimp actually is.
Building a genome-aware measuring stick
To solve this, the authors designed a “genome‑structure adaptive” framework that tailors ROH analysis to the quirks of shrimp DNA. They created thirteen highly inbred shrimp families under controlled matings, then deeply sequenced the genomes of five of these families plus their parents. Crucially, they aligned the same sequencing data to two very different reference genomes: one older, fragmented assembly and a newer, highly continuous version. Using the common analysis tool PLINK, they systematically tested how eight key settings affected ROH calls, focusing on three that were especially sensitive to genome structure: how densely markers must be packed, how large a gap between markers is allowed inside a run, and how long a run must be to count. They built empirical, non‑overlapping genomic windows to track local marker spacing and missing data, and used “genomic coverage” of these windows, along with stability of FROH and ROH lengths, as objective guides for choosing sensible thresholds.
Different dials, same inbreeding picture
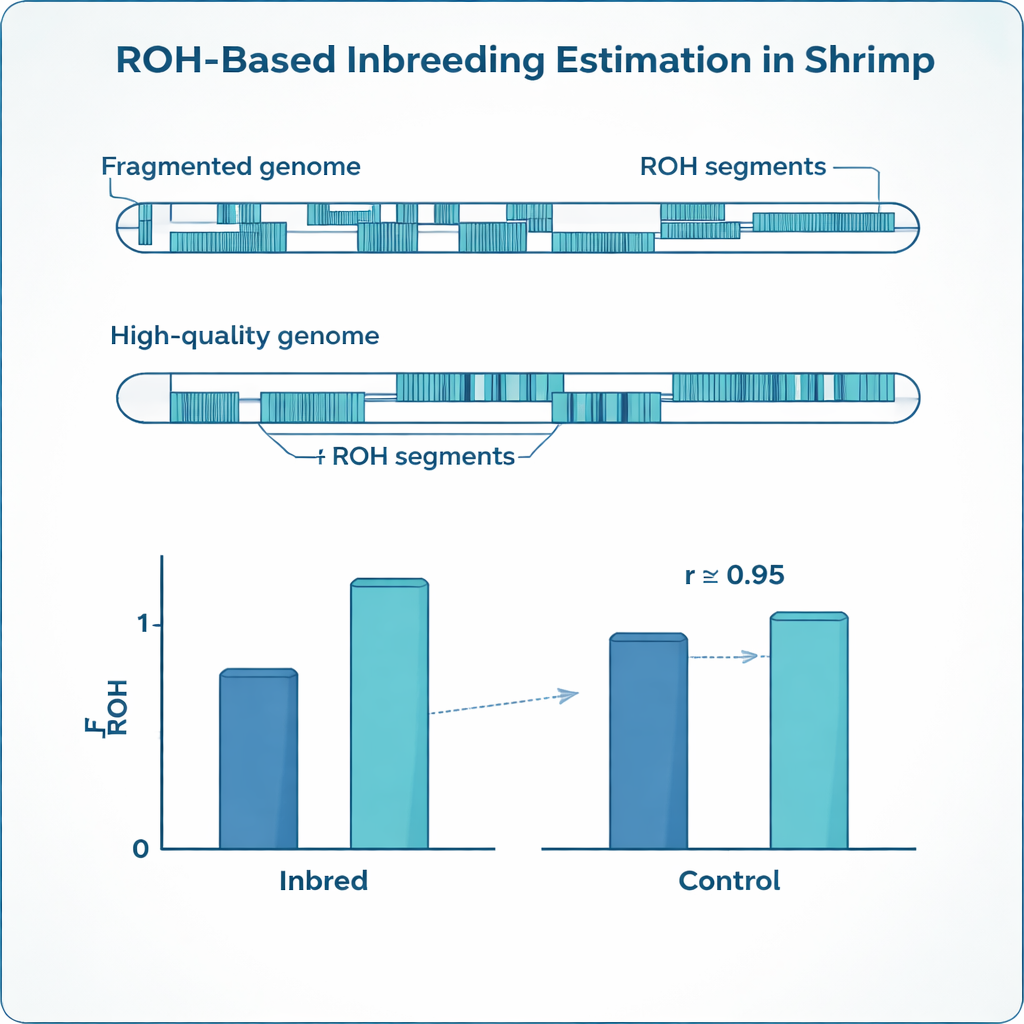
The optimized settings turned out to be very different for the two reference genomes. The fragmented genome required much denser markers, shorter allowed gaps and shorter minimum run lengths than the high‑quality genome to avoid breaking true ROH into many small pieces. Yet, after tuning these dials separately, the inbreeding estimates from both references converged: average FROH in the inbred shrimp was about 0.24 in both cases, closely matching the expected value from the planned matings and strongly agreeing with each other. At the same time, the more continuous genome revealed fewer but much longer ROH segments, while the fragmented map chopped many of those into short stretches. The study also exposed striking differences among full‑sibling shrimp: even within the same family, inbreeding levels varied widely, something that simple pedigree records cannot capture.
Sharper tools for healthier shrimp stocks
For non‑specialists, the take‑home message is straightforward: measuring inbreeding from DNA can be highly accurate in farmed shrimp, but only if the method respects the underlying genome’s structure. By providing a practical recipe for tuning ROH analysis to fragmented crustacean genomes, this work allows breeders to monitor inbreeding animal by animal, rather than relying on imperfect family charts. That, in turn, can help design mating plans that maintain genetic diversity while still improving growth and resilience, supporting more sustainable shrimp farming and offering a template for other aquaculture species facing similar genomic hurdles.
Citation: Zou, X., Zhou, H., Liu, M. et al. A genome-structure adaptive framework for ROH-based inbreeding estimation in Penaeus vannamei. Sci Rep 16, 6769 (2026). https://doi.org/10.1038/s41598-026-37622-8
Keywords: shrimp breeding, inbreeding, genomic selection, runs of homozygosity, aquaculture genetics