Clear Sky Science · en
Computational identification of multi-target natural compounds from Sesbania grandiflora as potential therapeutic agents against Klebsiella pneumoniae
Why an Old Remedy Matters for New Superbugs
Antibiotic-resistant infections are becoming harder and harder to treat, and one of the worst culprits is a hospital germ called Klebsiella pneumoniae. When this bacterium outsmarts our best drugs, doctors can run out of options. This study asks a hopeful question: could natural chemicals from a traditional medicinal tree, Sesbania grandiflora (also known as the vegetable hummingbird), be redesigned into modern medicines that hit the bacterium in several weak spots at once?
A Tough Bacterium That Outsmarts Drugs
Klebsiella pneumoniae causes serious illnesses such as pneumonia, urinary tract infections, sepsis, and meningitis, especially in people with weakened immune systems. Over the past decade, many strains have become resistant even to carbapenems, antibiotics usually saved as a “last resort.” The bacterium defends itself by breaking down drugs, pumping them out, sealing up its outer membrane, and forming protective biofilms. Because it uses many different tricks at once, a single-target drug is easy for it to evade. The authors argue that a better strategy is to block several vital processes together, making it much harder for the microbe to evolve resistance.
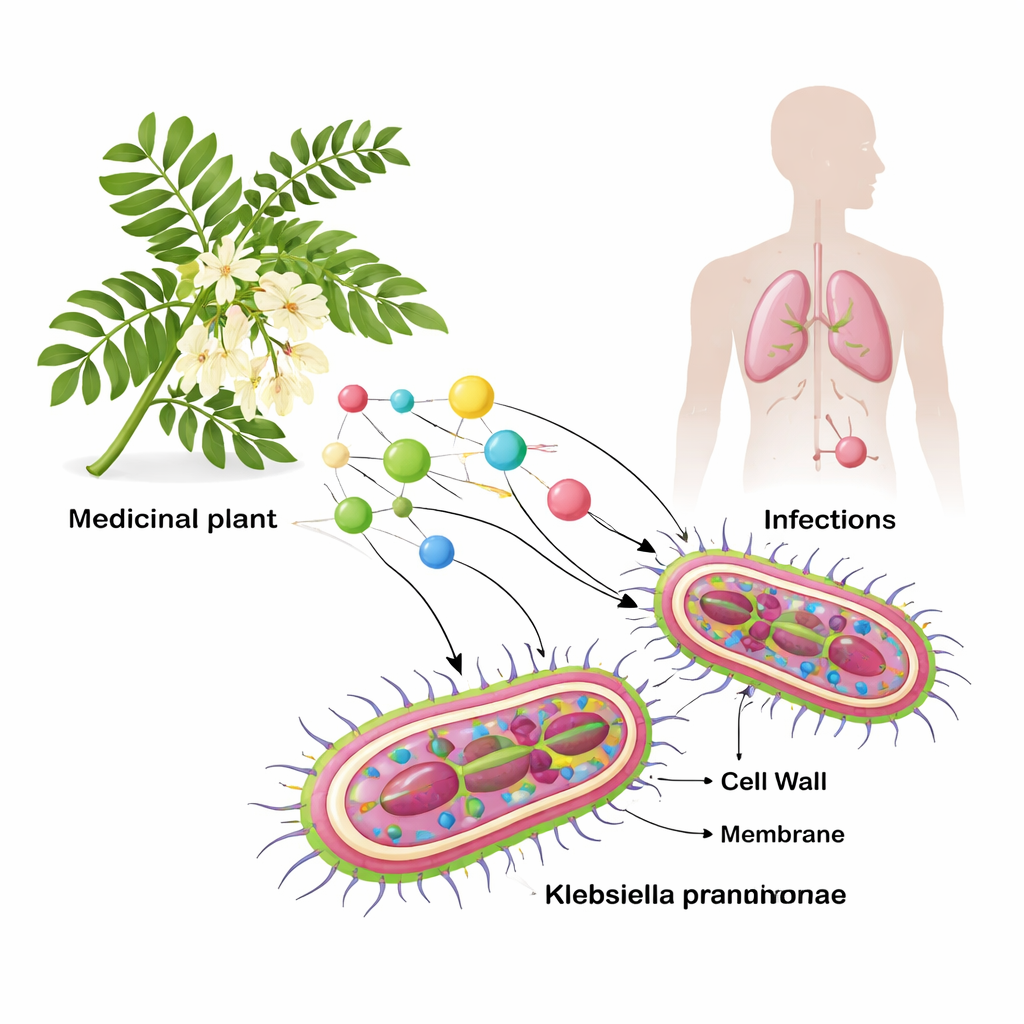
Looking to a Medicinal Tree for Clues
Sesbania grandiflora has a long history in traditional medicine and has shown activity against various disease-causing microbes. Instead of testing plant extracts directly in the lab, the researchers used computers to conduct a large virtual screening. They first combed through the complete set of Klebsiella proteins and, using strict quality and literature filters, narrowed the list down to six especially important bacterial components. These include enzymes needed for building the outer coat and cell wall, a resistance enzyme that destroys powerful antibiotics, and a structural protein that helps the bacterium interact with human cells. In parallel, they gathered 73 known plant chemicals from S. grandiflora and prepared three‑dimensional models of both the compounds and the bacterial targets.
Finding Plant Compounds That Hit Many Targets
The team used molecular docking, a technique that predicts how well each plant molecule might fit into the pockets of each bacterial protein. Most of the tested compounds could latch on to all six targets with favorable predicted strength, and a smaller group still showed especially strong binding. A network analysis revealed that three targets in particular—FabG, KPC‑2, and OmpA—were connected to almost all of the compounds, hinting at broad coverage. The researchers then applied standard “drug‑likeness” rules and computer models of absorption and toxicity to filter out molecules unlikely to work safely in the body. This winnowing process reduced the list to nine promising candidates, many sharing structural features—such as ring-shaped cores and multiple oxygen-rich groups—that help them form stable contacts with proteins.
Zooming In on a Star Candidate
Among these nine, one compound, Sonchuionoside A, stood out because it docked strongly to all six bacterial targets while also satisfying the safety and usability filters. To test how robust these interactions might be, the authors ran long molecular dynamics simulations, essentially watching how each protein and the compound move together over hundreds of nanoseconds in a virtual water box. Across all six targets, Sonchuionoside A remained bound without disrupting protein stability and often made the proteins slightly more compact and orderly. Detailed analyses of motion, surface exposure, hydrogen bonds, and estimated binding energies suggested especially strong and favorable interactions with two enzymes involved in building essential cell components (LpxH and FabG), along with solid binding to the resistance enzyme KPC‑2 and the outer-membrane protein OmpA.
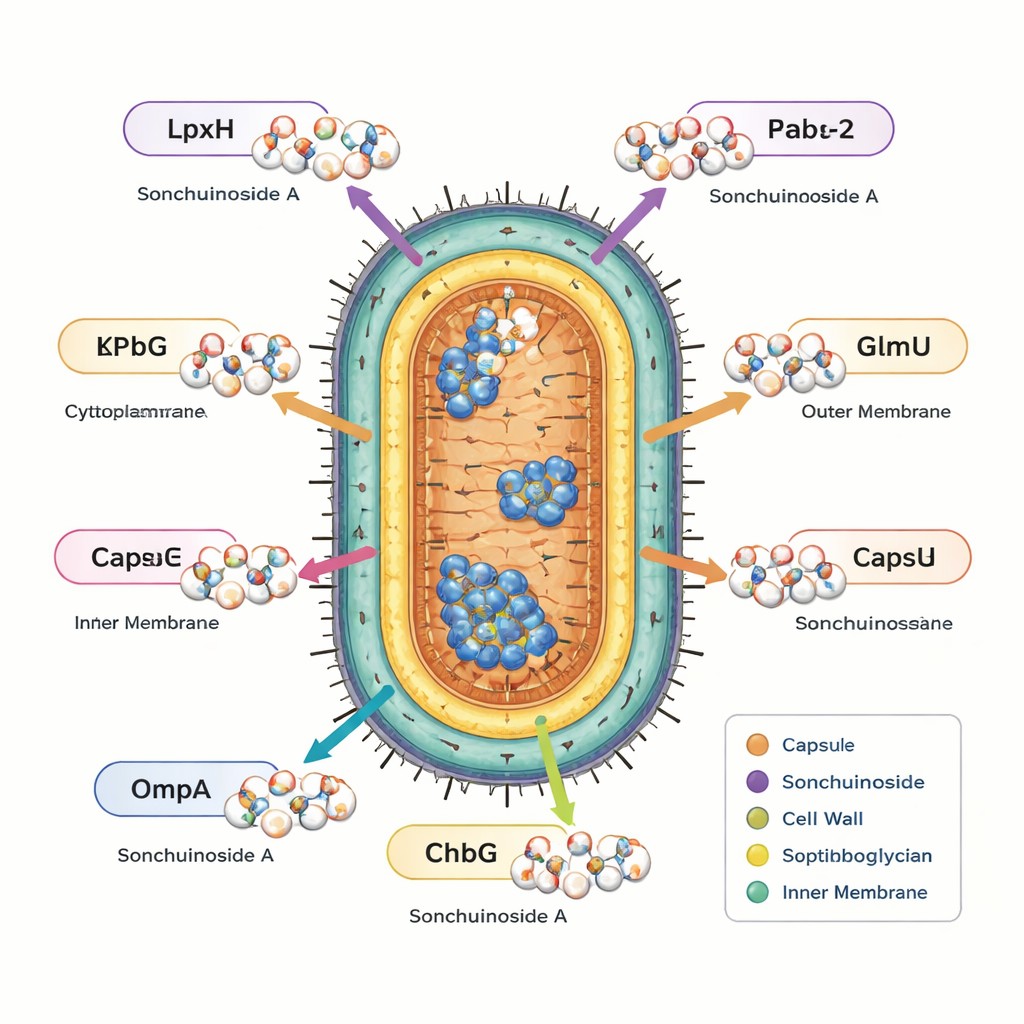
What This Could Mean for Future Treatments
This research does not yet deliver a ready-to-use drug, and all of the findings come from computer models rather than animal or human tests. Still, it provides a compelling roadmap. The work shows that a single plant compound like Sonchuionoside A might be engineered to attack Klebsiella pneumoniae on multiple fronts—weakening its cell defenses, undermining its resistance machinery, and blunting its ability to cause disease. In simple terms, the study suggests that a traditional medicinal tree could inspire future multi-pronged antibiotics, helping doctors stay one step ahead of dangerous, drug-resistant infections.
Citation: Sajal, H., Mohan, A., Ravi, V. et al. Computational identification of multi-target natural compounds from Sesbania grandiflora as potential therapeutic agents against Klebsiella pneumoniae. Sci Rep 16, 7782 (2026). https://doi.org/10.1038/s41598-026-37613-9
Keywords: antibiotic resistance, Klebsiella pneumoniae, medicinal plants, natural compounds, multi-target drugs