Clear Sky Science · en
DNA conjugation on functionalized plastic surfaces for sequential, iterative single molecule sequencing
Storing Tomorrow’s Data in Tiny Strands
Imagine backing up your photos, books, or scientific records in a form that could last for centuries and fit into a speck smaller than a grain of sand. DNA—the same molecule that carries our genes—is emerging as a powerful candidate for ultra-dense, long-lasting data storage. This paper explores a new way to “park” DNA data safely on the inside of ordinary plastic lab tubes, then read it out again and again without destroying the original molecules.
A New Kind of Data ‘Thumb Drive’
Today’s hard drives and flash memory wear out, and they pack far less information per gram than DNA potentially can. Researchers have already shown how to translate digital files into sequences of DNA “letters.” But each time those DNA strands are copied or sequenced, some of the original material is used up, like ink fading as you photocopy a page too many times. In this study, the authors turned the humble plastic PCR tube—already standard in biology labs—into a reusable physical storage device. They chemically attached DNA strands carrying encoded data to the inner surface of the tube, so the DNA stays fixed in place while copies are repeatedly made and read. 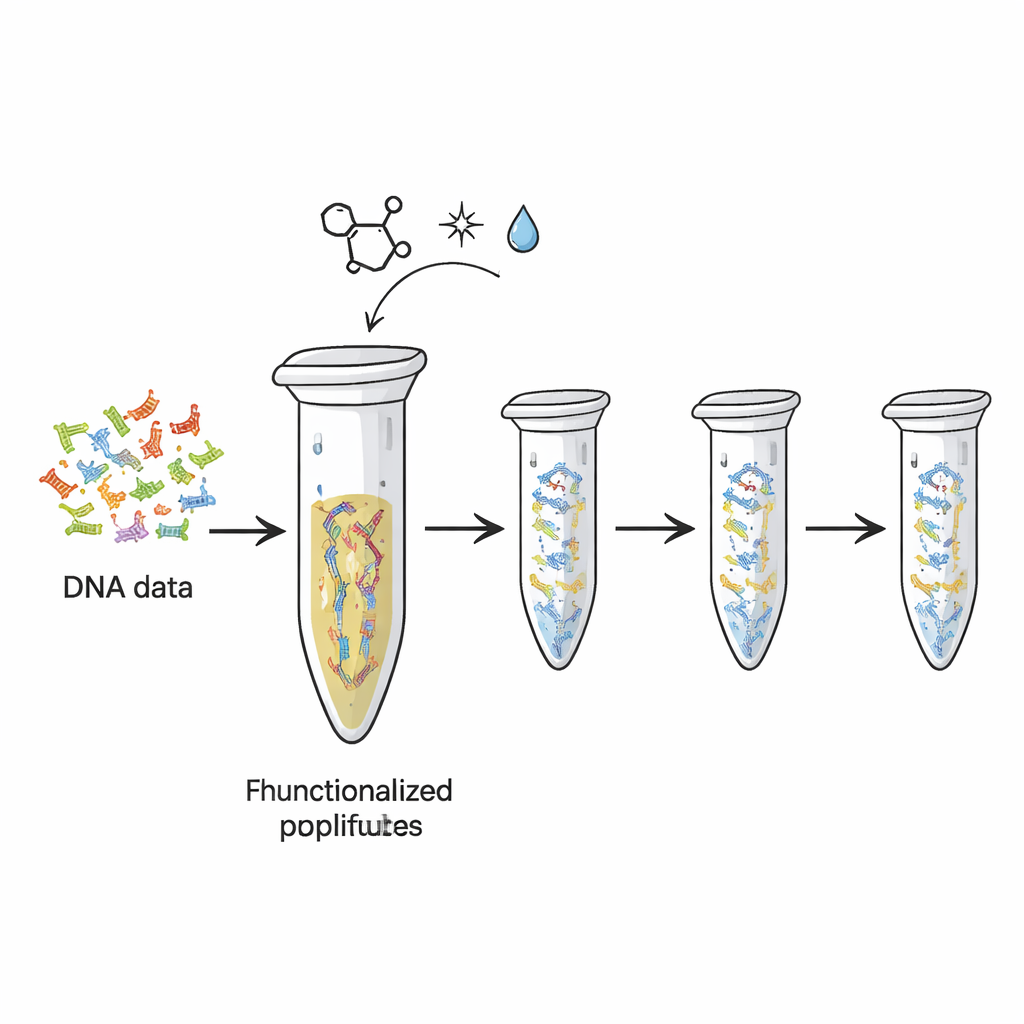
Clicking DNA onto Plastic
The team relied on a type of highly specific ‘click chemistry’—a set of reactions prized for being fast, reliable, and gentle. First, they added special chemical tails to the ends of DNA strands that represent different data “files.” These tails, based on a molecule called TCO, are designed to lock onto matching groups (MTz) that have been grafted onto the plastic surface of the tube. When the two meet, they form a stable covalent bond, effectively gluing the DNA to the plastic. Tests with a control DNA fragment showed that after incubation, nearly all of the DNA had disappeared from solution, indicating it was now immobilized on the tube wall. The platform could hold on the order of hundreds of femtomoles of DNA, suggesting ample capacity for practical data sets.
Retrieving Digital ‘Files’ on Demand
To see whether this surface-bound DNA still behaved like a workable archive, the researchers encoded text and other data into a pool of about 15,000 short DNA strands, grouped into 18 “file groups.” Each group could be selectively copied using its own pair of primers—short starting sequences that direct the copying machinery to the right targets. The team repeatedly performed standard PCR reactions in the same tube, each time choosing a different file group to amplify. After each run, they removed the copied DNA, cleaned the tube with enzymes that digest leftover products in solution, and moved on to the next file. Nanopore sequencing of the copied material showed that most file groups were retrieved with high accuracy, and cross-contamination from earlier runs remained extremely low, typically around 1 percent or less.
Finding a Gentler Way to Read
However, there was a catch: as they repeated PCR on the same tube up to 18 times, the amount of DNA retrieved steadily fell. PCR relies on rapid heating and cooling, and the authors inferred that the repeated high temperatures were damaging the immobilized DNA or its connection to the plastic, even though control tests suggested the DNA was not simply washing off into solution. To solve this, they turned to recombinase polymerase amplification (RPA), a newer method that works at a single, comparatively low temperature close to body heat. Using RPA on fresh DNA-coated tubes, they again queried all 18 file groups in sequence. This time, the yields were high—around 60 ng/µL—and did not show the same downward trend. The pattern of which strands were favored or disfavored during copying also closely matched what was seen when the DNA was free in solution. 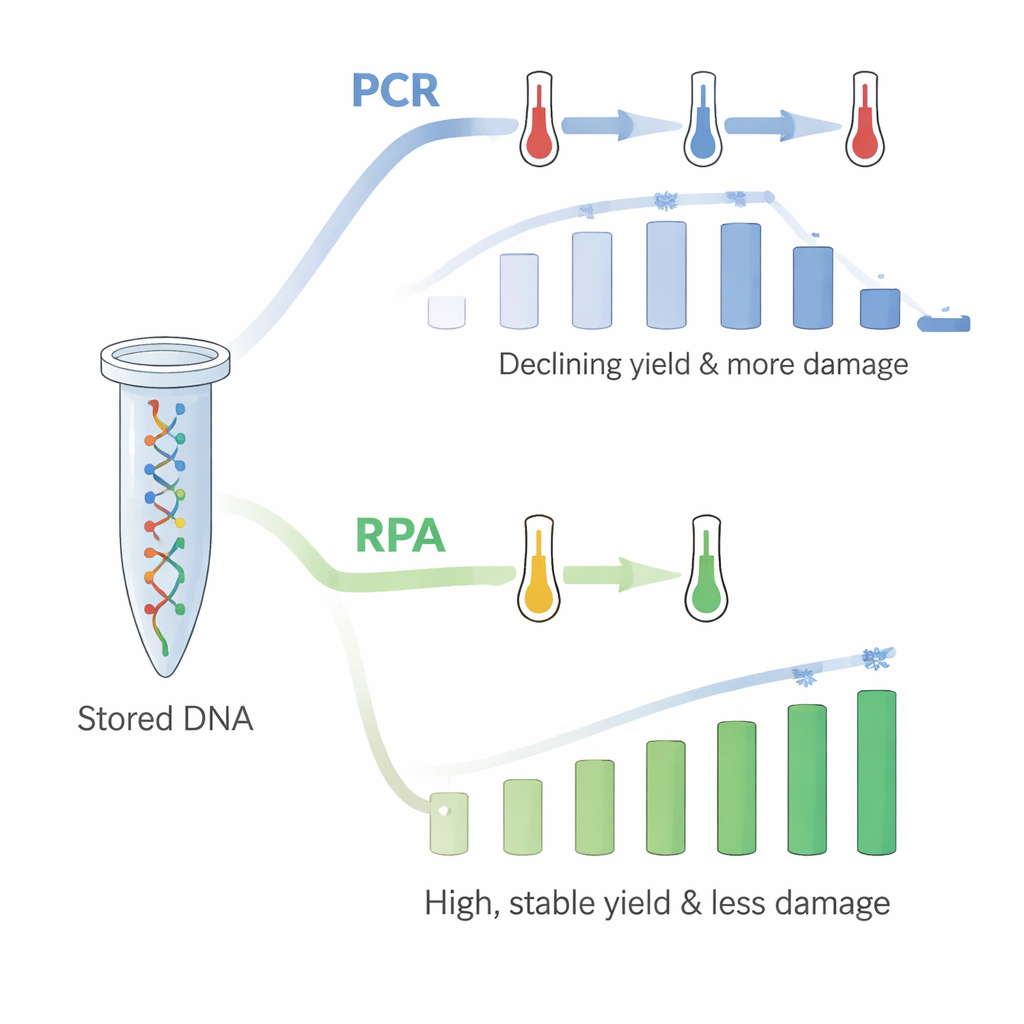
Toward Portable, Long-Lived DNA Memory
By combining robust click chemistry with low-temperature DNA copying, this work points to a practical way to turn simple plastic tubes into reusable DNA data cartridges. The DNA stays physically locked to the tube, where it can be repeatedly queried for specific “files” without consuming the original molecules, especially when using the gentler RPA method. For non-specialists, the key message is that DNA is not just the code of life—it can also act as a compact, durable storage medium for digital information. Methods like this bring us closer to a future where your long-term backups might reside not on spinning disks, but in carefully engineered molecules sitting quietly on a lab shelf.
Citation: Roy, S., Ji, H.P. & Lau, B.T. DNA conjugation on functionalized plastic surfaces for sequential, iterative single molecule sequencing. Sci Rep 16, 6467 (2026). https://doi.org/10.1038/s41598-026-37575-y
Keywords: DNA data storage, click chemistry, plastic surface conjugation, recombinase polymerase amplification, nanopore sequencing