Clear Sky Science · en
Prevalence and genotyping of Pseudomonas aeruginosa from food and human sources
Why a Common Germ Matters to Everyday Life
Pseudomonas aeruginosa is a mouthful to say, but it is a germ that can quietly travel from water and raw foods into hospitals and people, sometimes causing serious infections that are hard to treat. This study, carried out in an Egyptian city, traced where this bacterium shows up in everyday items like tap water, fish, meat, and milk, and in patients’ wound infections. It also examined how tough it is against antibiotics and whether it carries genetic “tools” that make it more dangerous. The findings help explain why food hygiene, clean water, and careful antibiotic use are all tied together in protecting public health.
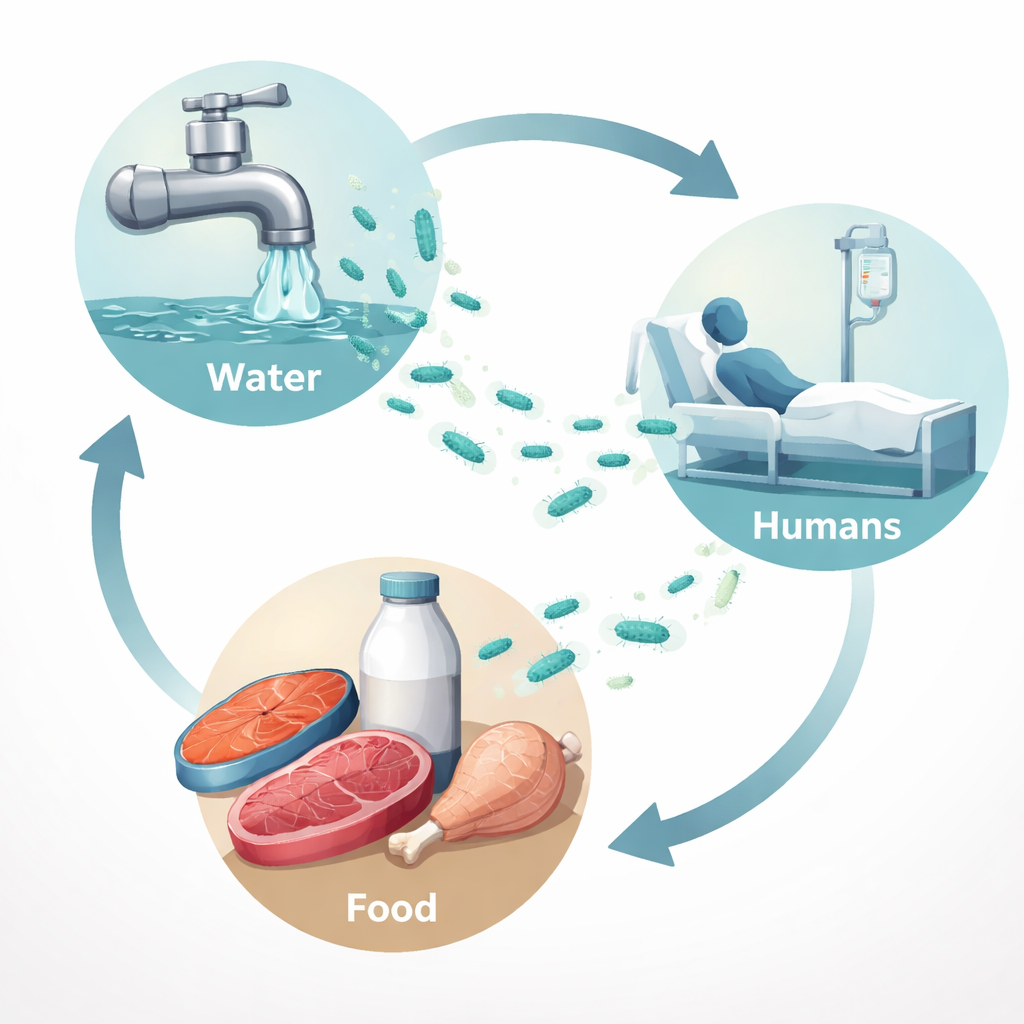
Following the Germ from Market to Hospital
The researchers collected 350 samples from around Sadat City in Menoufia Governorate, Egypt. These included human pus from infected wounds, tap water, fish swabs, chicken meat, minced meat, raw milk, and hospital surfaces. They were looking specifically for P. aeruginosa. Overall, they found the bacterium in about one in seven samples. It appeared most often in human wound samples (about one in four), but it was also present in tap water, fish, and raw milk, and to a lesser extent in chicken and minced meat. Interestingly, none of the hospital surface swabs tested positive, suggesting that in this setting, the bigger threat may come from food and water entering the hospital, rather than from contaminated counters and equipment.
How Dangerous Are These Bacteria?
Finding the germ is only part of the story; how it behaves matters just as much. The team tested each bacterial isolate using standard laboratory methods to see whether it could form a slimy protective layer called a biofilm. About one‑third of the isolates did so, with some forming strong, thick layers. Biofilms make bacteria much harder to kill because they shield cells from cleaning agents and antibiotics. Genetic tests on the biofilm‑forming strains showed that they carried multiple “virulence” genes, which encode proteins that damage host tissues or help the bacteria cling to surfaces and evade the immune system. In simple terms, many of the strains they found were not just present—they were well equipped to cause disease.
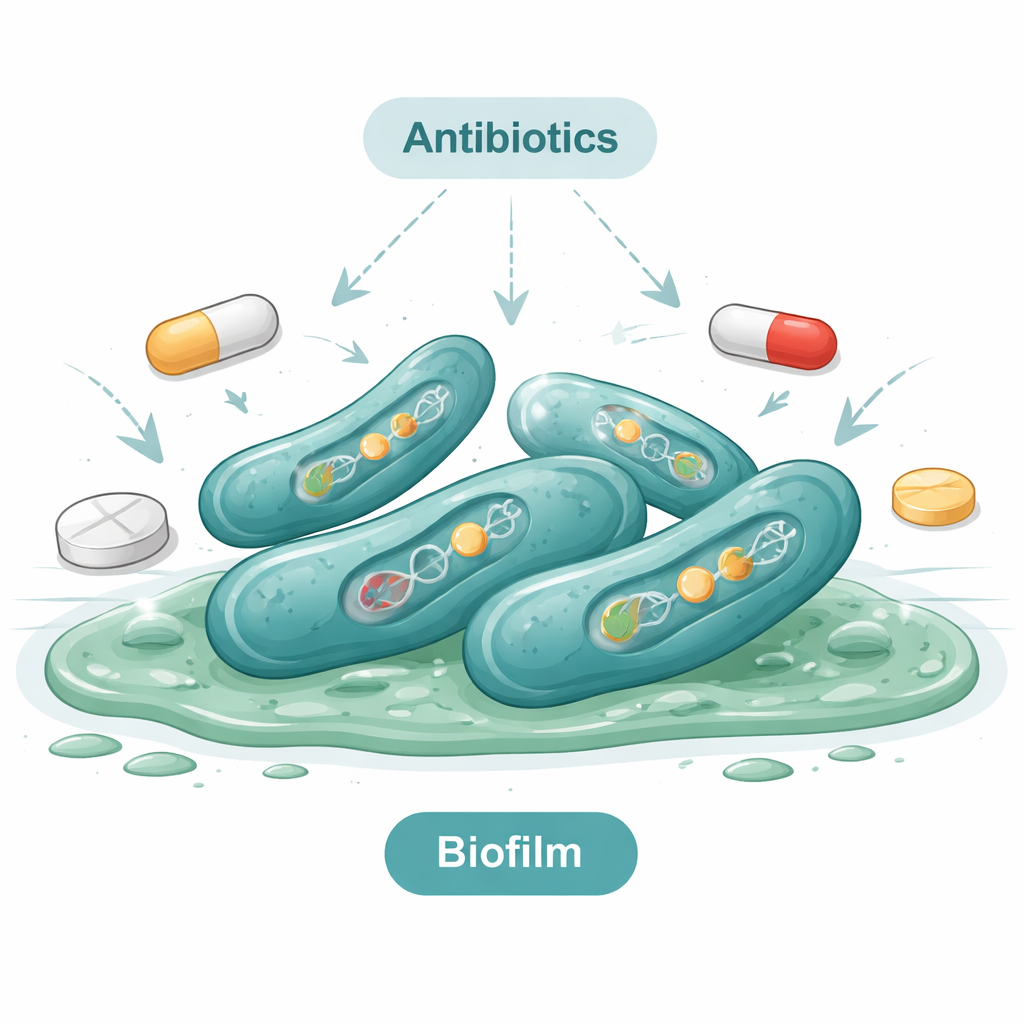
Antibiotics That Fail—and a Few That Still Work
The study then asked a pressing question: which antibiotics still work against these strains? When the researchers exposed 50 isolates to 16 different drugs, they discovered that many common antibiotics were almost useless. All isolates were resistant to amoxicillin, and nearly all resisted erythromycin and several other older drugs. Most were also unfazed by colistin, a drug often reserved as a last resort. On the brighter side, a handful of antibiotics—including imipenem and some related drugs—still worked against many isolates. Even so, more than four out of five strains were resistant to multiple drug types at once, a situation known as multidrug resistance. Genetic testing backed this up: the bacteria carried resistance genes that help them inactivate antibiotics or pump them out of their cells.
Linking Food, Water, and Human Infections
To explore how related the strains were across sources, the team used a DNA “fingerprinting” method called ERIC‑PCR on a subset of clinical isolates. This technique groups bacteria based on repeated patterns in their genomes. The analysis split the strains into clusters that contained samples from both people and foods such as fish, milk, chicken, and minced meat. This pattern suggests that food and water can act as stepping stones, allowing similar strains of P. aeruginosa to move between the environment and human patients. In practical terms, the bacterium detected in a piece of raw fish in the market may be closely related to the one causing a wound infection in a nearby hospital.
What This Means for Everyday Health
For a layperson, the message is straightforward: a common environmental germ is finding its way from water and animal‑based foods into people, and many of its strains are armed with genes that make them tough, stubborn, and harmful. Because so many isolates are resistant to several antibiotics, treating infections becomes more complicated, expensive, and risky. The authors argue that better hygiene in food production and handling, safer water systems, and more careful use of antibiotics in both people and animals are essential to slow the spread of these hardy strains. They also call for ongoing surveillance and more detailed genetic studies to map how P. aeruginosa travels through the food chain and into hospitals, so that future outbreaks can be better predicted and prevented.
Citation: Mousa, W.S., Abdeen, E.E., El-Gendy, H.F. et al. Prevalence and genotyping of Pseudomonas aeruginosa from food and human sources. Sci Rep 16, 7179 (2026). https://doi.org/10.1038/s41598-026-37559-y
Keywords: Pseudomonas aeruginosa, foodborne bacteria, antibiotic resistance, biofilm, water contamination