Clear Sky Science · en
Computational identification of Terminalia arjuna phytochemicals as potential 3α-HSD3 inhibitors
Why this research matters
Breast cancer is the most common cancer in women, and many tumors grow in response to hormones. This study asks a timely question: can natural substances from a traditional medicinal tree, Terminalia arjuna, be turned into modern drug candidates that gently "turn down" a key hormone-processing enzyme linked to tumor growth? Using advanced computer simulations instead of animal testing, the researchers search for plant compounds that might one day complement or improve current hormone therapies.
Hormones, a single enzyme, and tumor growth
In certain breast cancers, especially the widely studied MCF-7 type, growth is strongly driven by the female hormone estrogen. Normally, androgens (often thought of as male hormones) can counterbalance estrogen’s effects. The enzyme at the center of this work, called 3α-HSD3, helps convert a powerful androgen into weaker forms. When 3α-HSD3 is overactive, it strips away that protective androgen influence and leaves an estrogen-dominated environment that encourages tumor cells to multiply and resist treatment. Blocking this enzyme could therefore shift the hormone balance back toward restraining cancer growth.
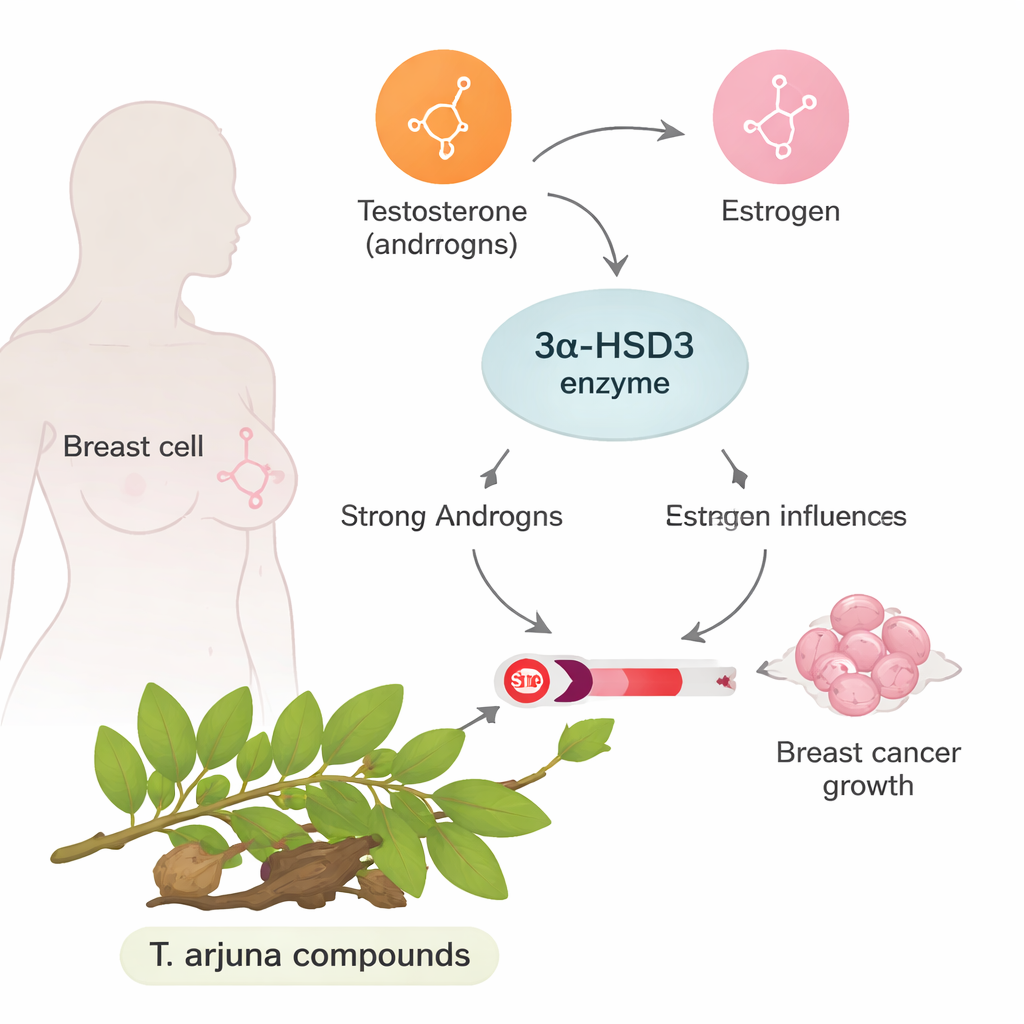
The promise of a traditional healing tree
Terminalia arjuna is a tree long used in South Asian traditional medicine, especially for heart health. Its bark and other parts contain many plant chemicals with reported antioxidant and anticancer properties. However, none had been carefully examined against 3α-HSD3. The authors selected nine well-known molecules from this tree, including flavonoids and tannins, based on earlier reports of anticancer and protective effects. Instead of immediately moving into lab animals or patients, they first used a suite of computer-aided drug design tools to ask: which of these natural molecules are most likely to fit and block the enzyme, behave like real drugs in the body, and avoid obvious toxicity?
Simulating how plant molecules meet their target
The team began with the three-dimensional structure of human 3α-HSD3, previously solved by X-ray crystallography, and checked its quality using standard structural tests. They then built and digitally "relaxed" the shapes of each plant molecule to reflect realistic forms they would take in the body. Next came virtual docking: the computer tried many ways of placing each molecule into the enzyme’s active pocket and scored how snugly and favorably it bound. Several Terminalia compounds showed strong predicted binding, often better than a known synthetic reference inhibitor. Four stood out in particular: luteolin, leucocyanidin, gallic acid, and ellagic acid. These were not only predicted to bind tightly, but also to form multiple hydrogen bonds and other stabilizing contacts with key amino acids that control the enzyme’s activity.
Testing stability, behavior, and safety in silico
Docking offers a snapshot; the researchers then asked if these complexes would remain stable over time in a watery, body-like environment. They ran 100-nanosecond molecular dynamics simulations—essentially, physics-based movies of atoms jostling at body temperature. Measures of how much the protein and ligands wobbled showed that the four plant molecules stayed firmly lodged in the enzyme’s pocket, often stabilizing it more than the control drug. Additional calculations estimated binding energies and confirmed that luteolin, leucocyanidin, and gallic acid in particular formed strong, favorable interactions. Parallel computer models predicted that all four candidates should be reasonably absorbable by the gut, distribute well in tissues, avoid major liver enzyme problems, and show low likelihood of genetic or organ toxicity at therapeutic doses.
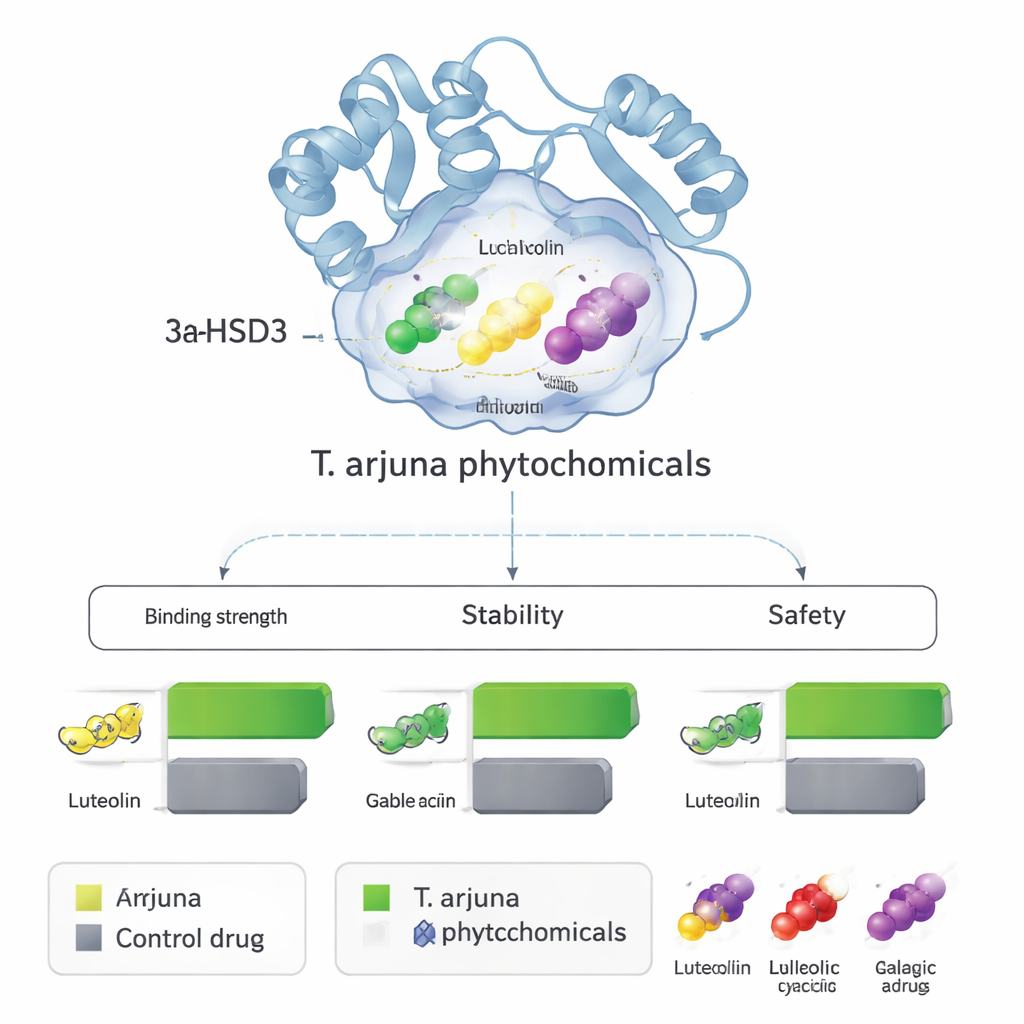
From computer predictions to future therapies
To a non-specialist, the key message is that this work does not claim to have discovered a new cure, but it does provide a focused shortlist of promising natural molecules. Luteolin, leucocyanidin, gallic acid, and ellagic acid from Terminalia arjuna appear, by multiple independent computer tests, to latch onto and potentially block 3α-HSD3, an enzyme that helps create an estrogen-rich environment in hormone-sensitive breast cancer cells. They also look drug-like and relatively safe on paper. The next essential steps will be laboratory experiments to see whether these predictions hold true: do these compounds really slow the enzyme in test tubes, and do they actually curb growth of breast cancer cells without harming healthy tissue? If so, they could guide the design of improved derivatives or inspire new combination therapies that harness both traditional plant wisdom and modern molecular science.
Citation: Al Mashud, M.A., Rahman, M.A., Kumer, A. et al. Computational identification of Terminalia arjuna phytochemicals as potential 3α-HSD3 inhibitors. Sci Rep 16, 8045 (2026). https://doi.org/10.1038/s41598-026-37558-z
Keywords: breast cancer hormones, Terminalia arjuna, natural enzyme inhibitors, computer-aided drug design, luteolin gallic acid