Clear Sky Science · en
A split biotin ligase approach reveals proteins associated with oligomeric alpha-synuclein during aggregation
Why clumped brain proteins matter
In Parkinson’s disease and related conditions, a normally helpful brain protein called alpha‑synuclein can clump into sticky deposits known as Lewy bodies. These deposits appear decades before symptoms and are a defining feature seen at autopsy, yet we still do not know which other proteins help push alpha‑synuclein from a harmless state into dangerous aggregates. This study uses a clever molecular "proximity tag" to map which proteins sit close to alpha‑synuclein as it shifts from single molecules to small clumps, offering new clues to how the disease begins and which pathways might be targeted to slow or prevent it.
A molecular “tag” that lights up neighbors
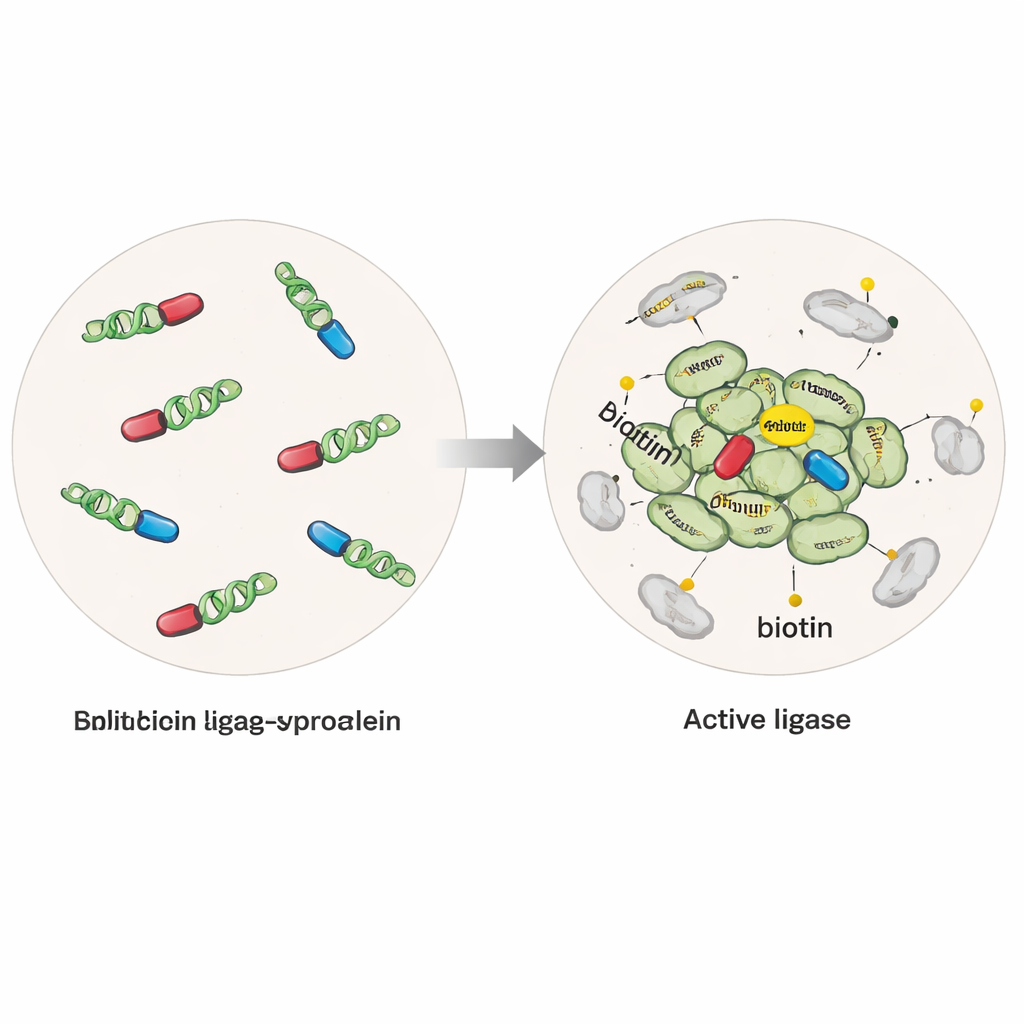
The researchers engineered human cells to make alpha‑synuclein fused to a modified enzyme that can attach a small biotin tag onto any protein within a few billionths of a meter. In one version, the enzyme was intact and active whenever alpha‑synuclein was present, capturing proteins near single molecules as well as clumps. In a second, more selective "split" version, the enzyme was cut into two inactive halves, each attached to a separate alpha‑synuclein molecule. Only when alpha‑synuclein molecules came together into multimers or oligomers did the halves reunite into an active enzyme, turning on tagging specifically around these early aggregates. By adding purified alpha‑synuclein fibrils to the cells, the team could also drive disease‑like aggregation and compare normal versus pathological states.
Building a map of nearby proteins
After allowing time for tagging, the team pulled down all the biotin‑labeled proteins and identified them using mass spectrometry, a technique that weighs and counts proteins with high precision. Across all conditions they found 1,277 distinct proteins, and after careful filtering they narrowed this to 581 that were likely to be genuinely near alpha‑synuclein. Comparing the intact and split enzyme systems, they could distinguish proteins that associate mainly with single molecules from those that prefer multimers. They also compared cells with and without added fibrils to see how the local protein environment of alpha‑synuclein changes once aggregation is triggered.
Key cellular pathways drawn into the clumps
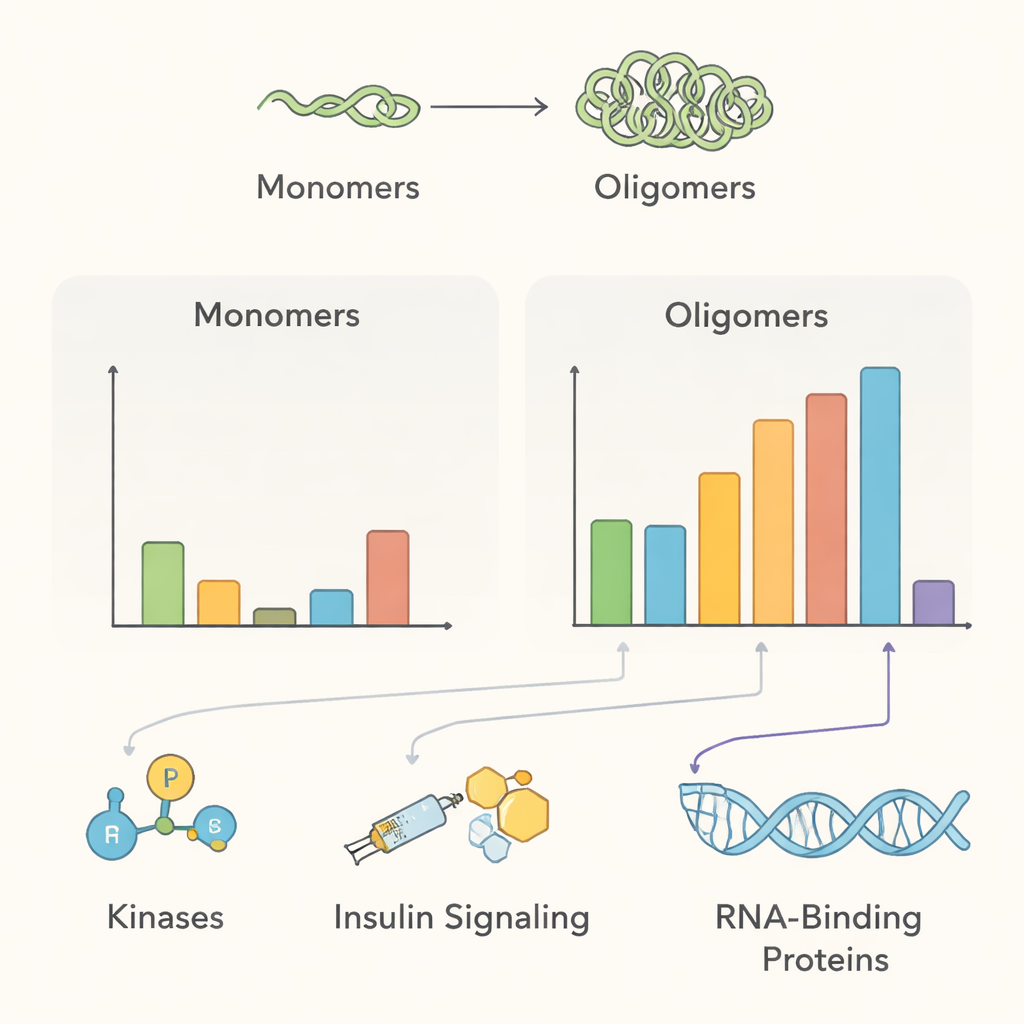
The resulting interactome sketch points to several major biological themes. Under normal conditions, multimeric alpha‑synuclein sat close to many membrane‑trafficking proteins and protein‑modifying enzymes—consistent with its proposed role in helping nerve cells manage synaptic vesicles and maintain protein quality control. Several enzymes that add phosphate groups, including kinases PAK4, RIPK2, and MAP4K4, were enriched near healthy multimers but appeared to lose contact once aggregates formed, hinting that disruption of these relationships might favor disease‑linked forms of the protein. The team also detected proteins tied to insulin‑like growth factor signaling, supporting clinical links between insulin resistance and Parkinson’s disease, and found that aggregation recruits proteins involved in disposing of damaged proteins, such as the 14‑3‑3 chaperone family and specific components of cellular degradation pathways.
Unexpected bridges to other brain disorders
Beyond Parkinson’s disease, the data highlight a surprising connection to amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD). An RNA‑binding protein called FUS, which mislocalizes and aggregates in those conditions, appeared near multimeric alpha‑synuclein under both normal and aggregation‑driven conditions. Other RNA‑related proteins also shifted their proximity as alpha‑synuclein aggregated. These findings raise the possibility that small alpha‑synuclein assemblies may trap RNA‑handling proteins in the wrong place within cells, potentially linking the biology of Lewy body diseases with that of ALS and FTD, although direct physical interactions still need to be verified.
What this means for understanding disease
For non‑specialists, the main takeaway is that this work provides an early‑stage "contact list" of proteins that travel with alpha‑synuclein as it moves from a normal helper to a harmful clump. The study suggests that losing certain protective relationships—such as with specific kinases or quality‑control enzymes—may be just as important as gaining new toxic partners. While the experiments were done in a simplified cell model and do not yet prove cause and effect, they highlight concrete candidates and pathways for future testing in neurons and brain tissue. In the long run, mapping these shifting protein neighborhoods could guide strategies to restore protective interactions, support insulin‑related signaling, or prevent the capture of RNA‑binding proteins, potentially slowing the march from subtle molecular changes to full‑blown Lewy body diseases.
Citation: Fernandes, A.R., Owen, A.P., Faroqi, A.H. et al. A split biotin ligase approach reveals proteins associated with oligomeric alpha-synuclein during aggregation. Sci Rep 16, 7270 (2026). https://doi.org/10.1038/s41598-026-37551-6
Keywords: alpha-synuclein, Lewy body disease, protein aggregation, proximity proteomics, Parkinson's disease