Clear Sky Science · en
New insights into the molecular basis of gametogenesis in the hybridogenetic water frog Pelophylax esculentus
Why strange frogs matter
Most animals, including humans, reshuffle their genes every generation when sperm and egg meet. But a few odd creatures bend these rules and pass on entire genomes as ready-made clones. European water frogs belong to this select group. They are living experiments in how sex, asexuality, and hybridization can intertwine. Understanding how these frogs make their eggs and sperm does not just satisfy curiosity about a quirky amphibian—it also sheds light on how genomes are controlled, repaired, and sometimes ruthlessly edited in all vertebrates.
Frogs that break the rules of heredity
In many European ponds, three closely related water frogs coexist. Two are ordinary sexual species, called here simply L and R. When they mate, they can produce a hybrid, LR, known as Pelophylax esculentus. This hybrid does something remarkable: in its germline—the cells that will become eggs or sperm—it throws out one parental genome and copies the other without shuffling it. The result is a clonal gamete carrying either an intact L or an intact R genome, which is then fertilized by a normal gamete from one of the parental species. Depending on the local “population system,” hybrids may routinely exclude the R genome, the L genome, or, in some places, both in different individuals. The study asks: which genes help orchestrate this targeted genome exclusion and clonal inheritance?
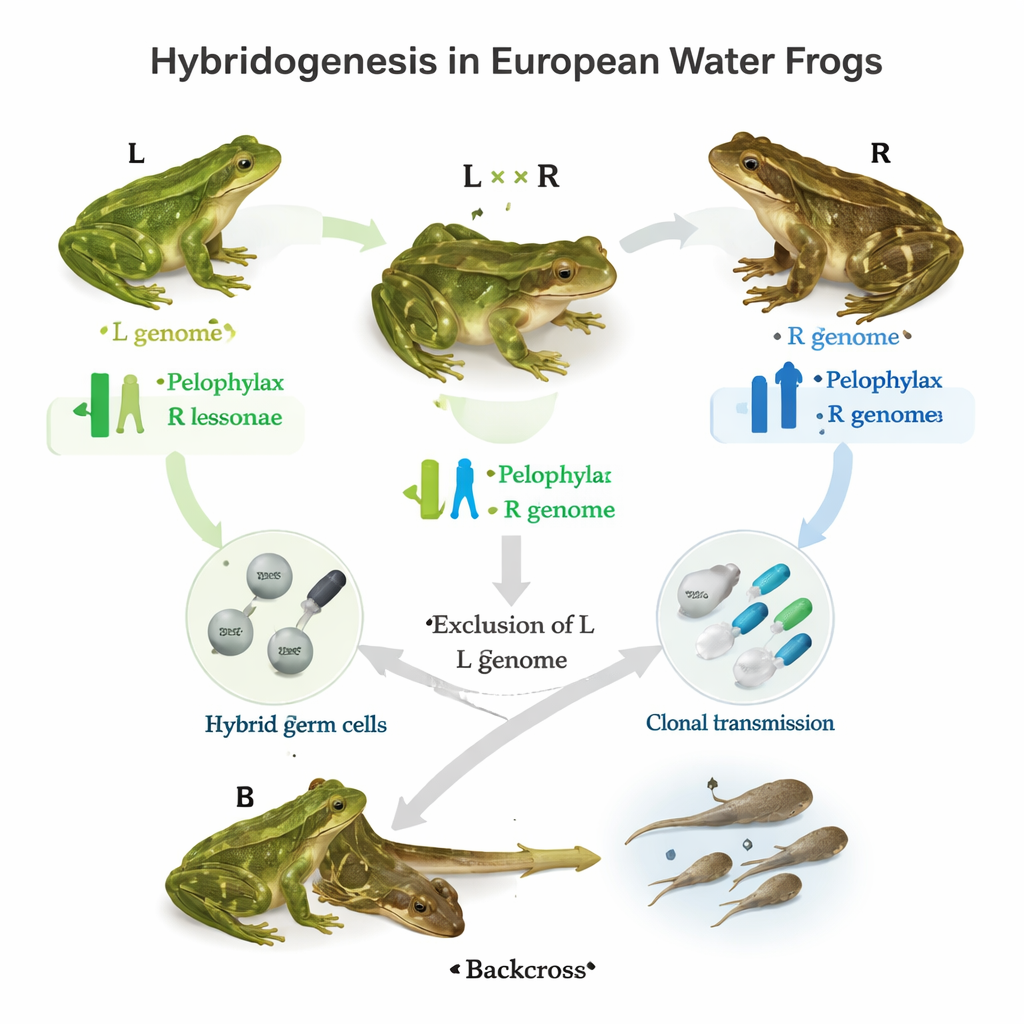
Hunting for the genes behind clonal sex
The authors first built a catalog of 160 genes known from other animals to take part in gametogenesis—the process of making eggs and sperm. They sequenced these genes from testes of L and R frogs and compared their basic properties, such as length, structure, and DNA base composition. Most of these genes turned out to be highly conserved between the two species, suggesting they perform important roles that tolerate little change. Next, the team focused on 52 of these genes and scanned them for single-letter DNA differences, known as single nucleotide polymorphisms (SNPs), in more than 650 frogs collected across Europe. These individuals came from different population systems: those where only L and hybrids coexist, those where only R and hybrids coexist, all-hybrid systems rich in triploid frogs, and purely R populations.
Signals of gene flow and hidden structure
By examining patterns of SNP variation, the researchers uncovered evidence of introgression—gene flow—between the L and R gene pools. In several genes, R versions were nearly indistinguishable from their L counterparts, best explained by past hybridization and backcrossing. Although the overall rates of gene flow were low, they were higher into R than into L, matching earlier genetic work. When the team compared SNP frequencies among population systems, they found that certain genetic variants clustered strongly with particular systems. For example, LR hybrids from all-hybrid populations were genetically more similar to LR frogs from R–hybrid systems than to LR frogs from purely diploid L–hybrid systems, revealing hidden structure linked to how reproduction is organized locally.
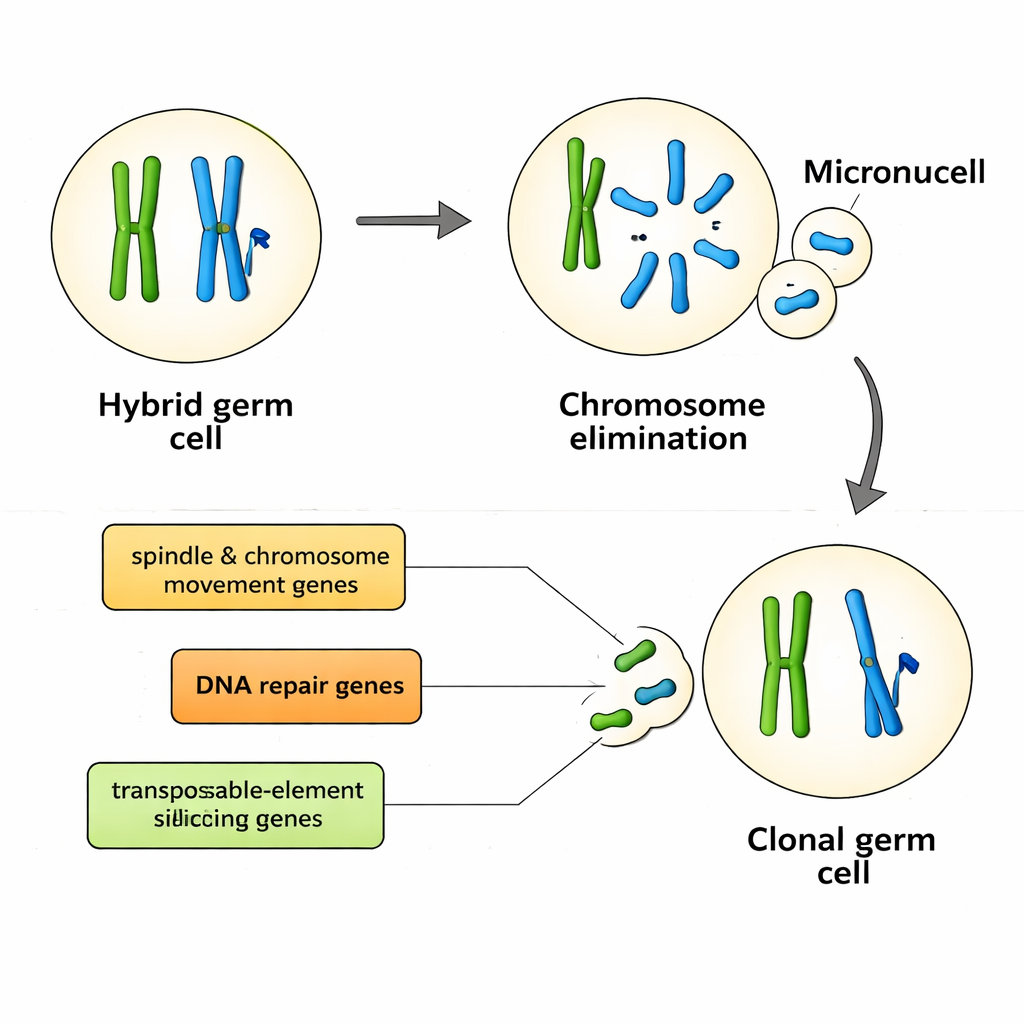
Key players in chromosome control and genome defense
Ten genes, in particular, showed strong associations with population system and thus with the way genomes are excluded and clonally transmitted. They fall into several functional groups. Some, such as kif22 and nusap1, help build and control the spindle, the cellular machine that moves chromosomes during cell division. Others, including hormad1, rad50, rad51ap1, and sfr1, are central to making and repairing DNA double-strand breaks—controlled cuts that normally promote recombination but could also trigger targeted elimination of chromosomes. Another gene, henmt1, stabilizes small RNAs that silence transposable elements, the mobile DNA pieces that can damage genomes or reshape centromeres, the anchor points needed for proper chromosome segregation. Together, these genes sketch a picture in which genome exclusion arises from the interplay of chromosome transport, DNA repair, and genome defense pathways, not from a single master switch.
Evolution at the edge of stability
The study suggests that rare recombination and two-way gene flow between L and R have helped generate distinct genome variants that favor different non-Mendelian inheritance modes. Some hybrid lineages may accumulate harmful mutations as their clonal genomes ratchet forward, while sporadic recombination and introgression occasionally refresh them, at the cost of developmental problems in some offspring. Rather than fitting a simple rule based on how divergent two species are, the Pelophylax system seems governed by a complex network of interacting factors that fine-tune whether a hybrid becomes a stable clonal lineage or a dead end.
What this means beyond frogs
For a general reader, the take-home message is that inheritance can be far more flexible than textbook Mendelian diagrams suggest. These water frogs reveal that vertebrate genomes can be selectively deleted, copied, and reused like modular building blocks, depending on context. By pinpointing real genes tied to this process, the study turns an odd reproductive trick into a tractable molecular problem. Future experiments, guided by this new gene catalog, will test how tweaks to chromosome-handling and genome-defense pathways can flip a population between ordinary sex and clonal hybrid reproduction—insight that may ultimately illuminate fertility, genome stability, and the evolution of sex in many other animals.
Citation: Plötner, M., Meixner, M., Poustka, A.J. et al. New insights into the molecular basis of gametogenesis in the hybridogenetic water frog Pelophylax esculentus. Sci Rep 16, 5012 (2026). https://doi.org/10.1038/s41598-026-37515-w
Keywords: hybridogenesis, water frogs, clonal inheritance, genome exclusion, gametogenesis