Clear Sky Science · en
Molecular docking and dynamic simulation of marine natural products from soft coral-derived microbes against SARS-CoV-2 main protease and spike protein
Hidden Help from Ocean Life
Long after vaccines arrived, new versions of the virus that causes COVID-19 kept emerging, challenging treatments and prolonging the pandemic. This study asks a surprising question with real-world importance: could chemical compounds made by tiny microbes living in soft corals help block the coronavirus, including major variants like Delta and Omicron? Using computer-based screening rather than lab animals or patients, the researchers explored whether these marine molecules might latch onto critical viral parts and slow the infection process.
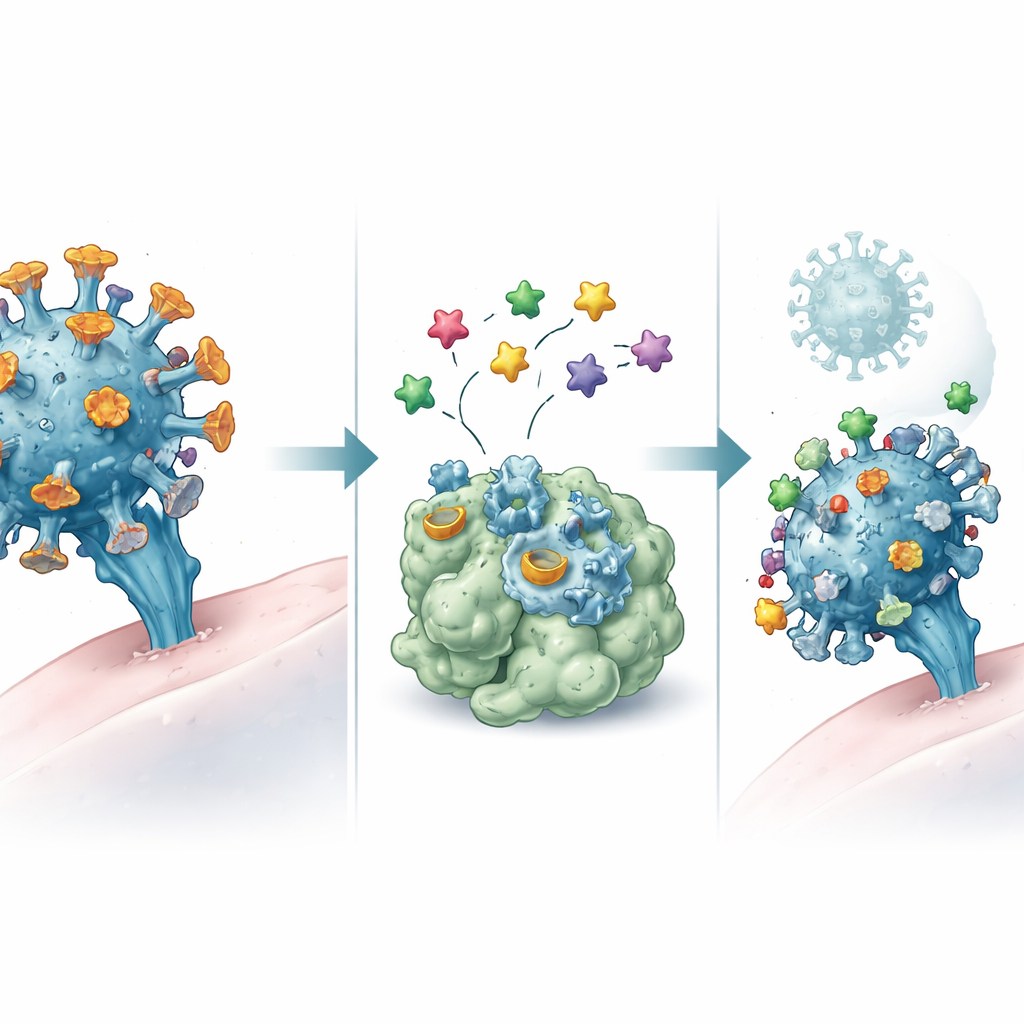
How the Virus Breaks In
The virus behind COVID-19 relies on two main tools to invade our cells and multiply. First is the spike protein on the virus surface, which acts like a key that fits into a lock on human cells, beginning with a region called the receptor-binding domain. Second is the main protease, an internal cutting tool the virus uses to process its proteins and build new virus particles. Variants of concern—Alpha, Beta, Gamma, Delta, and Omicron—carry small changes in the spike region that can make the virus spread more easily or dodge parts of our immune response, while the protease remains more stable. Because existing antiviral drugs do not always work well against these variants, both the spike and the protease are prime targets for new treatments.
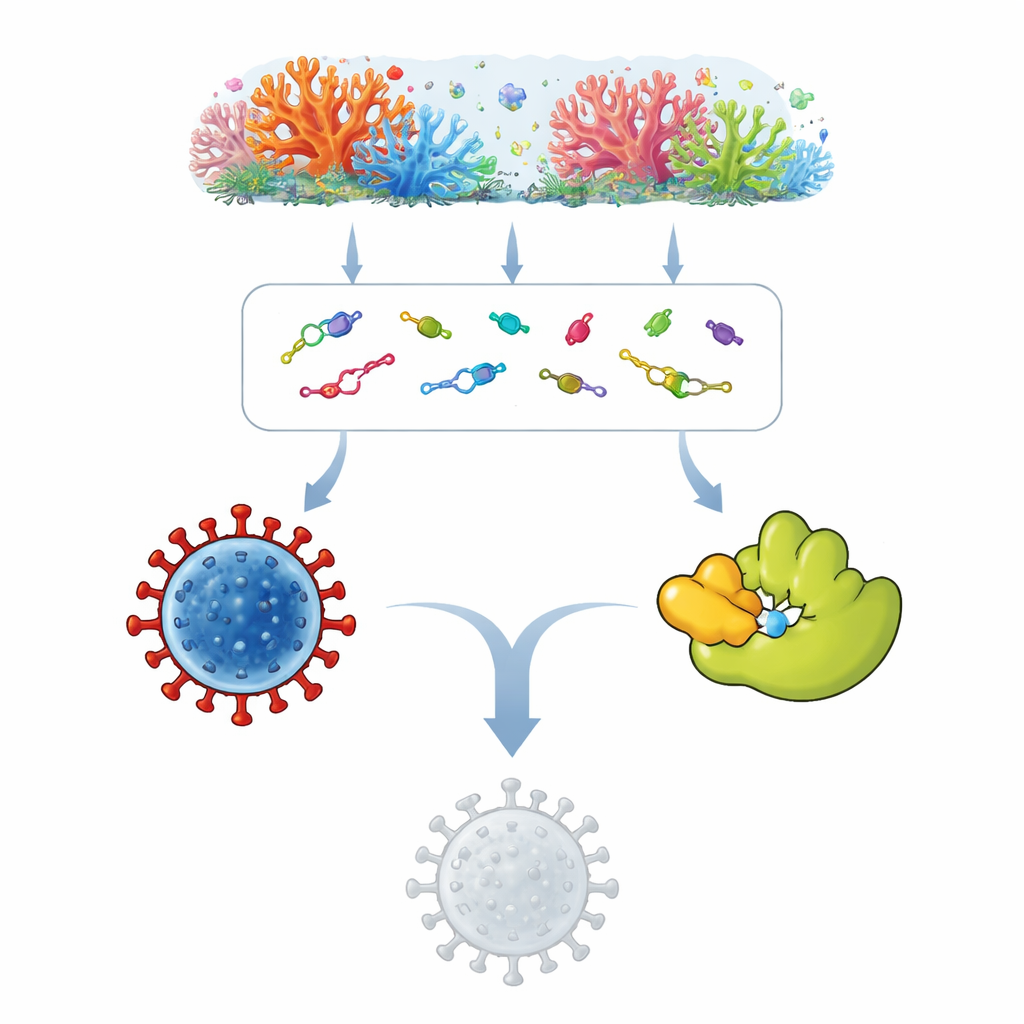
Treasure Hunt in Coral Reefs
Soft corals form vibrant underwater habitats and host a rich community of microbes such as fungi and bacteria. These tiny partners produce a wide array of natural substances as part of their own survival strategies, some of which have already led to anticancer or antimicrobial drugs. The team collected information on 119 such marine natural products and built three-dimensional models of their shapes. They then used molecular docking, a kind of virtual fitting exercise, to see which compounds could snugly attach to the spike protein and the main protease with stronger predicted attraction than known antiviral drugs like remdesivir or nelfinavir.
Virtual Matchmaking with the Virus
The computer docking runs highlighted several standout molecules, including Cottoquinazoline B and D, Tetraorcinol A, Versicoloritide A and C, Fumiquinazoline K, and Pencillanthranin A. These compounds were predicted to bind both the main protease and the spike region of the original virus and multiple variants more strongly than the control drugs. Many formed multiple stabilizing contacts, such as hydrogen bonds and hydrophobic interactions, at key spots on the viral proteins involved in cell entry or replication. To go beyond static snapshots, the researchers ran long molecular dynamics simulations, which mimic how these protein–compound pairs move in watery surroundings over time. Several top candidates, especially Cottoquinazoline B, Tetraorcinol A, and Versicoloritide A, remained tightly associated with their viral targets over hundreds of nanoseconds, suggesting stable attachment rather than fleeting encounters.
Early Safety Checks on the Digital Drawing Board
The study also examined basic drug-like features using established prediction tools. These tests estimate whether a compound is likely to be absorbed, distributed, broken down, and cleared in a way compatible with future medicines, and whether it might be toxic. Many of the promising coral-derived molecules met common rules of thumb for oral drugs and were flagged as non-carcinogenic, although a few raised concerns about potential toxicity or chemical behaviors that can interfere with lab tests. Overall, the most attractive candidates combined strong predicted binding to viral proteins with acceptable virtual safety profiles, marking them as especially interesting for follow-up work.
What This Could Mean for Future Treatments
This research does not claim to have discovered a ready-to-use COVID-19 drug. Rather, it provides a carefully filtered list of marine compounds that look promising on the computer screen: they appear able to stick to the virus’s spike and main protease, including in major variants, and many pass early checks for drug-likeness. The next steps would require real-world lab experiments and animal studies to see whether these molecules truly block infection and are safe in living systems. Still, the work underscores how overlooked ecosystems like coral reefs may hold valuable chemical tools against fast-changing viruses, and how computational methods can rapidly sift through nature’s library to guide smarter, faster drug discovery.
Citation: Anthikapalli, N.V.A., Patil, V.S., Alugoju, P. et al. Molecular docking and dynamic simulation of marine natural products from soft coral-derived microbes against SARS-CoV-2 main protease and spike protein. Sci Rep 16, 8252 (2026). https://doi.org/10.1038/s41598-026-37446-6
Keywords: marine natural products, coral reef microbes, SARS-CoV-2 spike, main protease inhibitors, COVID-19 drug discovery