Clear Sky Science · en
Comparative karyotype analysis of eleven species of Lilium from China using FISH with rDNA oligo-probes
Why lily chromosomes matter beyond the lab
Garden lilies are more than beautiful flowers and traditional remedies: they are also genetic models that help scientists understand how new plant species arise and how to classify them correctly. This study looks inside the cells of eleven Chinese lily species, many used as food or medicine, to see how their chromosomes are arranged and marked. By comparing these hidden structures, the authors clarify which lilies are truly close relatives and provide evidence to settle long-standing debates about lily family trees.
Untangling a confusing lily family tree
The genus Lilium contains about 125 species spread across the Northern Hemisphere, with China as one of its main centers of diversity. For more than a century, botanists have tried to group these species into natural sections based largely on visible traits such as flower shape. But look-alike flowers do not always mean close kinship, and genetic studies of chloroplasts and nuclei have sometimes given conflicting answers about how lilies are related. In particular, one group called Leucolirion and its two subsections (6a and 6b), plus several species from southwest and central China, have been difficult to place with confidence.
Reading barcodes on chromosomes
To tackle this problem, the authors turned to cytogenetics, the study of chromosomes. All eleven species they examined share the same basic set of twelve chromosomes, usually present as 24 in diploids; two populations of Lilium lancifolium were triploid with 36 chromosomes. Under the microscope, each species shows two larger chromosomes and ten smaller ones, but subtle differences in shape and length can be hard to see by eye. The team therefore measured chromosome sizes precisely and used a technique called fluorescence in situ hybridization (FISH) to attach glowing DNA tags to specific ribosomal DNA (rDNA) regions, which act like bright barcodes along each chromosome.
Finding patterns in lily genomes
The researchers mapped two types of rDNA—5S and 35S—on the chromosomes of each species. Most lilies had one 5S site and between two and six 35S sites, often at characteristic positions. For example, four species traditionally placed in subsection Leucolirion 6a (L. leucanthum, L. sargentiae, L. sulphureum, and L. regale) shared very similar patterns of rDNA marks on the same chromosome pairs, pointing to a close relationship, even though L. regale showed some extra changes. In contrast, species of the sister subsection 6b and some Sinomartagon lilies displayed distinctly different rDNA arrangements, underlining a real split between these two groups. Outside Leucolirion, strongly shared patterns linked the widely cultivated L. lancifolium to L. pumilum and L. davidii, matching earlier DNA-sequence studies that had grouped them together.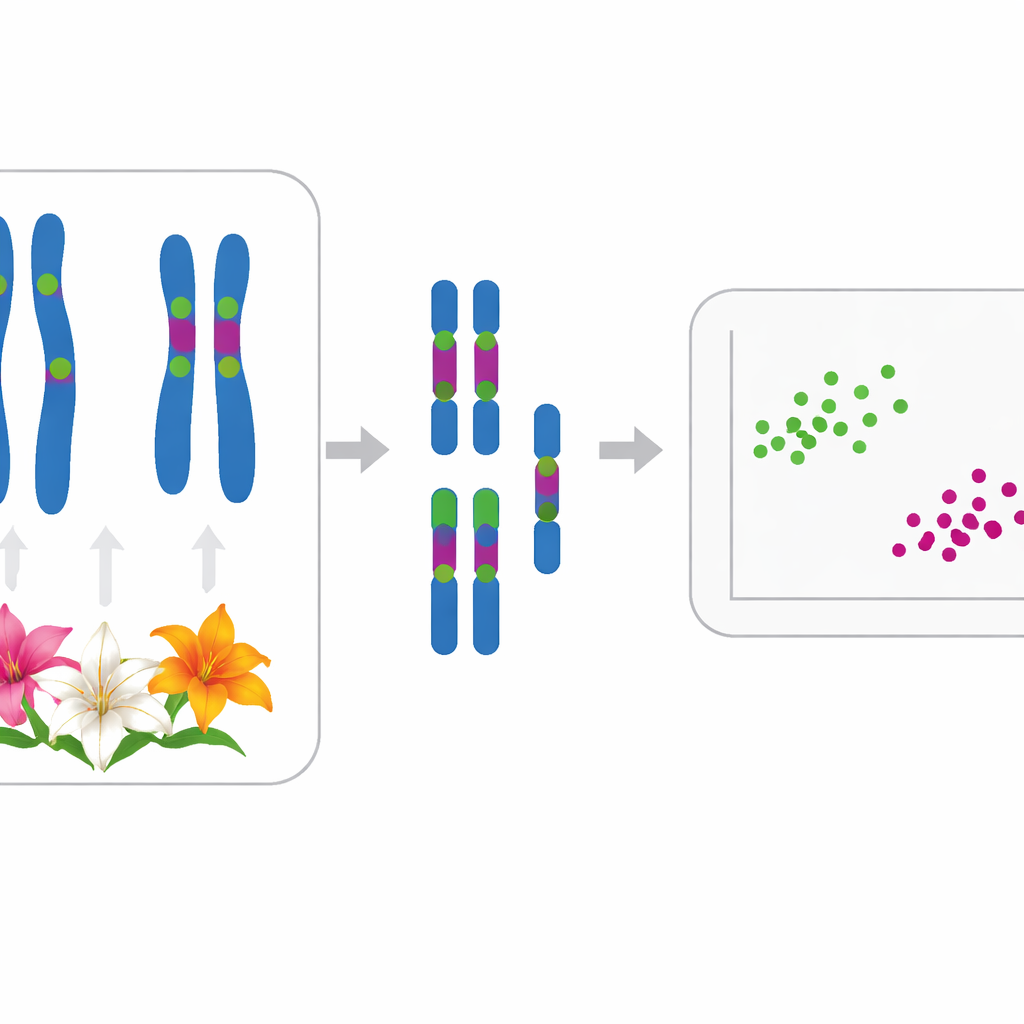
Turning measurements into a map of relationships
Beyond the barcode positions, the team quantified how uneven each karyotype was—both in the positions of centromeres (the “waist” of the chromosome) and in chromosome lengths. They used two statistical indices, MCA and CVCL, to capture these features and plotted the eleven species in a two-dimensional graph. Each species occupied its own distinct spot, showing that even apparently similar karyotypes can be told apart when measured carefully. A broader statistical analysis (principal coordinate analysis) then grouped the lilies into two main clusters that correspond well with their inferred evolutionary history. One cluster contained L. pumilum, L. jinfushanense, L. brownii var. viridulum, L. regale, and the L. lancifolium and L. davidii forms; the other grouped L. leucanthum, L. sargentiae, L. sulphureum, L. henryi, and L. rosthornii.
What this means for naming and using lilies
By combining detailed chromosome measurements with rDNA barcode maps, this study shows that lily species differ more in their karyotypes than previously thought and that these differences carry a clear evolutionary signal. In practical terms, the work supports treating subsection Leucolirion 6a as a natural, well-defined group within the genus Lilium, in line with recent classification proposals. For breeders, herbal practitioners, and conservation planners, having a more accurate picture of who is related to whom among lilies can guide cross-breeding, safeguard genetic diversity, and ensure that medicinal and edible varieties are correctly identified and used.
Citation: Yang, YD., Jiang, XH., Zhang, BY. et al. Comparative karyotype analysis of eleven species of Lilium from China using FISH with rDNA oligo-probes. Sci Rep 16, 9446 (2026). https://doi.org/10.1038/s41598-026-37297-1
Keywords: Lilium, chromosomes, fluorescence in situ hybridization, plant evolution, cytotaxonomy