Clear Sky Science · en
Impact of the defined Oligo-MM12 microbiota on intestinal colonisation and dissemination of Listeria monocytogenes
Why the germs in your gut matter
Many people know Listeria monocytogenes as a dangerous foodborne germ linked to contaminated ready-to-eat foods and severe illness in vulnerable individuals. What is less obvious is that whether this bacterium gains a foothold in the body depends not only on our immune system, but also on the bustling community of microbes already living in our intestines. This study uses specially designed mouse models and a miniature “artificial colon” to ask a simple but important question: how does the complexity of the gut microbiome influence Listeria’s ability to colonize the gut and spread through the body?
A tale of two microbial neighborhoods
To tackle this question, the researchers compared two kinds of mice. One group, called SPF mice, carried a rich, natural mix of gut bacteria similar to that found in many laboratory animals. The other group, Oligo-MM12 mice, hosted a far simpler community made up of just 12 well-characterized bacterial strains. Because every member of this community is known and stable across generations, Oligo-MM12 mice are a powerful tool for dissecting which microbes help block invaders and which might leave gaps in our defences. By contrasting these mice, the team could test how a dense, diverse gut ecosystem differs from a streamlined, minimalist one in handling an infectious challenge.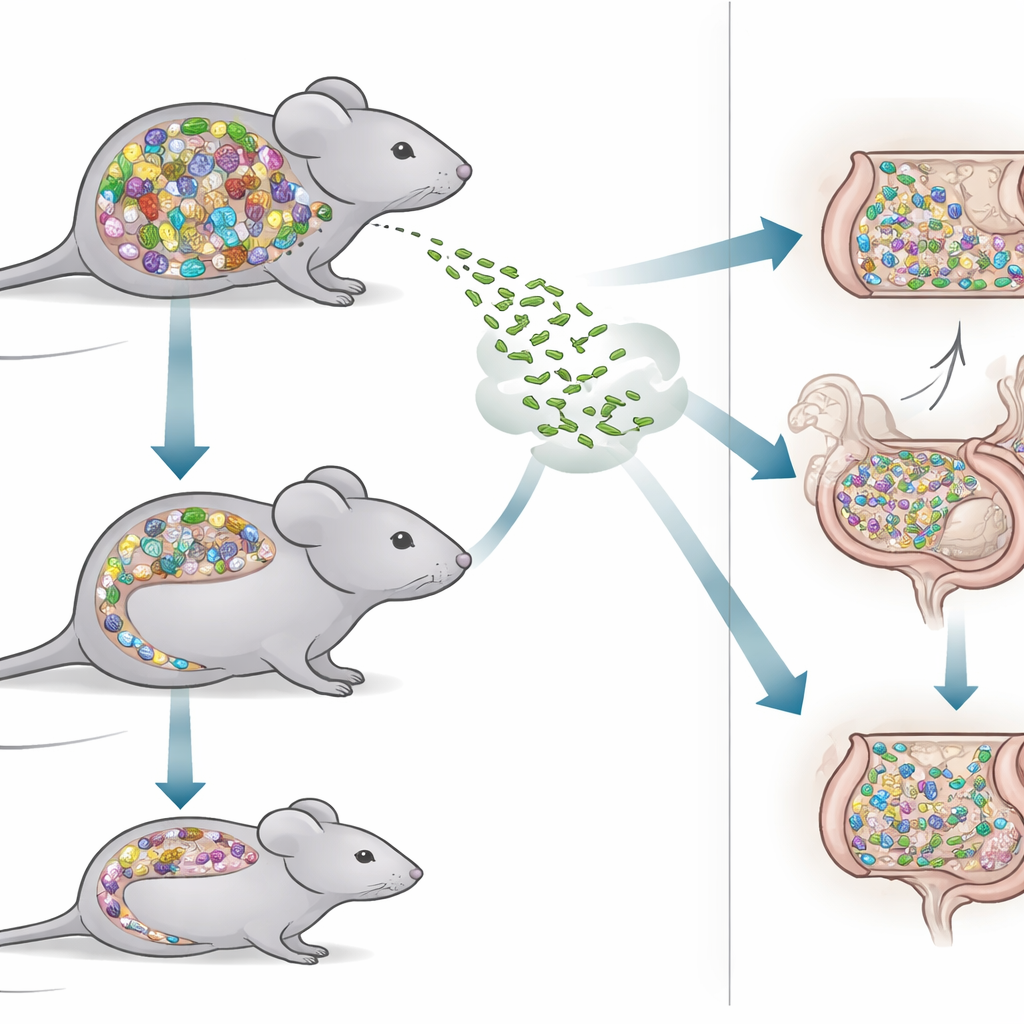
Following Listeria through the gut and beyond
The scientists infected both mouse types with the same oral dose of Listeria and tracked how many bacteria appeared in the feces and internal organs over three days. Interestingly, the simpler Oligo-MM12 community initially seemed better at holding Listeria in check: on the first day after infection, these mice shed fewer Listeria cells in their stool than SPF mice. However, that early advantage faded. By the third day, Oligo-MM12 mice were shedding more Listeria, while SPF mice had steadily pushed levels down. Despite these clear differences in the gut, the numbers of Listeria in internal organs such as the liver, spleen, and lymph nodes were similar between the two mouse models, suggesting that once Listeria reaches deeper tissues, microbiome complexity plays a smaller role.
Simulating the colon in a controlled box
To separate the effects of host tissues and immunity from the behaviour of the microbes themselves, the team also ran an ex vivo experiment using a device called the micro-Matrix bioreactor. They created faecal slurries from each mouse type, added them to a nutrient-rich medium in sealed, oxygen-free wells, and introduced Listeria to some of these miniature cultures while leaving others as controls. Over five days, they measured Listeria counts and sequenced bacterial DNA to follow how the communities evolved. In this artificial colon, Listeria grew to similar levels regardless of whether it faced the complex SPF microbiota or the simpler Oligo-MM12 community, and its presence caused only modest shifts in overall community composition. This contrast with the live animals highlights how strongly the immune system, gut lining, and bile and mucus flows shape infection in the real intestine.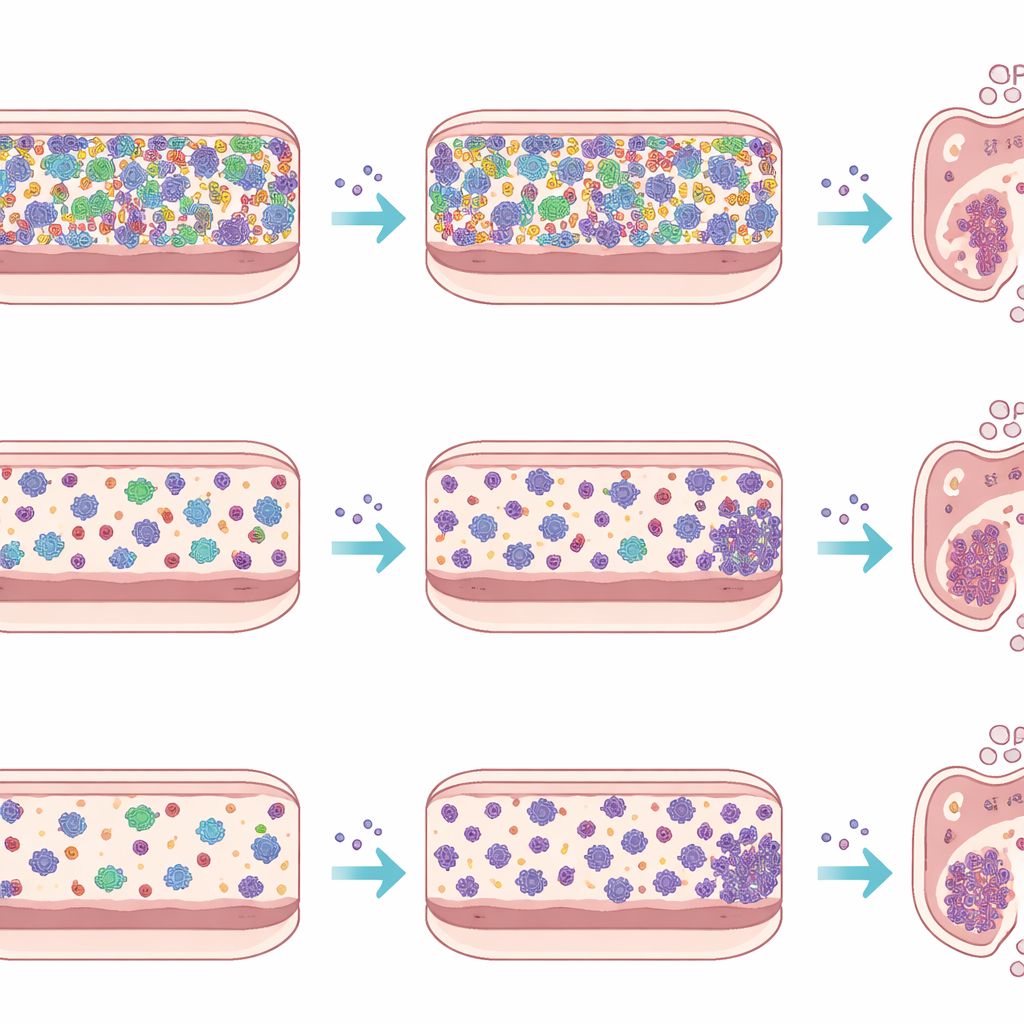
Key microbial winners, losers, and bystanders
By looking at which bacterial groups waxed or waned during infection, the study pinpoints candidates that may influence Listeria’s success. In Oligo-MM12 mice, levels of Akkermansia, a mucus-associated bacterium often linked to gut health, dipped on the second day then rebounded strongly by day three. In SPF mice, several genera known to produce beneficial metabolites or support barrier integrity, such as Blautia and Bacteroides, declined during infection, while certain Prevotella-like bacteria showed complex rises and falls that may reflect direct targeting by Listeria’s own antimicrobial compounds. Ex vivo, these patterns were largely muted, reinforcing the idea that many of the observed shifts arise from three-way interactions among pathogen, resident microbes, and the host environment rather than from simple microbe–microbe competition alone.
What this means for protecting people
For non-specialists, the central message is that not all healthy-looking microbiomes protect against infection in the same way, and that “more diversity” is not always better at every moment. The simplified Oligo-MM12 community provided strong early resistance to Listeria in the gut, but over time the more intricate SPF microbiome did a better job of clearing the invader. The work shows that carefully engineered, low-complexity microbiota models, combined with controllable ex vivo systems, can reveal which bacterial groups and environmental factors are crucial for blocking dangerous foodborne pathogens. Ultimately, such insights could guide targeted probiotics or microbiome-based therapies that bolster the right microbial allies at the right time to prevent serious Listeria infections in at-risk patients.
Citation: Cazzaniga, M., Bra, K.K., Herzog, M.K.M. et al. Impact of the defined Oligo-MM12 microbiota on intestinal colonisation and dissemination of Listeria monocytogenes. Sci Rep 16, 8434 (2026). https://doi.org/10.1038/s41598-026-37294-4
Keywords: Listeria infection, gut microbiome, colonization resistance, mouse model, probiotics