Clear Sky Science · en
Integrated analysis of global regulators and efflux genes associated with antimicrobial resistance reversal in multidrug resistant Klebsiella pneumoniae
Why these hospital germs are so hard to kill
In hospitals around the world, certain strains of the bacterium Klebsiella pneumoniae have become notoriously difficult to treat. They shrug off multiple antibiotics that once worked reliably, turning routine infections into life‑threatening emergencies. This study looks under the microbial hood to understand one important trick these bacteria use: microscopic pumps that spit antibiotics back out of the cell, and the genetic switches that seem to turn those pumps up or down.
Superbugs and their molecular escape hatches
Multidrug‑resistant Klebsiella pneumoniae is now encountered in many hospitals, including the Iraqi medical center where this research was carried out. These bacteria can survive several major antibiotic families at once, such as fluoroquinolones, aminoglycosides, and third‑generation cephalosporins. One way they do this is by installing powerful “efflux pumps” in their cell envelope. Among the most important is a system called AcrAB‑TolC, which spans the bacterial membranes and works like a tiny sump pump, grabbing antibiotic molecules inside the cell and pushing them back out before the drugs can do serious damage. When these pumps are highly active, the antibiotic concentration inside the bacterium never climbs high enough to be lethal.
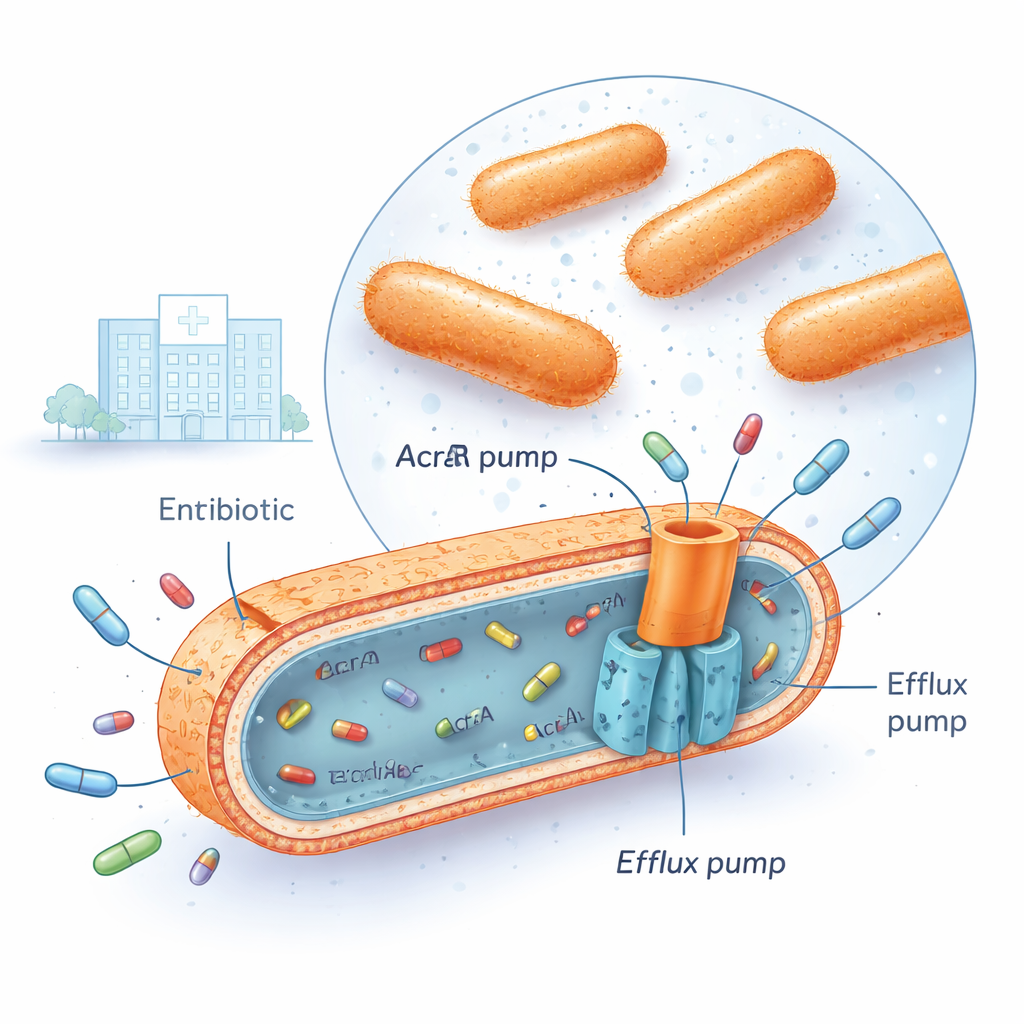
Gene switches that accompany pump overdrive
The researchers collected 30 multidrug‑resistant clinical isolates of Klebsiella pneumoniae and compared them with 10 antibiotic‑susceptible isolates and a standard reference strain. Using a technique called quantitative real‑time PCR, they measured how much certain genes were switched on. They focused on three "global regulators"—genes named marA, soxS, and rob—along with the three parts of the AcrAB‑TolC pump itself (acrA, acrB, and tolC). In the resistant strains, two regulators, marA and soxS, were typically turned up about four‑ to fivefold compared with susceptible bacteria, while acrB, a central pump component, was boosted nearly eightfold. Statistical analysis showed that when marA and soxS levels were high, acrB levels were usually high too, and those strains tended to withstand much higher doses of fluoroquinolone antibiotics like ciprofloxacin. A third regulator, rob, changed little and did not line up with pump activity, suggesting it plays a smaller role under the conditions tested.
Switching off the pumps to weaken resistance
Finding that pump and regulator genes are active together is suggestive but not enough to prove they truly drive resistance. To test whether the pumps were functionally important, the team turned to a chemical tool called PAβN. This compound is not used as a drug in patients, but in the lab it can plug AcrAB‑TolC‑like pumps and stop them from expelling antibiotics. The scientists chose ten resistant isolates that had especially high levels of the acrB gene and measured how much ciprofloxacin was needed to halt their growth, with and without PAβN. In eight of these ten strains, blocking the pump lowered the required ciprofloxacin dose at least fourfold, and sometimes as much as sixteenfold. This drop shows that, in many of these superbugs, active efflux pumps are indeed a major reason they can tolerate high antibiotic doses.
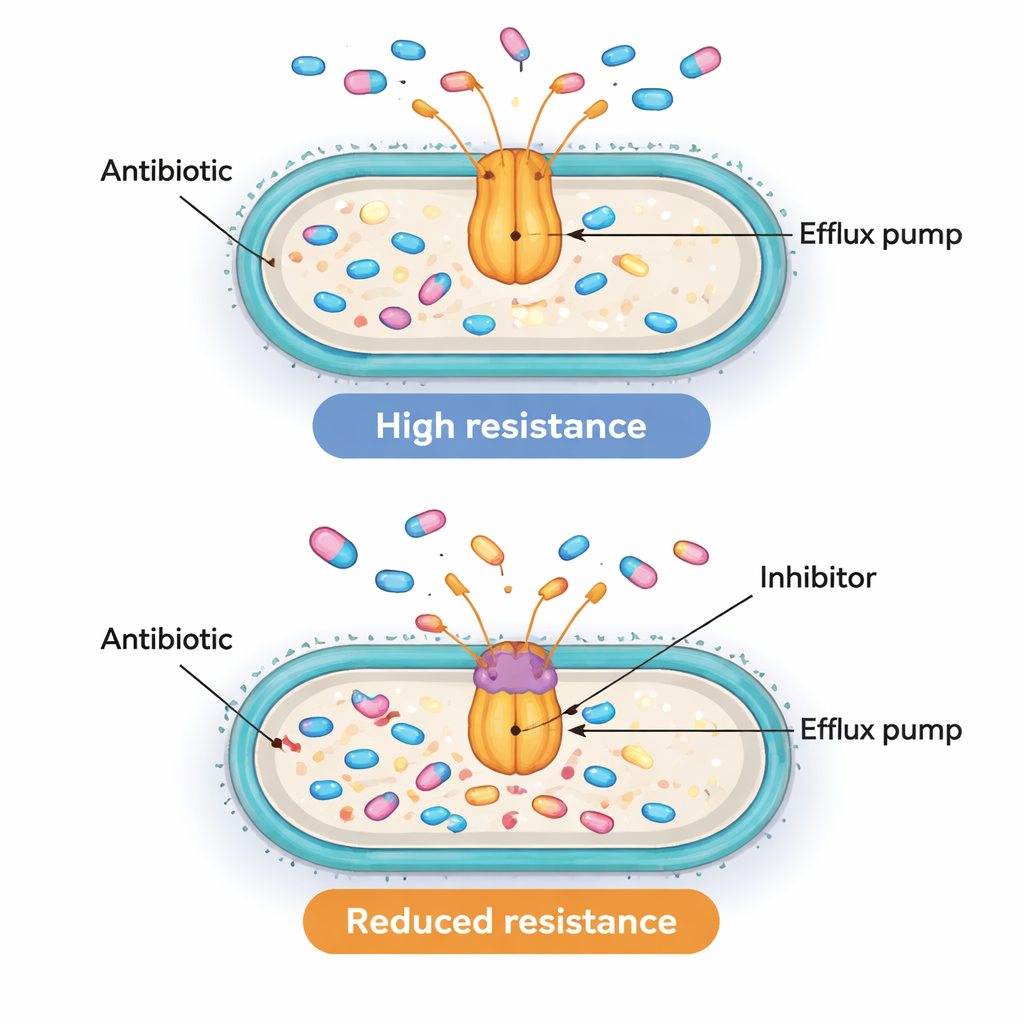
A complex resistance puzzle with promising targets
Not every strain behaved the same way. Some bacteria with strongly expressed pump genes only partly regained sensitivity when the pumps were blocked, implying that other resistance routes—such as changes to the antibiotic’s target inside the cell, production of drug‑breaking enzymes, or alternative pumps—also contribute. Gentamicin, an aminoglycoside antibiotic, for example, remained largely unaffected by the patterns seen in the marA/soxS and AcrAB‑TolC genes, underscoring that multidrug resistance is usually built from several overlapping defenses rather than a single mechanism.
What this means for future treatments
For non‑specialists, the key message is that some hospital bacteria can survive antibiotics not just by blocking or chopping up the drugs, but by actively throwing them out. This study links higher activity of two gene “switches,” MarA and SoxS, with more active efflux pumps and stronger resistance to a widely used class of antibiotics. When the pumps are experimentally plugged, many of these bacteria become at least partly sensitive again. While the work does not yet prove exactly how these switches control the pumps, it highlights both the regulators and the pumps as attractive targets. In the long run, pairing conventional antibiotics with safe pump blockers—or drugs that dial down these global regulators—could help restore the usefulness of existing treatments against otherwise daunting multidrug‑resistant infections.
Citation: Obaid, A.J., Alkawaz, A.J. & Naser, M.S. Integrated analysis of global regulators and efflux genes associated with antimicrobial resistance reversal in multidrug resistant Klebsiella pneumoniae. Sci Rep 16, 7435 (2026). https://doi.org/10.1038/s41598-026-37272-w
Keywords: antibiotic resistance, efflux pumps, Klebsiella pneumoniae, fluoroquinolones, resistance reversal