Clear Sky Science · en
Imaging intercellular biomolecules by using fluorescent protein indicators with lipid-PEG anchors
Watching Cells Talk in Real Time
Our brains and bodies rely on a constant chemical conversation between cells. Ions and neurotransmitters shuttle messages across tiny gaps, but these fast, fleeting signals are hard to see without disturbing them. This study introduces a simple way to "paint" living cells with glowing sensors that latch onto their outer surface, allowing scientists to watch these messages—especially potassium ions and the neurotransmitter glutamate—live and in detail. 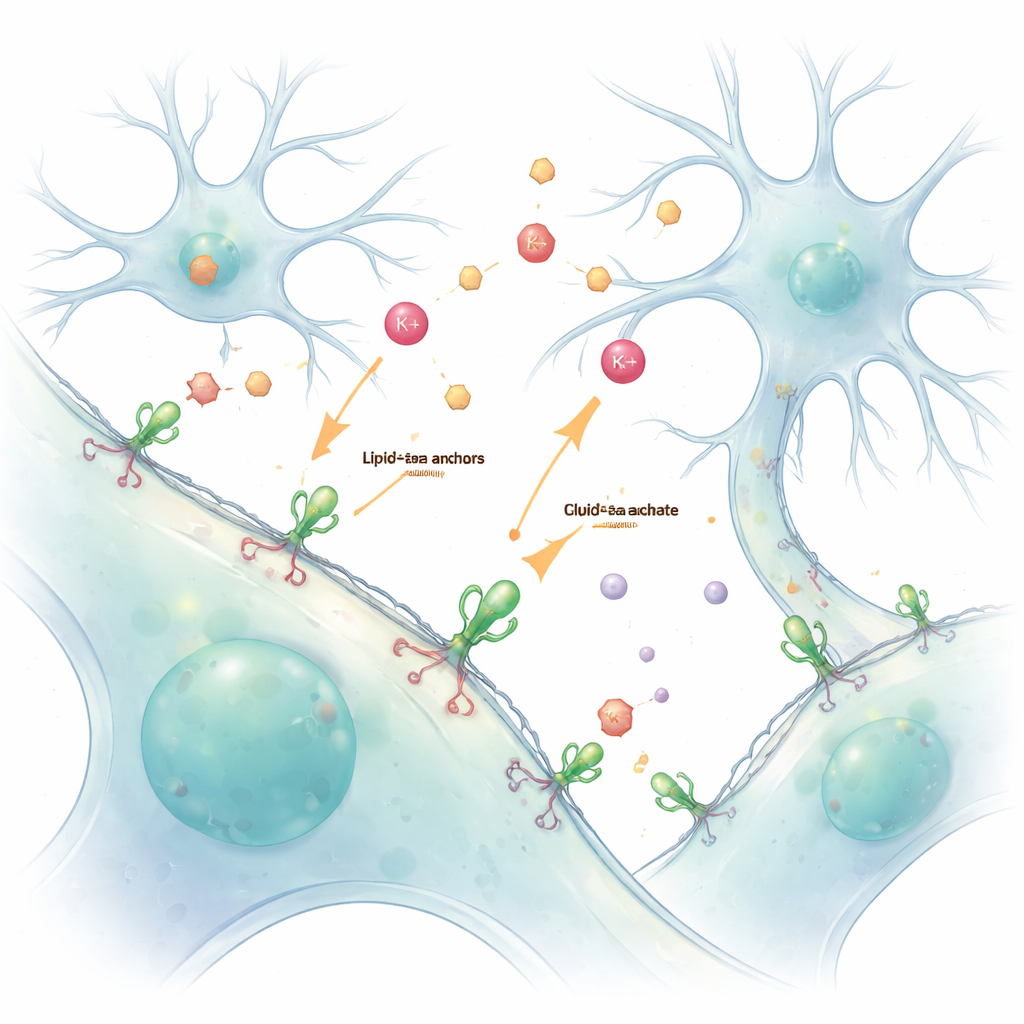
A New Way to Tag the Cell Surface
Traditionally, researchers engineer cells to make fluorescent protein sensors from their own DNA. While powerful, this genetic approach often misplaces the sensors inside the cell, making it difficult to separate what is happening inside from what is happening outside. The authors solved this by using a chemical handle called a lipid–PEG anchor. One end of this molecule embeds itself into the outer membrane, while the other end is chemically linked to a fluorescent protein sensor. When these prepared sensors are simply added to living cells, they spontaneously coat the outside surface, forming a stable, glowing layer that reports what is going on in the surrounding fluid.
Turning Potassium and Glutamate into Light
The team adapted two existing fluorescent indicators: GINKO2, which responds to potassium ions, and R-iGluSnFR1, which responds to glutamate. They purified these proteins from bacteria and then attached the lipid–PEG anchors. In cultured human cells, the anchored potassium sensor formed a smooth outline along the cell surface and brightened whenever extracellular potassium increased, then dimmed again when levels fell. Importantly, its light-emitting properties and sensitivity closely matched the original, unmodified sensor in test-tube experiments, showing that anchoring did not blunt its performance. The same was true for the glutamate sensor, which remained responsive after anchoring and clearly highlighted glutamate in the space just outside cells.
Listening to Neurons in Cultures and Slices
The researchers then moved to nerve cells from mouse hippocampus, a brain region important for learning and memory. They coated these neurons with the lipid–PEG–anchored glutamate sensor and loaded the insides of the same cells with a separate calcium dye that reports electrical activity. Using total internal reflection microscopy, they could watch red flashes from glutamate at the cell surface while green signals inside the cells reported firing. This dual-color view showed that the anchored sensors could track spontaneous neurotransmitter release without any genetic modification of the neurons themselves.
Mapping Chemical Waves Across Brain Tissue
Next, the team tested the method in acute brain slices, which preserve the natural wiring of the hippocampus. They applied the anchored potassium sensor to slices and electrically stimulated nerve fibers. The anchored sensor produced clear, repeatable fluorescence increases after each stimulus, while non-anchored sensors quickly diffused away and gave weak, unstable signals. By bathing the slices in solutions with known potassium levels, and correcting for the tissue’s own faint glow, the authors built a calibration curve and estimated actual extracellular potassium concentrations. Baseline levels were about 2.5 millimolar and rose to roughly 3.4 millimolar with modest stimulation, in line with classic electrode measurements. Anchored sensors for both potassium and glutamate revealed how these signals spread differently: glutamate changes stayed close to active synapses, while potassium increases extended hundreds of micrometers, hinting at broader effects on network excitability. 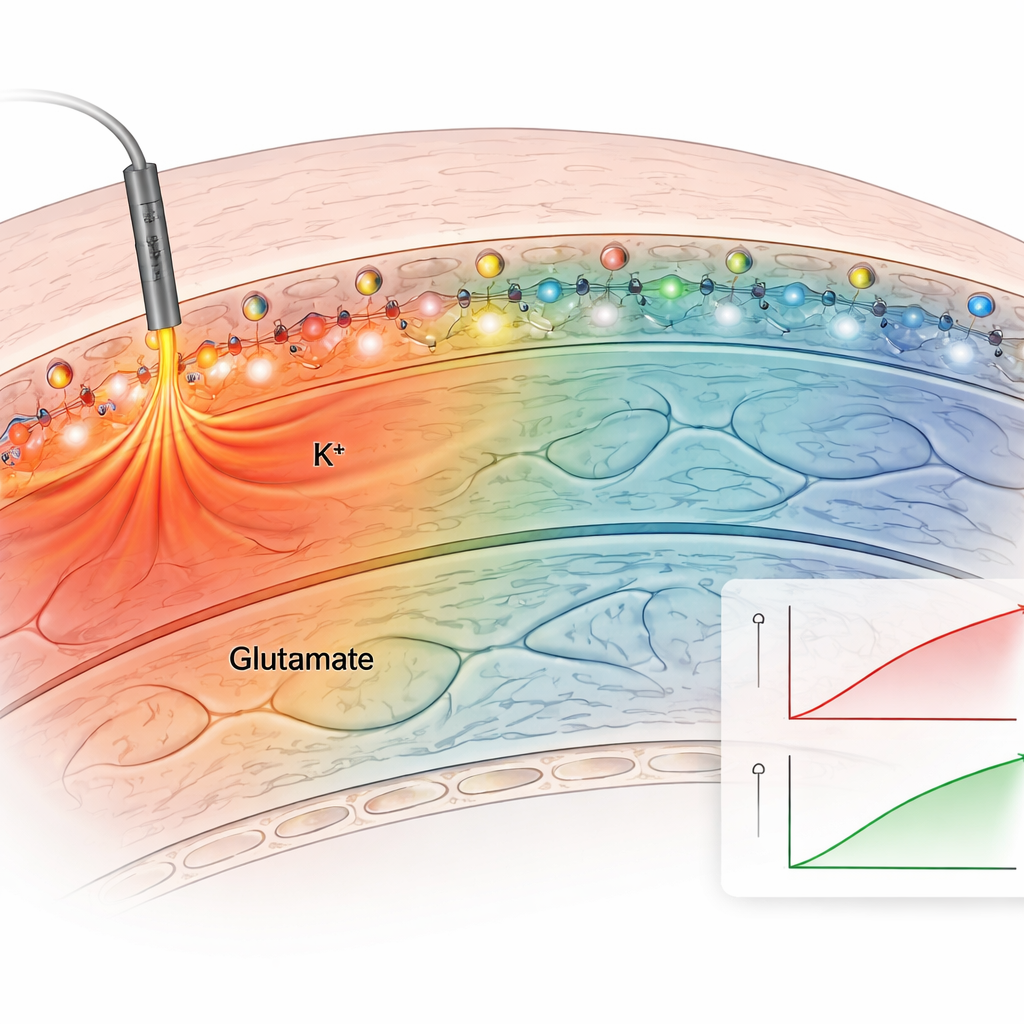
Probing How the Brain Clears and Generates Signals
Because the sensors are fixed at the cell surface, the authors could dissect which processes shape these extracellular waves. Blocking glutamate transporters, which normally mop up glutamate, caused the anchored glutamate sensor to show larger and longer-lasting signals, confirming that local uptake limits glutamate spread. Blocking AMPA-type glutamate receptors reduced both electrical responses and potassium signals, indicating that most of the potassium rise comes from postsynaptic neurons activated by synaptic input. Blocking sodium channels with tetrodotoxin nearly abolished potassium changes, tying them directly to action potentials. Together, these experiments illustrate how anchored sensors can turn complex, distributed chemistry into intuitive movies and maps.
Why This Matters for Future Brain Research
For non-specialists, the key message is that this work offers a quick, non-genetic way to coat living cells and tissues with highly sensitive molecular reporters. Instead of re-engineering each cell type, scientists can add pre-made sensor proteins from the outside and immediately visualize how ions and neurotransmitters change in real time at the cell surface. This approach improves control, reproducibility, and spatial precision while avoiding the challenges of gene delivery, especially in delicate preparations like acute brain slices. As more fluorescent indicators are developed for different chemicals, the same lipid–PEG anchoring strategy could become a widely used toolkit for watching the chemical language of cells across the body.
Citation: Mita, M., Kiyosue, K. & Tani, T. Imaging intercellular biomolecules by using fluorescent protein indicators with lipid-PEG anchors. Sci Rep 16, 6964 (2026). https://doi.org/10.1038/s41598-026-37240-4
Keywords: extracellular signaling, fluorescent biosensors, glutamate imaging, potassium dynamics, hippocampal neurons