Clear Sky Science · en
Dynamic graph convolution with comprehensive pruning and GNN classification for precise lymph node metastasis detection
Why tiny changes in lymph nodes matter
When breast cancer spreads, its first stop is often the lymph nodes, tiny filters that sit along the body’s drainage system. Spotting whether cancer cells have reached these nodes is one of the most important clues doctors use to choose surgery, chemotherapy, and radiation. Yet even expert pathologists can miss very small clusters of cancer cells on digital microscope images, especially when they look almost identical to healthy tissue. This study introduces a new artificial-intelligence framework that treats tissue as a network of connected regions, allowing it to find subtle signs of spread with remarkable accuracy.
Turning tissue images into networks
The researchers work with enormous digital slides of stained tissue, called whole-slide images, from breast cancer lymph node biopsies. These images contain millions of pixels and a bewildering mix of cell types, colors, and textures. To tame this complexity, the team first cleans the images: they normalize brightness and color, reduce noise, and generate augmented copies by rotating and flipping patches, so the computer learns to cope with natural variation. Each image patch is then broken into small, fairly uniform regions ("superpixels"), which become the dots—or nodes—in a graph, while the relationships between neighboring regions form the connecting lines—or edges. This network view preserves the irregular shapes and layouts of real tissue better than traditional grid-based image methods.
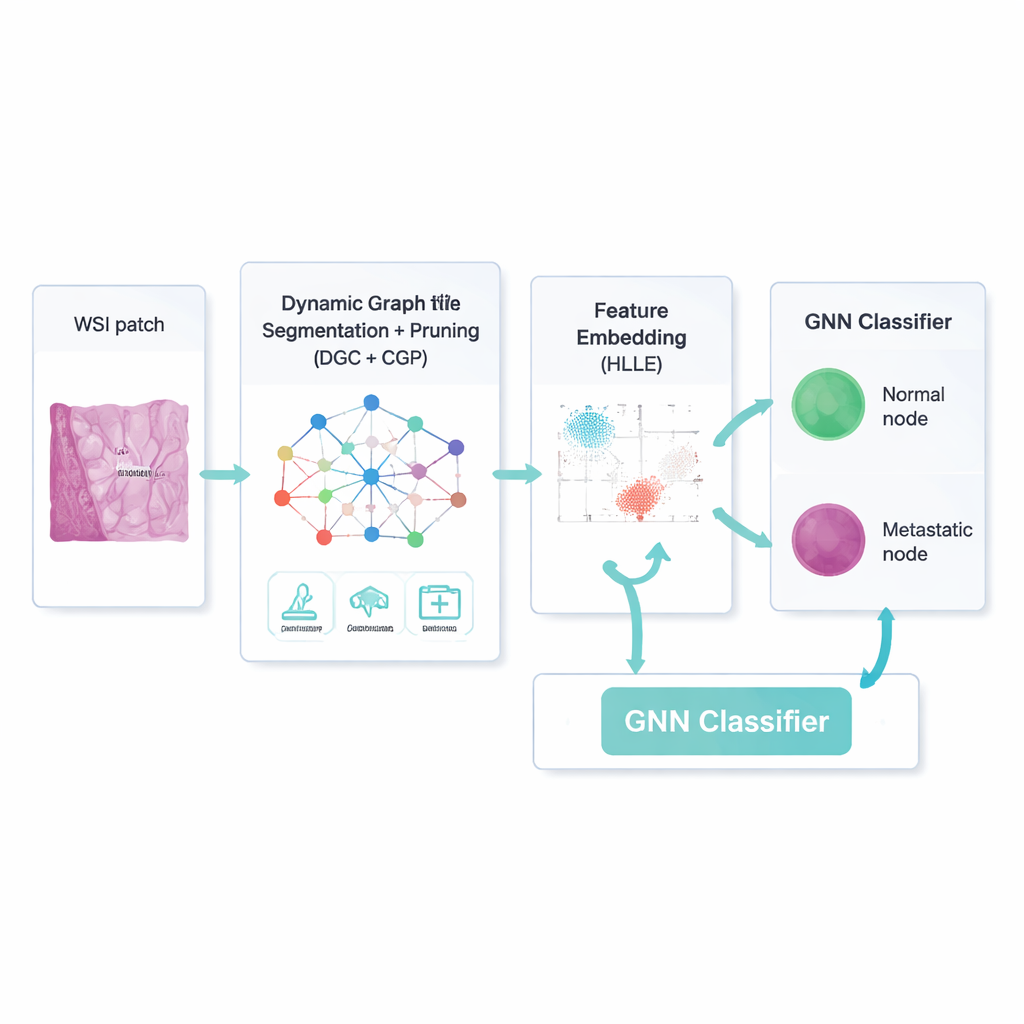
A smarter way to focus on what matters
Simply building a network is not enough; many connections and image features are irrelevant or even misleading. The framework therefore uses a dynamic graph convolutional autoencoder—a type of neural network that learns both what each region looks like and how regions influence one another. An added "attention" mechanism teaches the model to weigh some image channels more than others, for example emphasizing edges at a tumor boundary. At the same time, a strategy called Comprehensive Graph Gradual Pruning steadily trims away unhelpful pieces: weak connections between regions, less useful numerical features, and low-impact model weights. This pruning happens during training, not after, so the system learns to do more with less, ending up both faster and easier to interpret.
Compressing patterns without losing their shape
After the model has separated likely lymph node regions from background tissue, it still has to describe each region in a way that is compact but meaningful. To do this, the authors use a technique called Hessian-based Locally Linear Embedding. In simple terms, it squeezes many numerical features down into a smaller set while trying to preserve the curved "shape" of how real examples are arranged in feature space—for instance, how tiny metastases differ from normal immune cells along subtle texture or color patterns. These compressed descriptions become the input to a graph neural network classifier, which again works on the network of regions and their cleaned-up connections, deciding for each node whether it is metastatic or not.
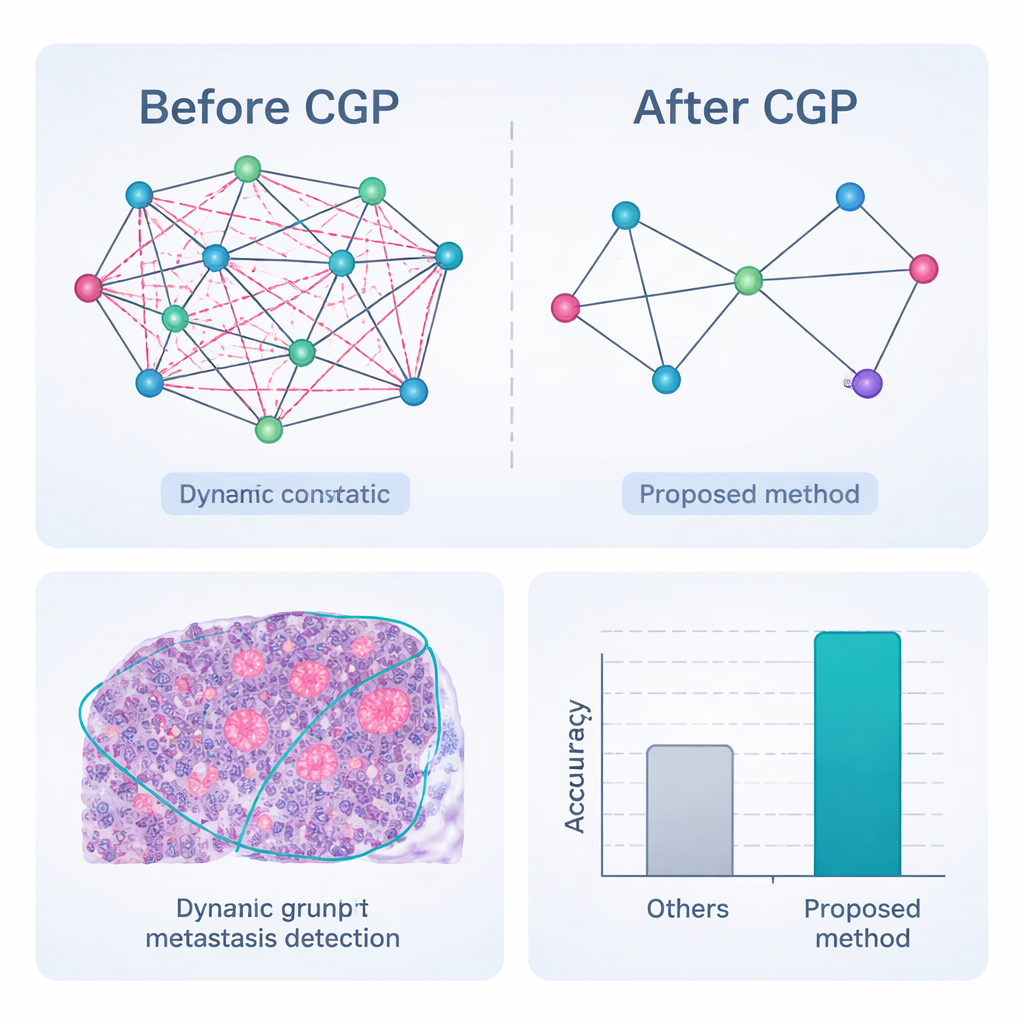
Putting the framework to the test
The complete pipeline—pre-processing, dynamic graph segmentation with pruning, feature embedding, and graph-based classification—was evaluated on CAMELYON17, a public collection of 1,000 expertly annotated lymph node slides from breast cancer patients. Compared with a range of strong deep-learning competitors, including popular convolutional networks and hybrid transformer models, the new method achieved the highest scores across nearly all measures. It correctly classified nodes as cancerous or not in 98.65% of cases and showed better agreement with expert outlines of tumor regions, especially for very small or faint metastases. Crucially, because the graph is aggressively pruned, the system reaches these results with far fewer calculations and lower memory use, making it more suitable for real-time use in busy pathology labs.
What this means for patients and clinicians
In everyday terms, this work shows how thinking of tissue as a smartly trimmed network of connected regions can help computers act as extremely careful second readers of lymph node slides. By focusing attention and computational power on the most informative structures while discarding noise, the framework is better at spotting tiny seeds of cancer that might otherwise be missed, and it does so efficiently enough to be practical. Although further clinical validation is needed, such tools could support pathologists in delivering faster, more consistent decisions about whether cancer has spread—information that directly shapes treatment plans and, ultimately, patient outcomes.
Citation: H. N., C., N., S., S., A.R. et al. Dynamic graph convolution with comprehensive pruning and GNN classification for precise lymph node metastasis detection. Sci Rep 16, 6682 (2026). https://doi.org/10.1038/s41598-026-37193-8
Keywords: lymph node metastasis, digital pathology, graph neural networks, medical image segmentation, breast cancer