Clear Sky Science · en
Genome-wide screens identify core regulators of cell surface prion protein expression
Why this matters for brain health
Prion diseases, such as Creutzfeldt–Jakob disease in humans and "mad cow" disease in cattle, are rare but always fatal brain disorders. A central culprit is a normal brain protein, called the prion protein, that can misfold and spread damage from cell to cell. The more of this protein sits on the surface of nerve cells, the easier it is for disease to take hold. This study set out to map, across the entire genome, which genes control how much prion protein appears on the outside of neuron-like cells. That map may help scientists design new ways to lower this protein and potentially slow multiple neurodegenerative diseases.
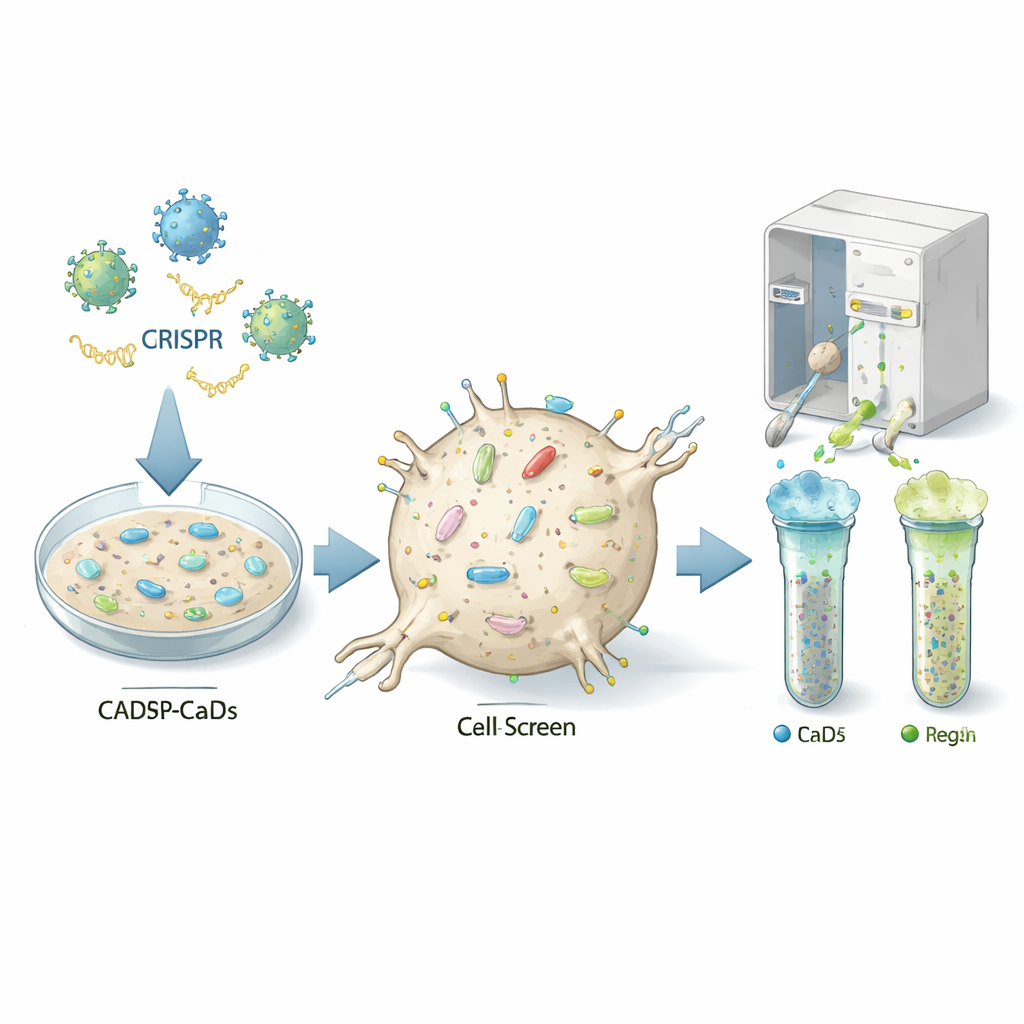
Finding the cell’s control knobs
The authors used a powerful gene-editing approach called CRISPR to switch off almost every gene, one by one, in a mouse neuron-like cell line that can be infected by prions (called CAD5 cells). Each cell received a different genetic "hit," so the resulting population contained millions of variants, each missing a specific gene. The team then stained the cells with fluorescent antibodies that recognize the normal prion protein on the cell surface and used a cell-sorting machine to separate cells with unusually low or high levels of this protein. By sequencing which guide RNAs were enriched in the low or high groups, they could infer which knocked-out genes normally act as on- or off-switches for prion protein at the cell surface.
Two cell states, overlapping answers
Neurons do not all look or behave the same over their lifetime, so the researchers asked whether the same genes control prion protein in different cell states. CAD5 cells can be kept in a fast-growing, less specialized state or pushed, by removing serum from the culture medium, to adopt a more mature, neuron-like form. The team ran the same genome-wide CRISPR screen in both conditions. In the undifferentiated (less mature) cells they validated 46 genes that increase, and 21 that decrease, surface prion protein when present. In the differentiated (more neuron-like) cells they confirmed 41 positive and 13 negative regulators. Twenty-three genes—mostly those helping attach a lipid “anchor” to the protein—were shared between both cell states, highlighting a core regulatory machinery that operates regardless of maturity.
Key assembly lines that matter most
Further analysis revealed that many of the newly identified genes belong to known cellular “assembly lines” that modify proteins as they travel to the cell surface. One major pathway builds the GPI anchor, a small fat-rich structure that tethers prion protein to the outer face of the cell membrane. Disrupting almost any step in this pathway reduced how much prion protein reached the surface, in both immature and mature cells. A second pathway involves N-linked glycosylation, in which complex sugar chains are added to proteins as they pass through the cell’s internal membranes. Genes in this sugar-adding pathway mainly emerged as important in the less mature cells. When the researchers treated cells with small molecules that block specific glycosylation steps, surface prion protein levels dropped by roughly one-third without killing the cells, confirming the genetic findings.
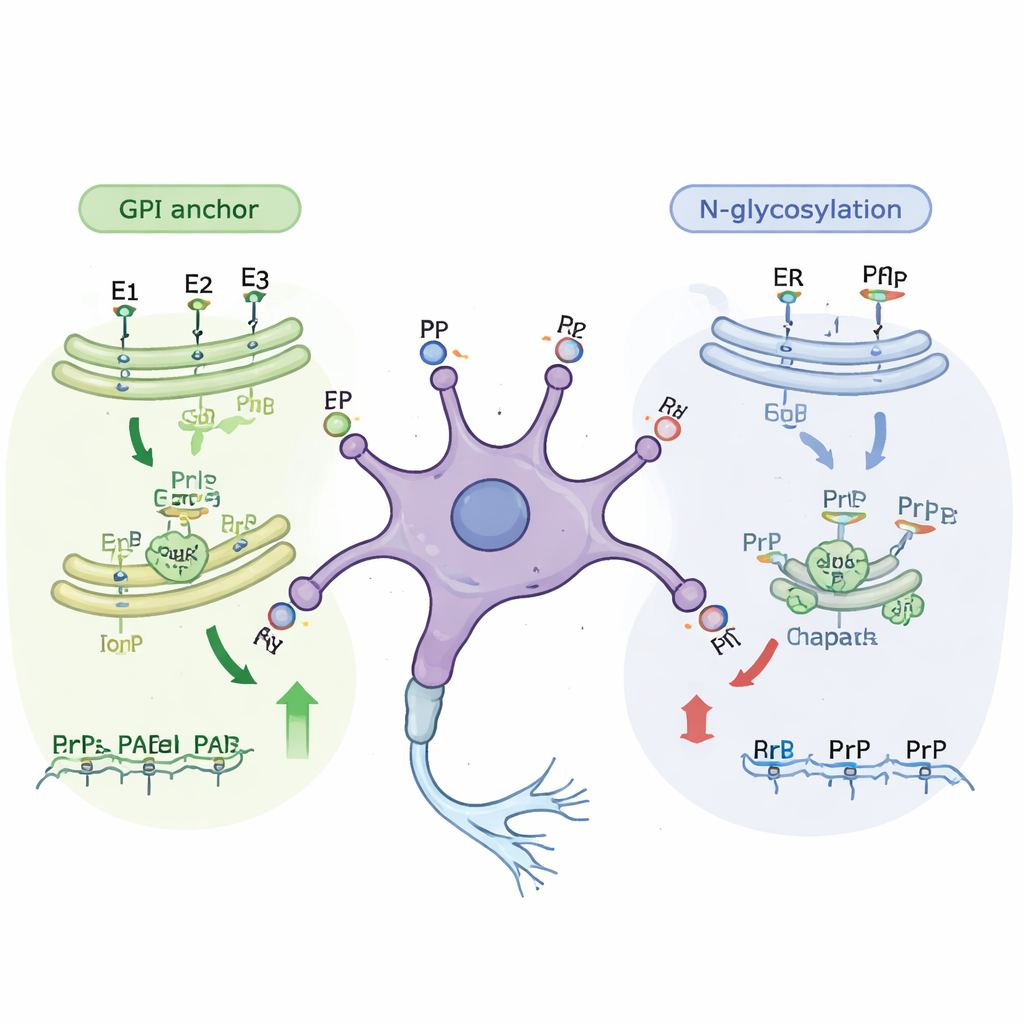
Helper proteins and stress responses
The screens also highlighted molecular chaperones—proteins that help other proteins fold correctly—as important prion regulators. In particular, Hspa5 (also called BiP), a central chaperone in the cell’s protein-folding compartment, emerged as a positive regulator in the more neuron-like cells. When the researchers used a drug to inhibit Hspa5, surface prion protein levels fell in both cell states, again without obvious harm to the cells. Other hits included genes involved in moving proteins through the cell, controlling how genes are turned on or off, and several proteins linked to synapse function and other brain diseases such as Alzheimer’s and ALS. Together, these results show that prion protein levels at the cell surface are shaped by a web of pathways spanning protein production, modification, trafficking, and quality control.
What this means for future treatments
This work provides the first comprehensive catalogue of genes that control how much prion protein appears on the surface of neuron-like cells that are susceptible to prion infection. Some of these genes, especially those in the GPI-anchor and N-glycosylation pathways and the Hspa5 chaperone system, emerge as promising starting points for drug discovery: dialing down their activity should lower the amount of prion protein available to misfold, and past studies show that even partial reductions can significantly delay disease in animals. At the same time, the clear differences between immature and mature cells underline that brain cell state matters when choosing targets. While more work is needed to test how manipulating these genes affects actual prion infection and other neurodegenerative conditions in living brains, this study offers a roadmap of cellular levers that researchers can explore to slow or prevent these devastating diseases.
Citation: Beauchemin, K.S., Supattapone, S. Genome-wide screens identify core regulators of cell surface prion protein expression. Sci Rep 16, 5895 (2026). https://doi.org/10.1038/s41598-026-37137-2
Keywords: prion protein, CRISPR screen, neurodegeneration, protein glycosylation, GPI anchor