Clear Sky Science · en
DANEELpath open source digital analysis tools for histopathological research in neuroblastoma models
Why tiny tumors in gel matter
Cancer is not just a mass of rogue cells; it is a whole neighborhood where cells talk to each other and to the scaffolding around them. In childhood cancers like neuroblastoma, this surrounding support structure can help tumors grow and resist treatment. This study introduces DANEELpath, a free, open-source set of digital tools that turns microscope images of lab-grown mini-tumors into rich, quantitative maps. By doing so, it helps scientists understand how tumor cells arrange themselves, how they respond to experimental drugs, and how those insights might translate to better therapies for children.
Building small tumor worlds in the lab
To mimic the complex environment of a real tumor, researchers grow neuroblastoma cells inside soft 3D hydrogels made from gelatin and silk. In this setting, cells naturally form round clusters, much like tiny tumors. A key player in this environment is vitronectin, a sticky protein in the extracellular matrix that helps cells attach and can drive more aggressive disease. The team prepared hydrogels with and without extra vitronectin and exposed some of them to cilengitide, an experimental drug that blocks vitronectin-related cell receptors. After weeks of growth, the gels were sliced, stained, and scanned as high-resolution whole-slide images, creating a digital version of each mini-tumor world.
Turning complex images into measurable patterns
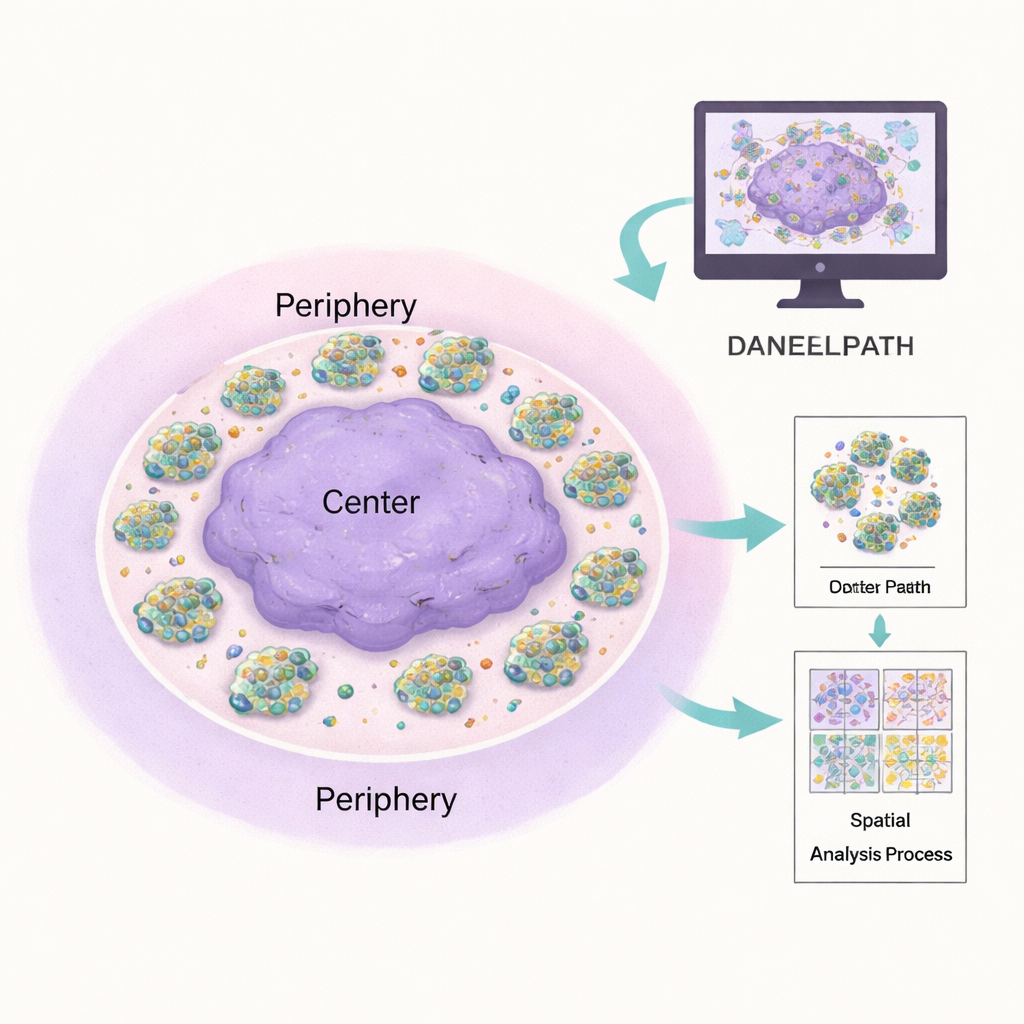
Traditional pathology depends on expert eyes at a microscope, which is powerful but hard to standardize and scale. DANEELpath plugs into the popular open-source platform QuPath and automates much of this work. Using deep-learning models called U-Nets, the toolkit can accurately outline each cell cluster in common stains, even when clusters vary from a few cells to hundreds. It then applies mathematical methods to split each irregularly shaped hydrogel into a balanced "center" and "periphery" ring, ensuring that comparisons between inner and outer regions are fair regardless of the gel’s shape. This allowed the authors to confirm that neuroblastoma clusters tend to be denser toward the hydrogel edge and that this pattern changes depending on vitronectin content and drug treatment.
Measuring cell spacing and cluster neighbors
Beyond simply counting clusters, DANEELpath examines how cells and clusters are arranged in space. Inside each cluster, another tool finds individual cell nuclei and measures the distance to each cell’s nearest neighbor, summarizing how tightly packed the cells are. Between clusters, the toolkit offers three ways to define "neighbors," including a method based on Voronoi diagrams that draws influence zones around each cluster. By comparing these measures under different conditions, the researchers showed that cilengitide changes how many neighboring clusters a large cluster has and how those neighbors are distributed, but in ways that depend on whether vitronectin was added to the gel. This highlights how the surrounding matrix and mechano-drugs together shape tumor organization.
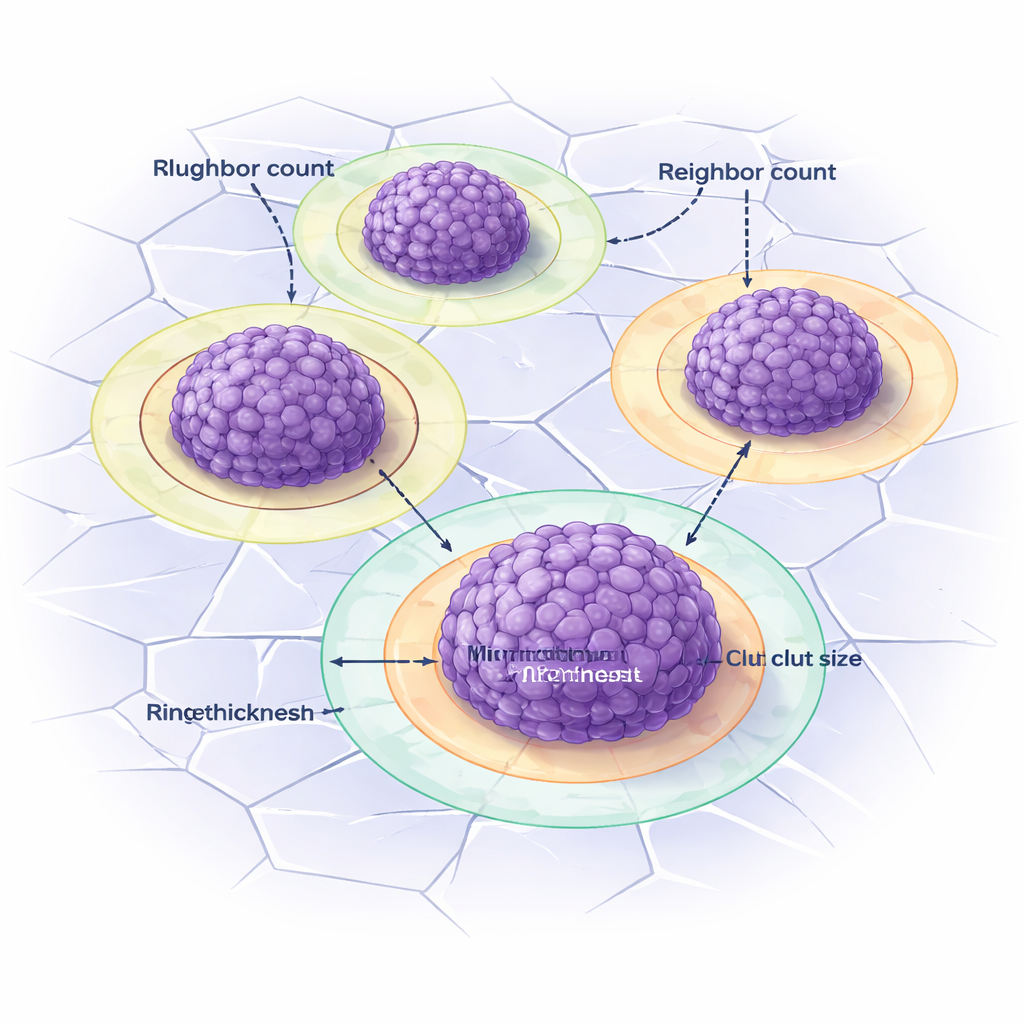
Seeing the halo around tumor clusters
Vitronectin in these models often forms striking rings around clusters: a bright "corona" when only cell-made vitronectin is present, and a pale halo when extra vitronectin is built into the gel. Separating these rings from the cluster core is tricky by eye. DANEELpath tackles this by first drawing Voronoi-based influence regions around each cluster, then expanding outward just enough to capture the ring without overlapping neighbors. Simple image-contrast rules are used to detect coronas and halos, and the toolkit automatically measures their thickness. Similar strategies were also applied to real patient samples, for example mapping immune cells at the edge of tumor regions or defining zones around blood vessels to study how vitronectin and other markers are arranged in human neuroblastoma.
What this means for future cancer research
In essence, this work turns static pathology images into detailed, reproducible measurements of how tumor cells and their surroundings are organized. For non-specialists, that means researchers can now more easily test how new drugs, scaffold materials, or genetic changes alter not just whether tumors grow, but how they grow in space. Because DANEELpath is open-source, works through a graphical interface, and runs on ordinary computers, it lowers the barrier for labs worldwide to adopt advanced image analysis. Over time, such tools could help link patterns seen in 3D models and patient tissue to outcomes in the clinic, guiding more targeted and effective treatments for children with high-risk neuroblastoma.
Citation: Vieco-Martí, I., López-Carrasco, A., Navarro, S. et al. DANEELpath open source digital analysis tools for histopathological research in neuroblastoma models. Sci Rep 16, 6162 (2026). https://doi.org/10.1038/s41598-026-37134-5
Keywords: neuroblastoma, digital pathology, 3D hydrogels, extracellular matrix, deep learning