Clear Sky Science · en
Development and preliminary evaluation of real-time PCR assays for six lactic acid bacteria
Friendly Germs Behind Everyday Foods
Many of the foods that line supermarket shelves—from yogurt and cheese to pickles and fermented vegetables—owe their flavor and possible health benefits to friendly bacteria known as probiotics. But to harness these "good bugs" safely and reliably, companies must be absolutely sure which bacterial species are really present in their products. This study describes new laboratory tests that can quickly and accurately identify six promising probiotic bacteria used in food, helping bridge the gap between cutting‑edge microbiology and the everyday foods we eat.
Why These Probiotic Bacteria Matter
The six bacteria at the heart of this work all belong to a broader group called lactic acid bacteria, long used in fermentation. Recent research suggests they can do much more than sour milk or cabbage. Some strains of Ligilactobacillus agilis and Ligilactobacillus salivarius may help fend off harmful gut germs, calm inflammation, and support a healthy intestinal barrier. Limosilactobacillus fermentum has been linked to better blood pressure control and antioxidant effects in animal studies. Lactobacillus johnsonii appears to help balance gut microbes in ways that could protect several organs. Pediococcus pentosaceus and Weissella cibaria show promise for lowering cholesterol, fighting spoilage, and even supporting oral health. With such a range of potential benefits, the food industry is eager to use these species more widely—but only if they can be identified with confidence.
The Challenge of Telling Look‑Alike Microbes Apart
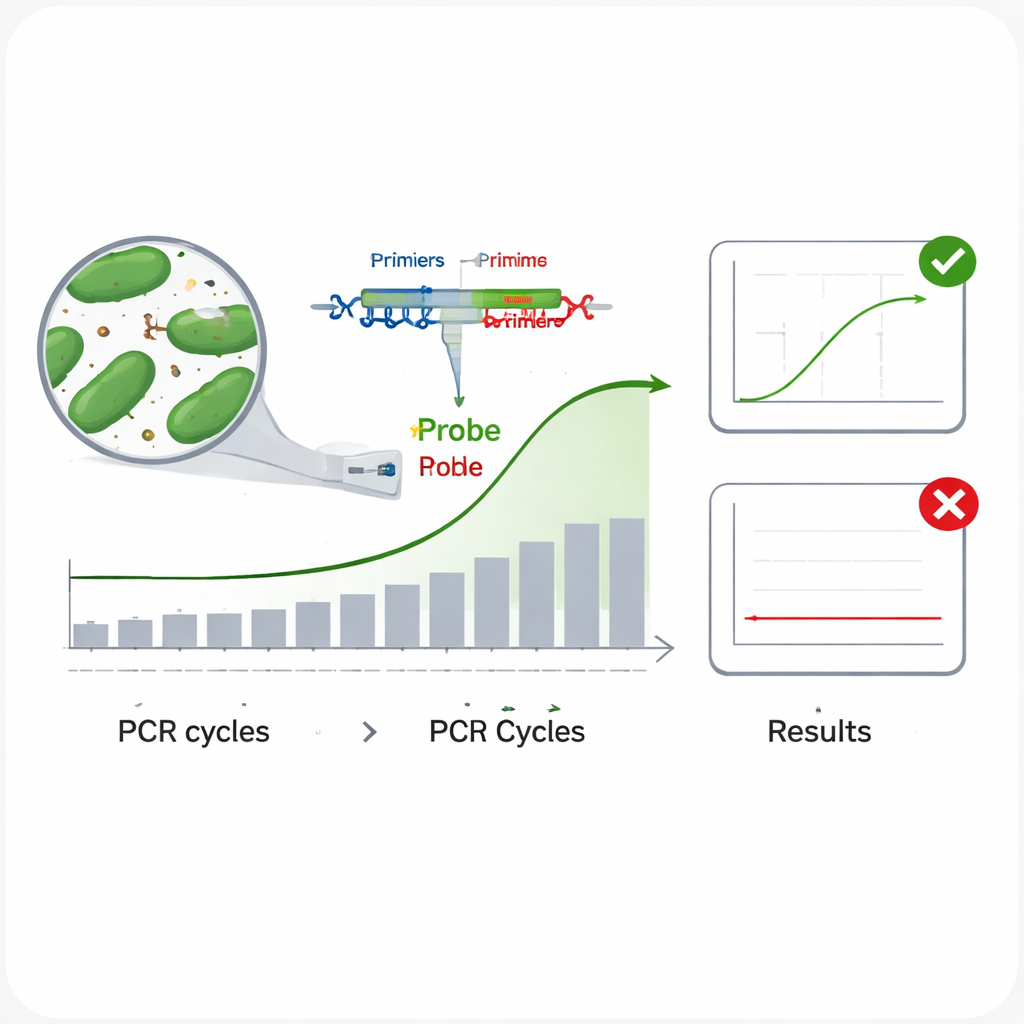
Traditional ways of identifying bacteria—growing them in the lab and running biochemical tests—are slow and labor‑intensive. Modern DNA‑based methods, especially real‑time PCR, are far faster. In real‑time PCR, short pieces of DNA called primers and a fluorescent probe home in on a unique stretch of a microbe’s genetic code; when the right microbe is present, the machine detects a glowing signal as the DNA is copied cycle after cycle. The problem is that closely related bacteria can share very similar DNA, so commonly used genetic regions such as the 16S rRNA gene sometimes fail to distinguish one species from another. That can lead to tests that miss the target (false negatives) or light up for the wrong species (false positives), both of which are unacceptable for product labeling and safety.
Designing Sharper Molecular "Barcodes"
To overcome this, the researchers combed through complete genome sequences to find short DNA regions that were both highly conserved within each target species and clearly different from other bacteria. For each of the six probiotics, they designed a matched set of primers and a probe with carefully tuned length, base composition, melting temperature, and minimal tendency to fold back on themselves or stick to each other. Computer checks using the BLAST database and sequence alignment software confirmed that the selected DNA regions were stable within the species yet distinct from non‑targets. The team then worked out a standard reaction mix and heating program so that all tests could be run under consistent and practical conditions in the same type of real‑time PCR machine.
Putting the New Tests Through Their Paces
The scientists next evaluated how well each assay performed in practice. To test inclusivity—whether a test can detect many different strains of the same species—they ran each assay on multiple DNA samples from its target bacterium. In every case, all tested strains produced a clear amplification curve, suggesting a low risk of missing genuine targets. To check specificity—whether the assay ignores non‑target species—they challenged each test with DNA from 13 other intestinal bacteria, including related lactic acid bacteria and common gut microbes like Escherichia coli. None of these produced a signal, indicating a very low risk of false alarms. The team also examined amplification efficiency by running a series of ten‑fold dilutions of DNA and confirmed that all six assays copied their targets at roughly 95–100% efficiency, close to the ideal. Finally, they measured precision by repeating runs at two DNA levels and found that tiny variations between repeats and between separate experiments stayed well below accepted limits.
What This Means for Future Foods
In simple terms, the authors have built six finely tuned DNA "fingerprint" tests that can tell key probiotic species apart quickly, accurately, and reliably. While they caution that broader testing on more strains, more non‑target species, and additional PCR instruments is still needed before routine industrial adoption, the early results are encouraging. For consumers, advances like this help ensure that foods advertised as containing specific probiotics really do contain the right microbes, in turn supporting honest labeling, better quality control, and more trustworthy research on how these tiny partners affect our health.
Citation: Li, SJ., Cui, B., Li, W. et al. Development and preliminary evaluation of real-time PCR assays for six lactic acid bacteria. Sci Rep 16, 6165 (2026). https://doi.org/10.1038/s41598-026-37047-3
Keywords: probiotics, lactic acid bacteria, real-time PCR, food microbiology, microbial identification