Clear Sky Science · en
Predicting kidney graft survival with a machine learning model based on for-cause biopsy transcriptomics
Why this matters for transplant patients
For people with kidney failure, a transplant can mean the difference between life on a dialysis machine and a return to everyday activities. Yet many transplanted kidneys still fail years after surgery, often because the body’s immune system slowly attacks the new organ. This study explores whether patterns in gene activity from routine kidney biopsies can be combined with modern machine learning to spot which grafts are in real danger long before standard tests show trouble.
Looking inside the transplant at the molecular level
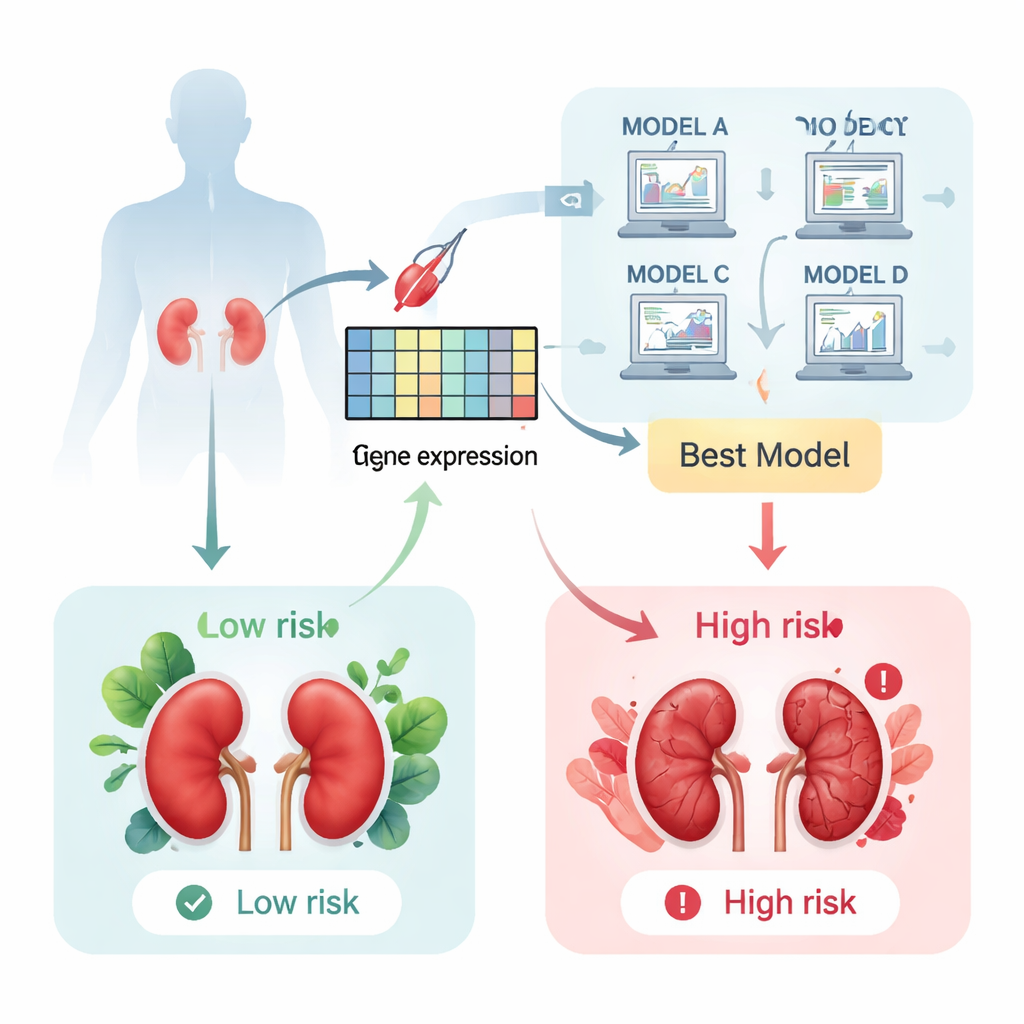
After a kidney transplant, doctors sometimes perform a biopsy “for cause” when blood tests or urine results suggest the organ might be under stress. Traditionally, pathologists look at these samples under a microscope to judge the level of damage. The authors of this paper asked a different question: could the genes turned on inside those biopsy samples reveal a clearer, earlier warning signal of future graft loss? To find out, they gathered gene activity data from more than 1,200 biopsies spread across six international research collections and focused on patients whose grafts either survived or later failed.
Finding an 11-gene warning signal
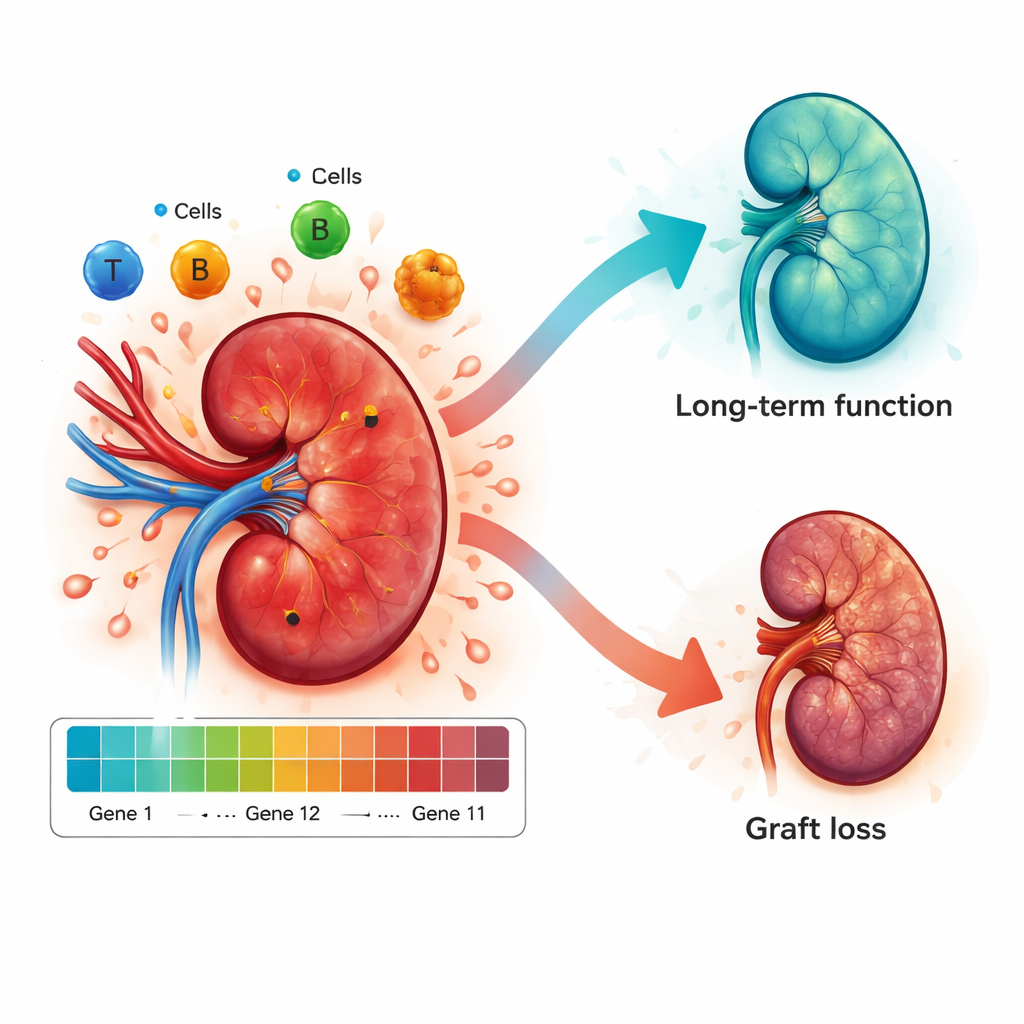
The researchers first compared biopsy samples from patients who eventually lost their transplanted kidney with those who kept good function. They searched thousands of genes and identified a small group of 11 that were consistently more active in the failing grafts. These genes were strongly linked to immune activation and inflammation, including signals that call white blood cells into the kidney and help them attack tissue. In other words, biopsies from kidneys that would later fail were already “lit up” with an aggressive immune signature, even when traditional measures might still look acceptable.
Training machines to forecast graft survival
Next, the team fed the activity levels of these 11 genes into a wide range of machine learning approaches designed to predict how long a transplanted kidney would keep working. They tested 117 different model setups and scored them on how accurately they could rank patients from lowest to highest risk of graft loss. A type of algorithm called a Gradient Boosting Machine emerged as the clear winner, correctly ordering patients with a high degree of accuracy. When the model assigned a high risk score, those patients were much more likely to lose their graft within the next few years than patients with low scores, as shown by survival curves that separated sharply over time.
Checking performance in new patient groups
To be useful in the clinic, a risk tool must work beyond the data it was built on. The authors therefore applied their 11-gene model to four completely independent biopsy collections from other centers. In these groups, detailed survival times were not always available, but experts had labeled each biopsy as showing rejection or not. The same gene-based score did a good job of distinguishing rejection from stable grafts, with accuracy measures that compare well to many medical tests used in practice. In a separate study where patients had scheduled biopsies at 0, 6, and 24 months after transplant, those who later developed chronic damage already showed higher risk scores months before the damage was obvious under the microscope.
What the model reveals about the immune battle
Beyond prediction, the researchers used the gene patterns to peek into the immune landscape inside the kidney. Biopsies with high risk scores showed more aggressive immune cell types, such as certain macrophages and T cells, and more of the chemical signals that help them swarm into the organ. Low-risk biopsies, in contrast, were richer in cell types associated with calming inflammation and promoting tissue repair. This suggests the model is not just a black box: its risk score mirrors real biological processes that either push the kidney toward rejection or support long-term peace between the organ and the recipient’s immune system.
How this could change transplant care
The study concludes that an 11-gene, machine learning–based score from routine kidney biopsies can reliably forecast which grafts are most likely to fail and can flag problems earlier than standard methods. For patients and clinicians, such a tool could one day guide personalized care: high-risk individuals might receive closer monitoring or tailored drug adjustments, while low-risk patients might avoid unnecessary procedures. Because the signature uses only a small set of genes, it could be translated into practical lab tests using existing technology. Although more prospective trials are needed before it becomes part of everyday practice, this work points toward a future in which molecular readouts and artificial intelligence help protect transplanted kidneys for the long term.
Citation: Filho, V.O.C., Passos, P.R.C., de Andrade, L.G.M. et al. Predicting kidney graft survival with a machine learning model based on for-cause biopsy transcriptomics. Sci Rep 16, 6157 (2026). https://doi.org/10.1038/s41598-026-37038-4
Keywords: kidney transplantation, graft survival, machine learning, biopsy gene expression, immune rejection