Clear Sky Science · en
Amplicon sequencing with Oxford nanopore technologies as a diagnostic alternative for small ruminant lentiviruses in sheep
Hidden infections in everyday sheep
Sheep around the world quietly carry viruses that can sap their health, shorten their lives, and cost farmers money—often without obvious symptoms for years. This study explores a new way to uncover those hidden infections using a portable DNA sequencing technology, offering a potential step-change in how we protect animal welfare, farm income, and even food security.
A slow, costly disease that is hard to spot
The work focuses on small ruminant lentiviruses (SRLV), a group of viruses that infect sheep and goats. In sheep, they cause Maedi-Visna disease, a long-lasting infection that can lead to breathing problems, arthritis, brain disease, and chronic udder inflammation. Many infected animals never show clear signs, yet the virus still lowers milk production, increases lamb deaths, and forces early culling. In some European dairy flocks, including in Spain and Greece, roughly half of the animals may be infected, making this one of the most important sheep diseases in intensive farms.
Why current tests miss many infected animals
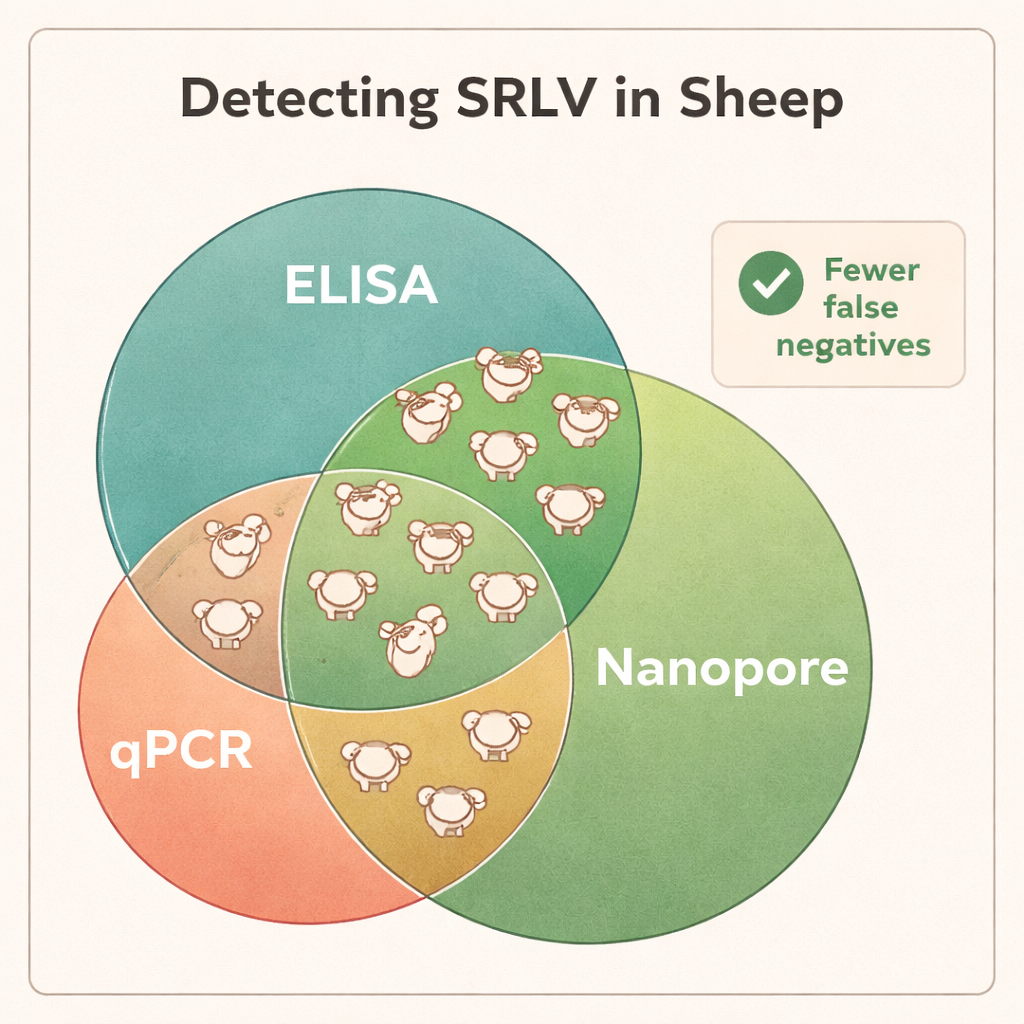
Today, farms mainly rely on blood tests that look for antibodies (ELISA) or on standard DNA tests (qPCR) to decide which animals are infected and should be removed from the flock. But SRLVs mutate and recombine rapidly, creating many slightly different viral variants. Some variants are poorly recognized by the antibody tests, and some infected sheep never develop strong antibody responses at all. qPCR, which targets short and very specific pieces of viral DNA, can also fail if those target regions change. As a result, many truly infected animals test negative and remain in the flock, quietly spreading the virus.
Using real-time DNA reading to find the virus
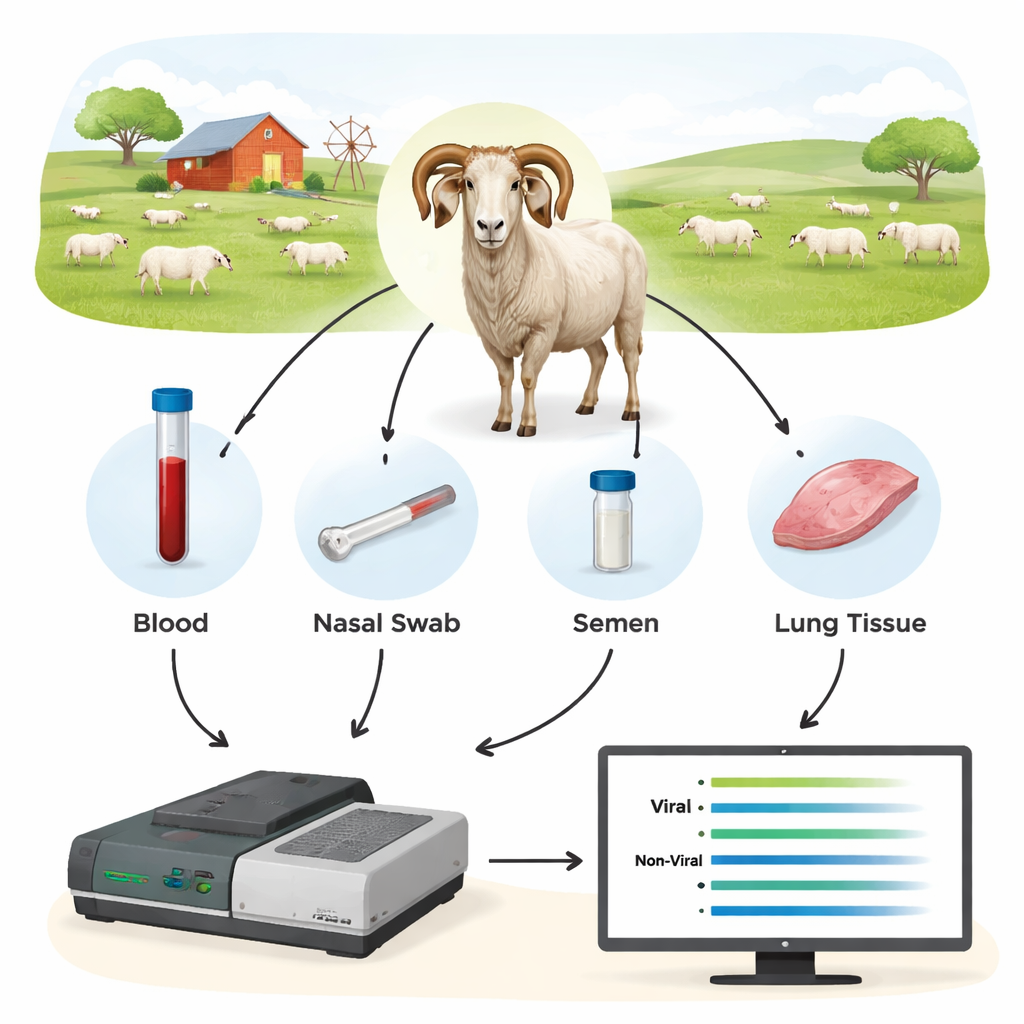
The researchers tested a third-generation DNA method called Oxford Nanopore sequencing as a new kind of diagnostic tool. Instead of searching for a single, tiny piece of viral DNA, they first amplified longer stretches of key viral genes from the animals’ samples and then sequenced those pieces in real time on a Nanopore device. They collected blood, nasal swabs, semen, and cells from blood and lungs from 44 rams and additional sheep, many of which had already been tested by conventional methods. By focusing on viral regions that are relatively conserved but still long enough to reveal the exact strain, the team could both detect infection and identify which viral types were present.
Blood works best—and finds what other tests miss
Sequencing showed that DNA from whole blood was the most reliable material for discovering SRLV, even though the virus mainly lives in a small subset of white blood cells. Lung tissue from heavily infected animals yielded very high amounts of viral DNA, but such samples are only available after slaughter. In contrast, nasal swabs, semen, and purified white blood cells offered too little viral material for consistent diagnosis. When the scientists compared Nanopore results with the standard ELISA and qPCR tests, the differences were striking: Nanopore sequencing confirmed infection in all ELISA-positive rams but also revealed that many ELISA-negative animals were in fact infected. Across different flocks, roughly 40–45% of animals labeled “negative” by ELISA turned out to carry the virus, and qPCR missed an even larger fraction. The sequencing data also uncovered coinfections with different SRLV types in some rams, information that traditional tests cannot easily provide.
From simple yes/no answers to deeper insights
Because Nanopore reads the actual viral sequences, it can go beyond a simple yes-or-no diagnosis. The team used the data to compare viral strains, build family trees of the viruses circulating in their flocks, and study subtle differences in viral proteins that might explain why some animals escape detection by standard ELISA kits. They showed that certain versions of a key viral protein, targeted by commercial antibody tests, differ notably between antibody-positive and antibody-negative animals. Over time, such information could help refine both serological tests and breeding programs aimed at selecting animals that are naturally more resistant to infection.
What this means for farmers and animal health
For non-specialists, the core message is straightforward: by reading longer pieces of viral DNA directly, Nanopore sequencing can uncover more infected sheep, earlier and more accurately, than current routine tests. It also tells us which exact virus strains are present in a flock. Although this approach is still more complex and costly than a single blood test, the technology is becoming faster, cheaper, and more portable. If integrated into control programs, it could sharply reduce the number of “hidden” carriers, improve the design of vaccines and tests, and support breeding of more resistant animals—making sheep farming more sustainable and humane.
Citation: Serrano, M., González, C., Roy, R. et al. Amplicon sequencing with Oxford nanopore technologies as a diagnostic alternative for small ruminant lentiviruses in sheep. Sci Rep 16, 6212 (2026). https://doi.org/10.1038/s41598-026-36989-y
Keywords: sheep health, lentivirus, nanopore sequencing, veterinary diagnostics, Maedi-Visna