Clear Sky Science · en
Orbitrap Astral–based proteome and phosphoproteome analysis identifies candidate proteins associated with the phosphatidate phosphatase MoPah1 in Magnaporthe oryzae
Why a rice disease enzyme matters
Rice is a staple food for billions of people, yet a single microscopic fungus, Magnaporthe oryzae, causes rice blast disease that can wipe out entire fields. This study digs into the inner chemistry of that fungus, focusing on one enzyme, MoPah1, that helps manage the cell’s fats and signaling molecules. By mapping thousands of proteins and their on–off switches, the researchers uncover how this enzyme connects basic metabolism to the fungus’s ability to infect rice plants.
A fungus that breaks into rice plants
Magnaporthe oryzae infects rice leaves using a specialized structure called an appressorium, which builds up enormous internal pressure to punch through the leaf surface. To power this process, the fungus rapidly burns through stored sugars and fats. Earlier work showed that MoPah1, an enzyme that converts a fatty molecule called phosphatidic acid into another called diacylglycerol, is vital for this lifestyle: removing the MoPah1 gene weakens the fungus and reduces its ability to cause disease. But which other proteins are linked to MoPah1, and how this enzyme fits into broader cellular networks, remained unknown.
Using next‑generation protein mapping
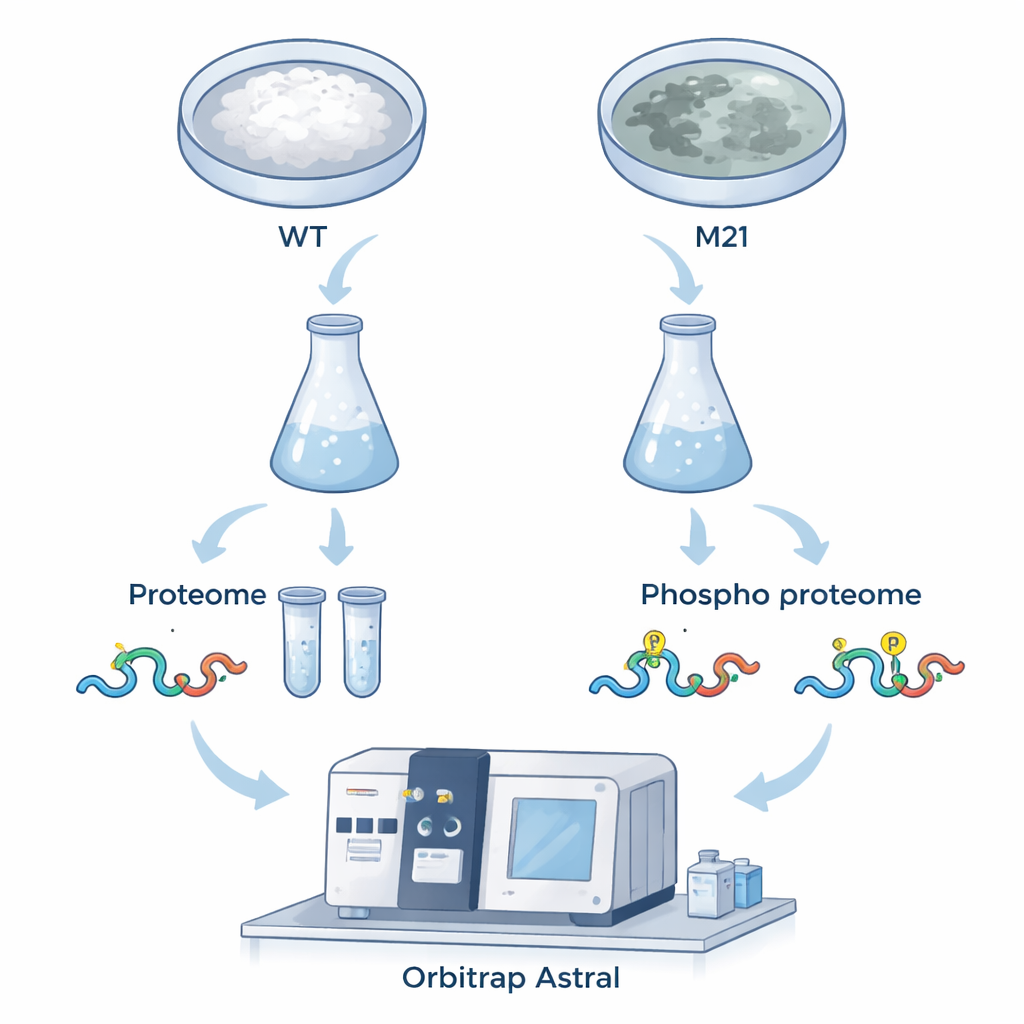
To answer these questions, the team compared normal fungi with a mutant lacking MoPah1. They focused on thread‑like growth (mycelium), because the mutant cannot produce the spores usually used for infection studies. Using an advanced mass spectrometer called the Orbitrap Astral and a data‑independent acquisition (DIA) strategy, they measured both the abundance of proteins and the presence of phosphate tags that act like molecular switches. In total, they identified 6,799 proteins and more than 15,000 phosphorylation sites, with hundreds showing clear increases or decreases in the mutant. This large, high‑quality dataset offers a detailed snapshot of how removing MoPah1 reshapes the fungal cell.
Shifts in energy use and cellular recycling
When the researchers examined which types of proteins changed, two themes stood out: metabolism of membrane fats and a “self‑eating” process called autophagy, which cells use to recycle components and survive stress. Many proteins involved in glycerophospholipids—the building blocks of cell membranes—and in related energy pathways were altered. Proteins linked to autophagy showed especially strong changes in their phosphorylation patterns, hinting that MoPah1 affects not just how fats are made and broken down, but also how the fungus recycles its own material during growth and infection. A subset of 72 proteins changed both in quantity and in phosphorylation state, marking them as prime candidates for direct or indirect control by MoPah1.
Finding MoPah1’s protein partners
To move beyond correlations, the team asked which proteins physically bind to MoPah1. They produced a MoPah1 protein fused to a “handle” (GST) in bacteria, then used it like bait to fish out interacting proteins from fungal extracts. Mass spectrometry identified 183 candidates, many of which cluster into large protein complexes, reinforcing the idea that MoPah1 sits in the middle of extensive cellular networks. Among these partners, one stood out: Pmk1, a key signaling protein in the MAPK pathway that controls appressorium formation, growth inside the plant, and responses to stress. Additional yeast two‑hybrid tests confirmed that MoPah1 and Pmk1 can interact directly.
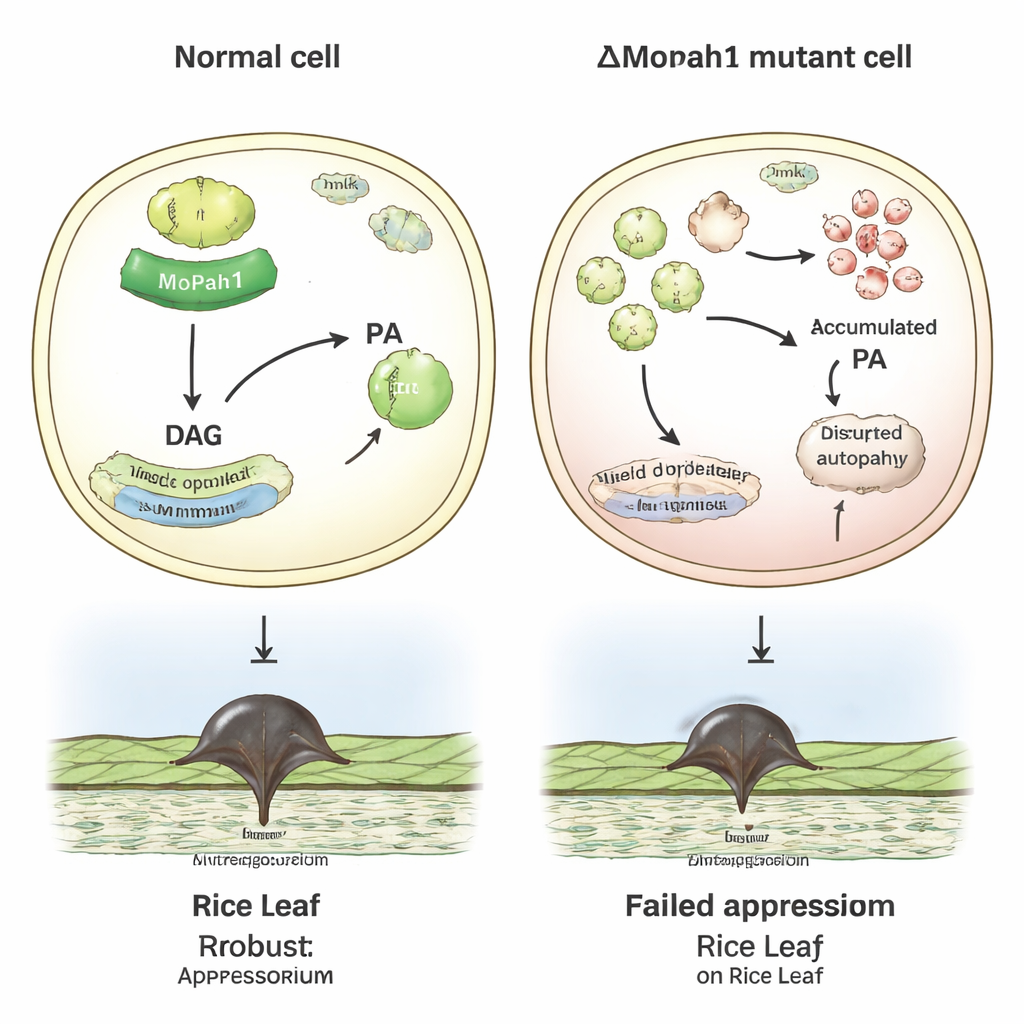
Linking fat control to infection signals
Putting these strands together, the authors propose that MoPah1 helps tune both membrane fat balance and major signaling routes in Magnaporthe oryzae. In fungi lacking MoPah1, broad disturbances in lipid metabolism, protein recycling, and MAPK signaling appear to undermine the formation and effectiveness of the infection structures needed to penetrate rice leaves. While the study was limited to mycelial samples—because the mutant cannot form spores—it still delivers a rich resource of protein and phosphorylation data, plus a shortlist of proteins most likely to work alongside MoPah1. For non‑specialists, the key takeaway is that controlling a single enzyme in a crop pathogen can ripple through many cellular systems, offering new angles for understanding, and perhaps one day disrupting, the fungus that threatens one of the world’s most important food sources.
Citation: Zhao, J., Yang, L., Shi, X. et al. Orbitrap Astral–based proteome and phosphoproteome analysis identifies candidate proteins associated with the phosphatidate phosphatase MoPah1 in Magnaporthe oryzae. Sci Rep 16, 6901 (2026). https://doi.org/10.1038/s41598-026-36940-1
Keywords: rice blast fungus, Magnaporthe oryzae, protein networks, lipid metabolism, fungal pathogenicity