Clear Sky Science · en
A hybrid learning framework for automated multiclass electrocardiogram classification with SimCardioNet
Why teaching computers to read heartbeats matters
Every time a doctor orders an electrocardiogram (ECG), they receive a squiggly line that can reveal heart attacks, dangerous rhythm problems, and early signs of disease. But reading these traces correctly takes years of training, and in many hospitals—especially in low‑resource settings—there simply are not enough heart specialists. This study presents SimCardioNet, a new artificial intelligence system designed to read ECG images automatically and accurately, even when only a small amount of expert-labeled data is available. By learning first from unlabeled ECGs and then refining itself with a modest set of labeled examples, SimCardioNet aims to bring reliable, fast ECG interpretation closer to everyday clinical practice.
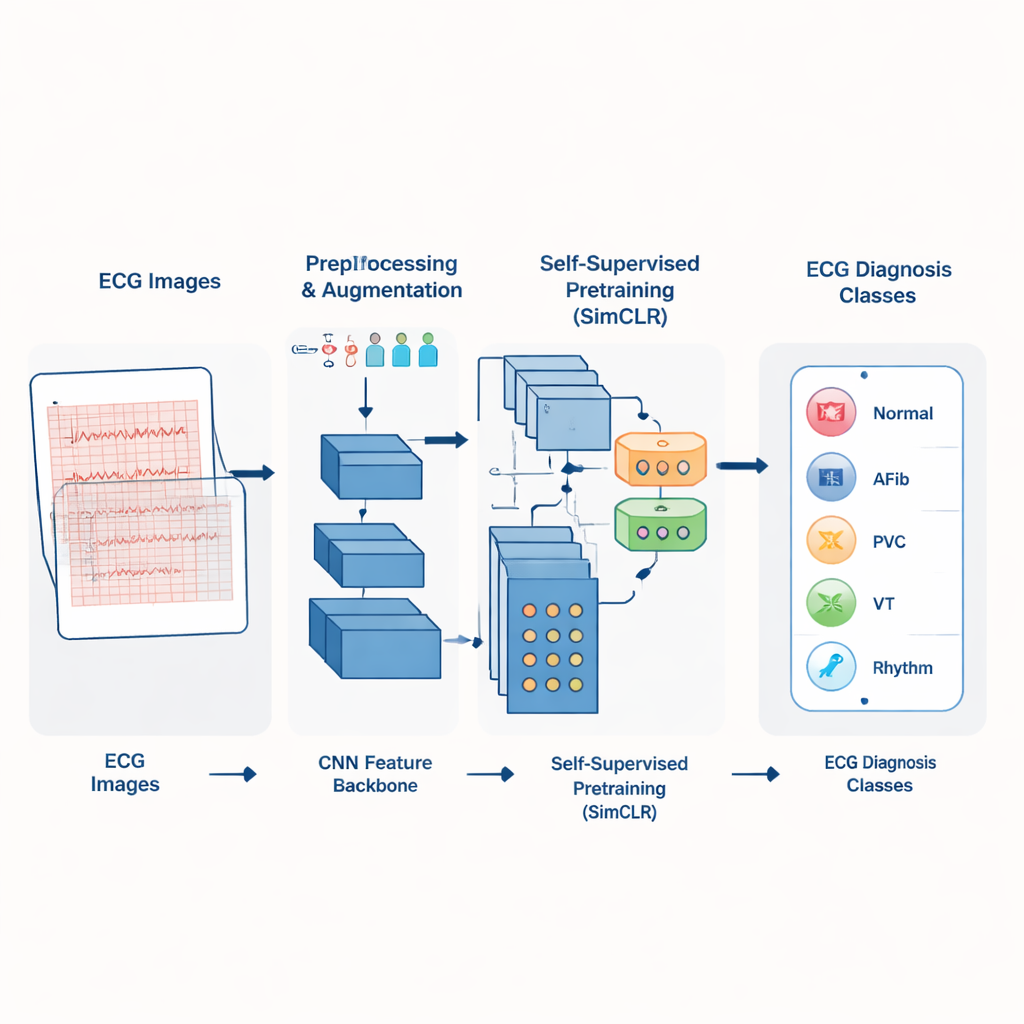
From paper printouts to smart pattern recognition
In many clinics, ECGs are stored not as clean digital signals but as scanned images or paper printouts. SimCardioNet is built to work directly with these images. The system first standardizes each ECG picture to a fixed size and applies a variety of subtle changes—small rotations, color shifts, cropping, and flips—that mimic real‑world variations in how ECGs are printed or scanned. These “augmented” versions help the model become robust to differences between hospitals and machines, so it learns to focus on the heart’s electrical patterns instead of superficial details like grid color or page layout.
A two‑step way of teaching the model
Rather than starting by asking the computer to jump straight into diagnosis, the authors use a two‑step learning process. In the first step, called self‑supervised learning, the model is shown many unlabeled ECG images and asked to recognize when two different views come from the same underlying ECG. It does this with a method known as contrastive learning: pairs of images from the same heartbeat are pulled closer together in its internal representation, while pairs from different patients are pushed apart. SimCardioNet uses a custom stack of convolutional layers (a standard deep‑learning building block for images), residual connections that ease training of deep networks, and a multi‑head attention module that helps the model focus on the most informative parts of each waveform.
Fine‑tuning the system to name heart conditions
After this “unsupervised” practice phase, the model has learned a rich sense of what ECGs typically look like. In the second step, supervised fine‑tuning, it is given labeled examples—ECGs tagged by experts as normal, heart attack, abnormal heartbeat, or history of heart attack, and, in a larger database, several broader disease groups. The authors gradually “unfreeze” the layers of the network, first training only the final layers and then allowing earlier layers to adjust. This careful schedule helps preserve the useful patterns learned from unlabeled data while adapting them to the specific task of diagnosis. A final classification module then assigns each ECG image to one of several clinically meaningful categories.
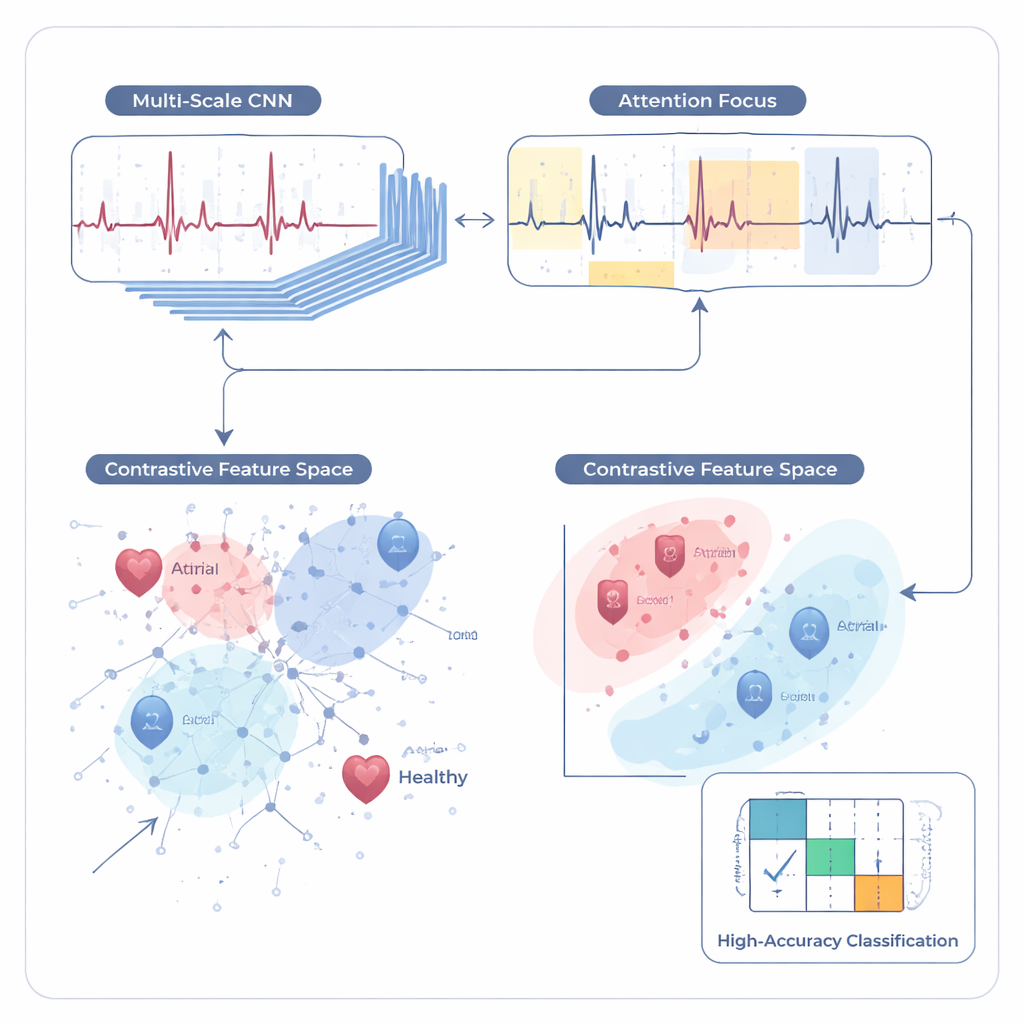
How well does it work in practice?
The team tested SimCardioNet on three separate image collections. On a four‑class dataset from Pakistani hospitals, the system correctly classified about 97.5% of ECGs, with similarly high scores for precision and recall—meaning it rarely missed disease and rarely raised false alarms. On an external Kaggle dataset it achieved perfect scores on the test split, suggesting that the features it learned transfer well to new sources, although the authors caution that such flawless numbers can sometimes reflect an easier task. On PTB‑XL, a large, widely used benchmark with five broad diagnostic groups, the model reached about 92% accuracy and F1‑score, outperforming several recent deep‑learning approaches, including specialized convolutional and recurrent networks. Visualization tools like Grad‑CAM showed that the model usually bases its decisions on clinically relevant waveform regions, such as the sharp QRS spikes and ST segments, though the authors also detected and propose fixes for occasional “shortcuts,” like focusing on page headers.
What this means for patients and clinicians
To a non‑specialist, the main message is that SimCardioNet shows how machines can be trained to interpret heart tracings accurately without demanding massive, fully labeled datasets that are expensive and slow to create. By first learning general structure from unlabeled ECG images and then refining that knowledge with a smaller labeled set, the system delivers reliable multi‑class diagnosis while remaining relatively efficient and explainable. Although more testing across hospitals, devices, and patient groups is needed before such tools can be trusted in routine care, this work suggests that automated ECG readers could one day help triage patients faster, support overburdened clinicians, and extend expert‑level cardiac assessment to regions where cardiologists are scarce.
Citation: Majid, M.D., Anwar, M., Bilal, S.F. et al. A hybrid learning framework for automated multiclass electrocardiogram classification with SimCardioNet. Sci Rep 16, 7621 (2026). https://doi.org/10.1038/s41598-026-36932-1
Keywords: electrocardiogram, deep learning, self-supervised learning, cardiovascular disease, medical imaging