Clear Sky Science · en
Generalizability and transferability of machine learning models using hyperspectral reflectance data for maize traits
Why scanning plant leaves matters for our future food
Feeding a growing population under a changing climate requires crops that can thrive in heat, drought, and other stresses. Breeders want to know which plants have the right mix of leaf structure, chemistry, and photosynthetic performance—but directly measuring these traits for thousands of plants is slow and destructive. This study explores whether simply scanning maize leaves with a hyperspectral sensor and using machine learning can reliably stand in for laborious lab measurements, even when plants are grown in different years and under shifting field conditions.
Light fingerprints of maize leaves
Every leaf reflects light in a pattern that depends on its pigments, water content, and internal structure. Hyperspectral sensors capture this pattern across hundreds of wavelengths from visible to shortwave infrared, creating a detailed "fingerprint" of each leaf. The researchers collected such fingerprints from a diverse maize population grown in three consecutive field seasons, along with 25 traits describing leaf anatomy (such as specific leaf area and carbon–nitrogen balance), gas exchange (how leaves take in CO2 and lose water), and chlorophyll fluorescence (a window into the efficiency and regulation of photosynthesis). This rich dataset allowed them to test how well different statistical models could turn light spectra into trait estimates.
Teaching machines to read leaves
The team focused on two widely used, relatively simple machine learning approaches: partial least squares regression (PLSR) and linear support vector regression (SVR). Both methods compress the highly detailed spectra into a smaller set of informative features before linking them to measured traits. The scientists carefully compared ways to tune the models, especially how many components PLSR should use, and how to avoid overfitting. They also examined whether it is better to feed models individual leaf measurements, averages from a single plot, or averages across all plants of the same genotype. A rigorous, nested cross-validation framework—essentially repeated train–test cycles—was used to check performance and uncertainty.
What traits are easiest to predict
Some leaf traits proved much more "readable" from light spectra than others. Structural and biochemical traits, such as specific leaf area and nitrogen content, were predicted with high accuracy, especially when data were averaged at the genotype level to reduce measurement noise. Certain photosynthetic capacity traits and some chlorophyll fluorescence indicators of how photosystem II behaves under light also showed moderate predictability. In contrast, traits tied to rapid, short-lived processes—like the speed at which leaves ramp up or relax protective energy dissipation—were poorly captured. For these, the spectral signal is either weak or easily swamped by environmental variation at the moment of measurement.
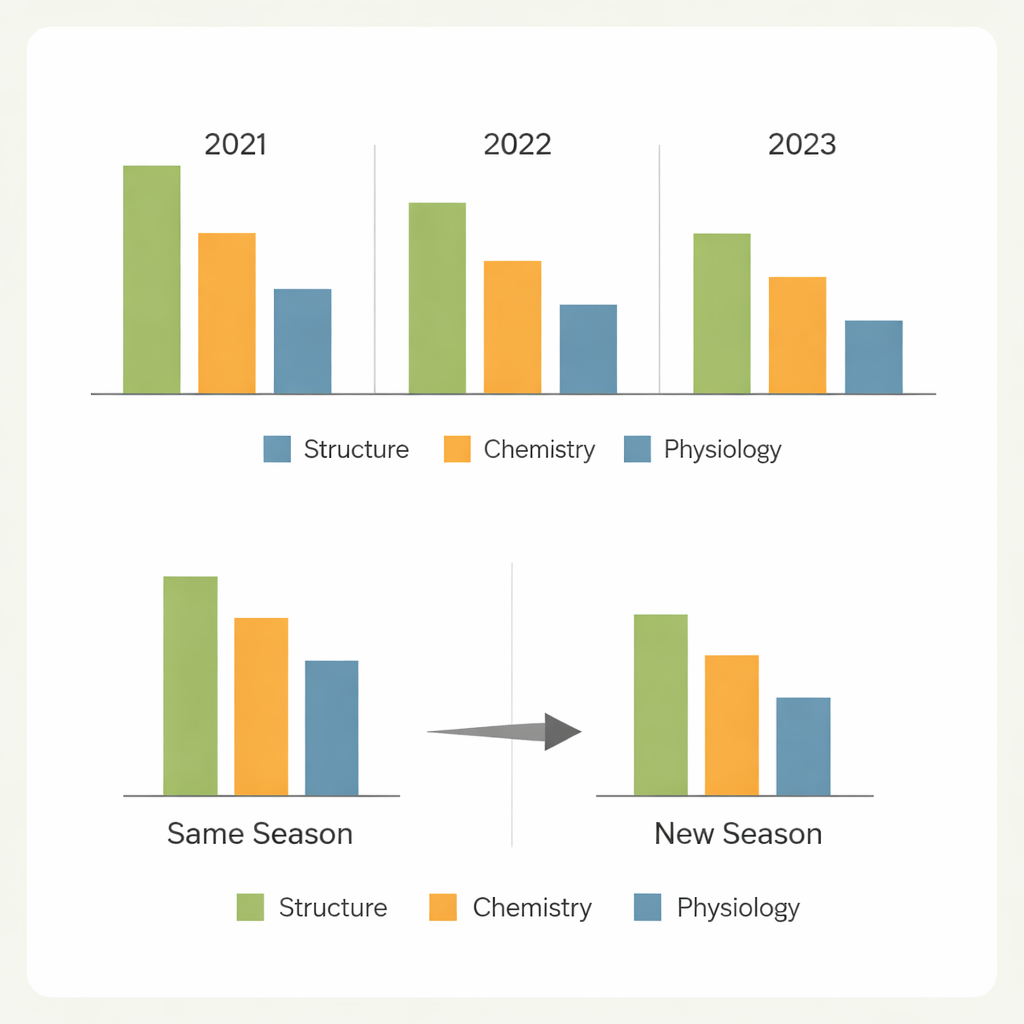
From one season to the next
A key question for real-world breeding is whether a model trained in one set of conditions can be trusted in another. When the models predicted random plants within the same season, performance was generally strong for the easier traits. Predicting entirely new genotypes grown in the same season led to only modest drops for structural and nitrogen-related traits, but much sharper declines for gas-exchange traits. The most severe test—predicting new genotypes in a different year—revealed large losses in accuracy, particularly for traits strongly shaped by the environment. Differences in weather, field conditions, and the mix of genotypes shifted the spectral patterns enough to limit transferability, with one season standing out as especially hard to predict from the others.
What this means for breeding and remote sensing
For breeders and crop scientists, the study offers both encouragement and caution. Hyperspectral scanning combined with relatively simple machine learning is already a powerful tool for high-throughput estimation of stable, integrative traits like leaf structure and nitrogen status, and can generalize across genotypes and years reasonably well for these targets. However, the same approach is far less reliable for fast, environmentally sensitive physiological traits when models are applied beyond the conditions they were trained on. The authors conclude that hyperspectral methods are ready to support large-scale screening of some key maize traits, but that predicting dynamic physiological behavior across environments will require richer training data, more advanced modeling, and perhaps additional types of measurements.
Citation: Xu, R., Ferguson, J., Breil-Aubert, M. et al. Generalizability and transferability of machine learning models using hyperspectral reflectance data for maize traits. Sci Rep 16, 5865 (2026). https://doi.org/10.1038/s41598-026-36819-1
Keywords: hyperspectral reflectance, maize, machine learning, plant phenotyping, photosynthesis