Clear Sky Science · en
Kernel mean matching enhances risk estimation under spatial distribution shifts
Why model risk under shifting maps matters
Machine-learning models are increasingly used to forecast where species will live, how tumours are organised in tissue, or how pollution spreads. Yet the data used to train these models are often collected in very specific places—dense sampling near cities, hospitals or easy-to-reach field sites—while the models are applied over much larger, different regions. This mismatch in where data come from versus where predictions are made can make models look safer and more accurate than they really are. The paper "Kernel mean matching enhances risk estimation under spatial distribution shifts" asks a deceptively simple question: when the world looks different from your training data, how wrong might your model be, and how can you tell?
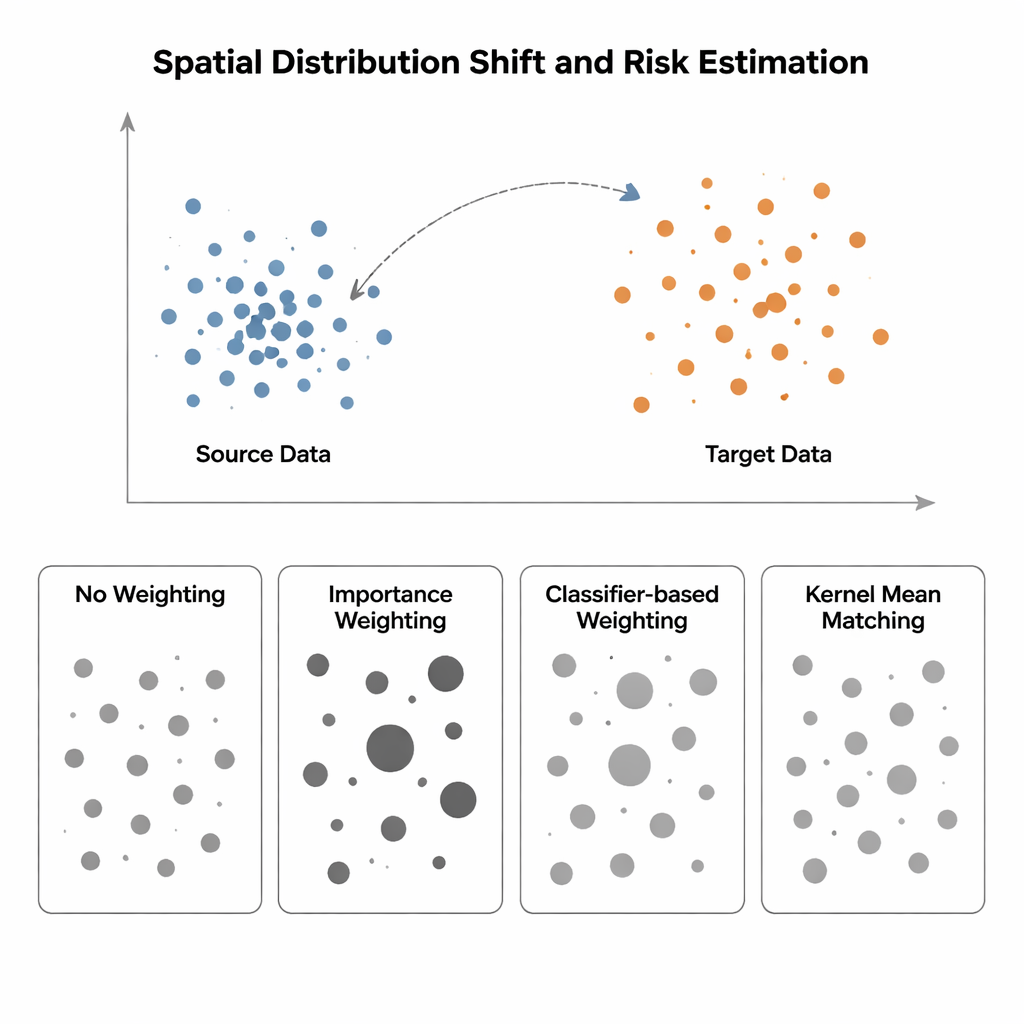
When training and testing live in different worlds
In statistics, the "risk" of a model is its expected error on new, unseen data. Standard evaluation tricks—like cross-validation or holding out a random test set—quietly assume that training and test data are drawn from the same distribution. Spatial data break this assumption. Environmental gradients, clustered sampling and changing climates mean that the conditions where we train a model can differ sharply from those where we deploy it. For example, species observations are often concentrated near roads, while conservation decisions concern remote areas; tumour samples may be taken from one part of a tissue, but predictions are needed elsewhere. In such cases, conventional estimates of risk tend to be over-optimistic, hiding how badly a model might fail in new locations.
Old tools struggle with spatial bias
The study compares four ways to estimate model risk when the input distribution shifts from a "source" region (where labels are known) to a "target" region (where labels are scarce or missing). The simplest method, called No Weighting, just measures average error on available data and assumes source and target are similar—an assumption that breaks down under spatial bias. Importance Weighting tries to fix this by scaling each source sample according to how common that kind of point is in the target relative to the source. In theory this recovers the correct risk, but in practice it requires estimating high-dimensional probability densities. When source data are tightly clustered and target data are more spread out—a typical situation in spatial ecology or medical imaging—these density estimates become unreliable, and a few samples get enormous weights, making the risk estimate wildly unstable. Classifier-based approaches, which train a classifier to distinguish source from target points and convert its probabilities into weights, avoid explicit density estimation but often produce miscalibrated risks because they optimise classification accuracy, not distribution alignment.
A different route: matching distributions directly
The authors advocate Kernel Mean Matching (KMM), an approach that bypasses density estimation altogether. Instead of trying to compute how likely each point is under source and target distributions, KMM searches for weights on source samples that make their average "signature" in a flexible kernel-defined feature space match that of the target samples. Intuitively, it stretches or shrinks the influence of each source point so that, taken together, the weighted source cloud looks like the target cloud. Once these weights are found, the risk is estimated as a weighted average of source errors. A complementary tool, the Local Correlation Function, quantifies how clustered the data are in space; it serves as a diagnostic to tell when distribution shifts are strong enough that reweighting is likely to help.
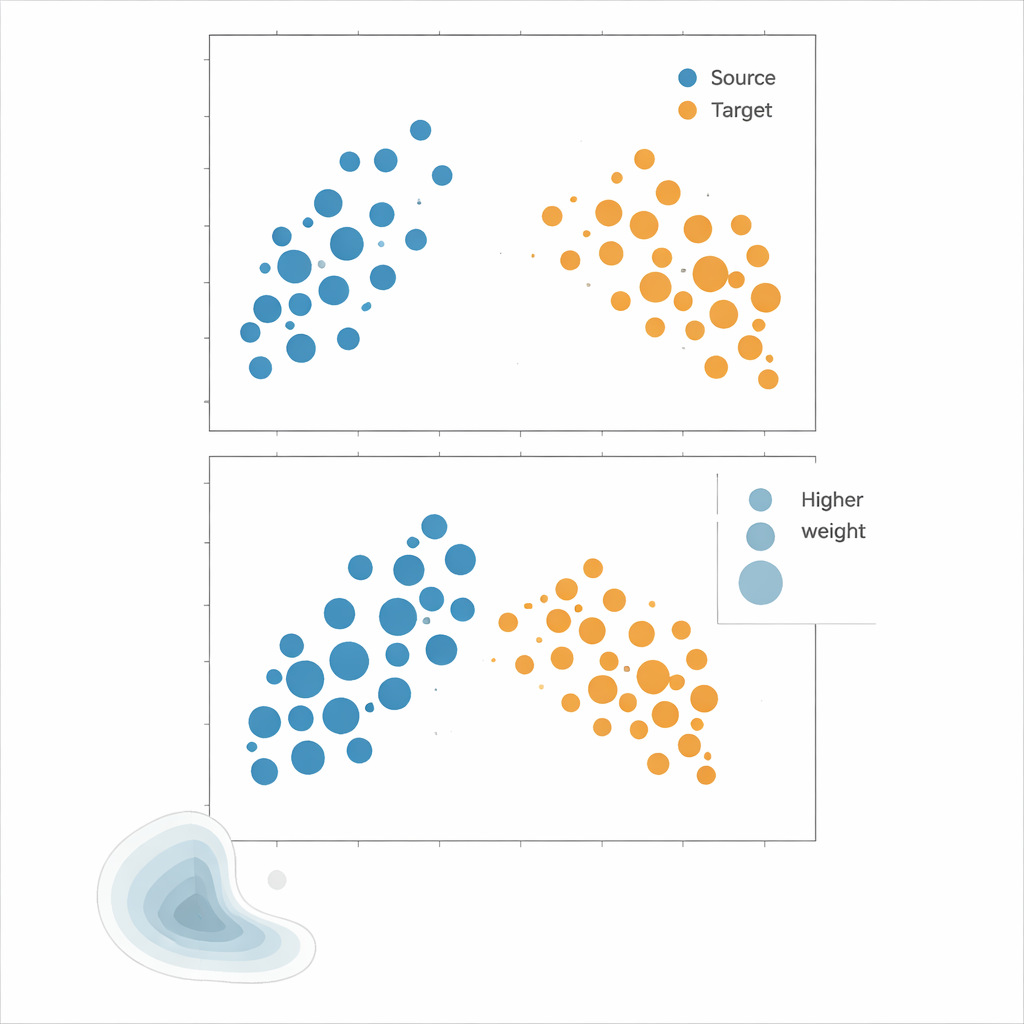
Putting the methods to the test
To see which strategy works best, the authors run extensive experiments on both synthetic and real-world data. Synthetic "landscapes" are built from mixtures of Gaussian clusters whose spread, shape and domain coverage can be precisely controlled, allowing structured tests like cropping part of the domain, changing correlation patterns between features, or switching between tightly clustered and almost uniform point patterns. Real datasets include Nordic plant species occurrences, described by climate and location, and spatial layouts of immune cells within tumours. Across these scenarios, models are trained on clustered source data and evaluated on less-clustered target data, mimicking common sampling biases. Performance is assessed using several error metrics, focusing on how closely each method’s estimated risk tracks the true error on the target.
More reliable risk in messy, high‑dimensional spaces
Across almost all synthetic setups and real datasets, KMM delivers the most accurate and stable risk estimates. It reduces mean absolute percentage error by roughly 12 to 87 percent compared with the alternatives, and crucially avoids the "weight explosion" that plagues importance weighting in high dimensions. In challenging tumour cell layouts, for example, importance weighting can yield errors exceeding several thousand percent, while KMM remains within manageable bounds. Classifier-based reweighting typically improves on naive methods but still lags behind KMM, reflecting its focus on discrimination rather than faithful distribution matching. These results suggest that for spatial applications—where data are clustered, biased and high-dimensional—KMM offers a principled way to estimate how much trust to place in a model’s predictions.
What this means for real-world decisions
For non-specialists using machine learning in ecology, environmental science or biomedicine, the message is straightforward: standard test scores can be dangerously misleading when your deployment region differs from where your data came from. Kernel Mean Matching provides a way to correct for this by rebalancing the influence of training samples so they statistically resemble the places or tissues you care about. The study shows that this approach consistently yields more honest estimates of model error, even under severe spatial bias and with many input variables. In practice, that means more reliable guidance when choosing between models and a clearer picture of where predictions are trustworthy—and where caution is warranted.
Citation: Serov, E., Koldasbayeva, D. & Zaytsev, A. Kernel mean matching enhances risk estimation under spatial distribution shifts. Sci Rep 16, 6921 (2026). https://doi.org/10.1038/s41598-026-36740-7
Keywords: distribution shift, spatial modeling, kernel mean matching, model risk estimation, ecological and biomedical data