Clear Sky Science · en
Clinical validation of lightweight CNN architectures for reliable multi-class classification of lung cancer using histopathological imaging techniques
Why this research matters for patients and doctors
Lung cancer is often deadly because it is discovered late or misclassified, which can delay the right treatment. This study explores how small, efficient computer programs—rather than huge, power-hungry ones—can reliably recognize different types of lung cancer from microscope images of tissue. If such lightweight tools work well, they could be used in hospitals around the world, including those with limited computing resources, to support pathologists in making faster and more consistent diagnoses.
Looking closely at cancer with digital microscopes
When a suspicious lung nodule is removed or biopsied, pathologists examine thinly sliced, stained tissue under a microscope to decide whether it is harmless or one of several cancer types. In this work, the authors focus on three key categories: benign lung tissue, lung adenocarcinoma, and lung squamous cell carcinoma. These subtypes matter because they respond differently to treatments. The team uses digital snapshots of these slides—histopathology images—and asks whether compact neural networks can learn the subtle visual patterns that distinguish each class, from cell shapes to tissue architecture, as reliably as much larger models.
Building smaller yet smarter digital classifiers
Most state-of-the-art image recognition systems are extremely large and demand expensive graphics processors, making them hard to deploy in many clinics. The researchers instead design four “lite” image-analysis models, called Lite-V0, Lite-V1, Lite-V2, and Lite-V4, each a streamlined version of a convolutional neural network (CNN). All four follow the same basic recipe: they gradually extract visual features through a stack of simple building blocks, then summarize the image and output one of the three lung tissue labels. What changes from version to version is how many blocks are used and how wide they are—essentially, how much capacity the model has to learn complex patterns. This controlled design lets the team study how much complexity is truly needed for reliable cancer classification.
Training, testing, and choosing the fairest model
To teach and test these models, the authors assemble a balanced collection of 15,000 lung tissue images, carefully split into training, validation, and test groups with equal numbers from each class. Before training, each image is resized, normalized, and gently augmented with flips, small rotations, and zooms to mimic how slides can appear under different conditions. Crucially, the team does not judge models on raw accuracy alone, because that metric can hide poor performance on one class. Instead, they rely on a “macro-F1” score, which forces the model to do well on all three tissue types, not just the easiest ones. A custom training procedure continuously monitors this balanced score and automatically stops training when improvements level off, saving the best version of each model for comparison. 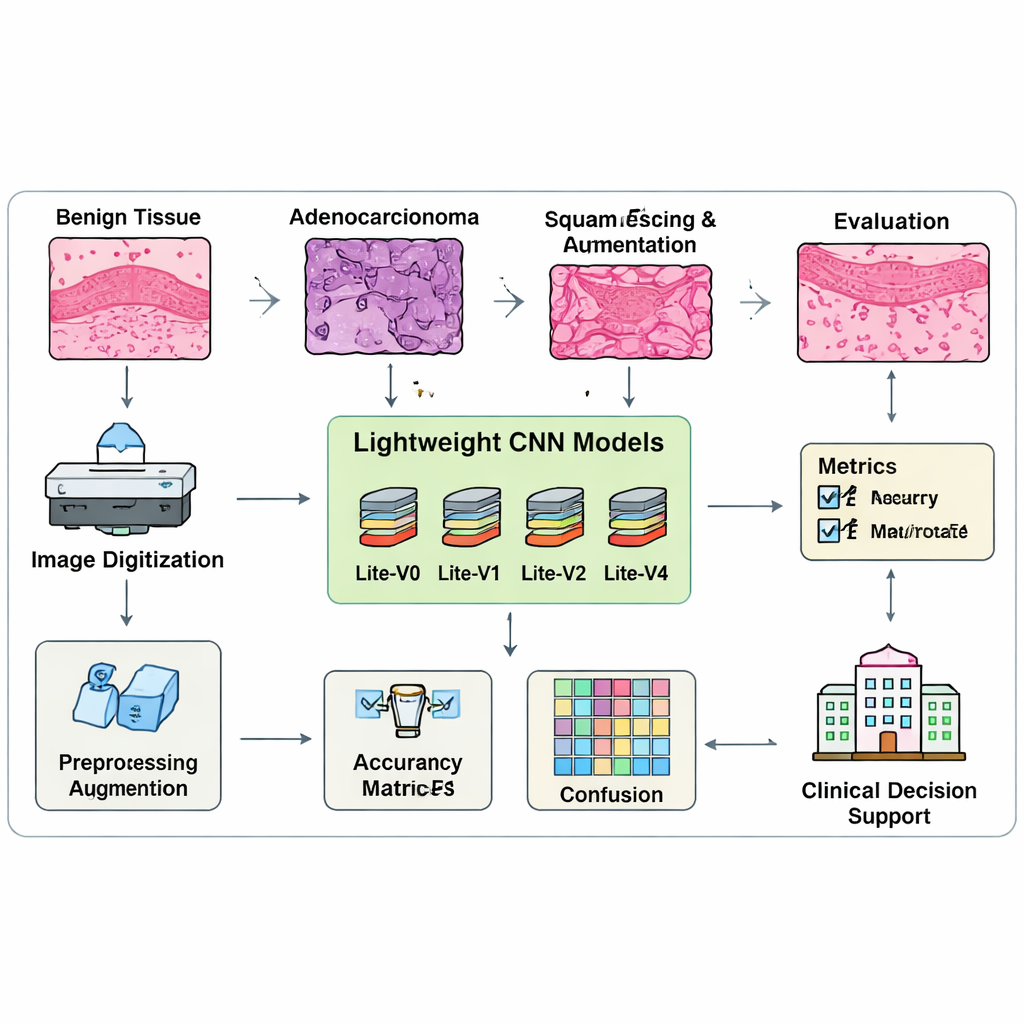
What the best lightweight model can really do
When the dust settles, one variant—Lite-V2—stands out. It is not the smallest network, nor the largest, but lands in the middle and achieves the best balance between accuracy and efficiency. On unseen test images, Lite-V2 correctly classifies benign tissue, adenocarcinoma, and squamous cell carcinoma with a high and evenly distributed performance, reaching a macro-F1 score of about 0.96. Confusion-matrix charts show that it rarely mixes up the three categories, while deeper versions begin to “overfit,” memorizing the training data but losing reliability on new cases. The authors further rerun Lite-V2 multiple times with different random starting points and use a statistical test to confirm that its advantage over the other variants is not a fluke. 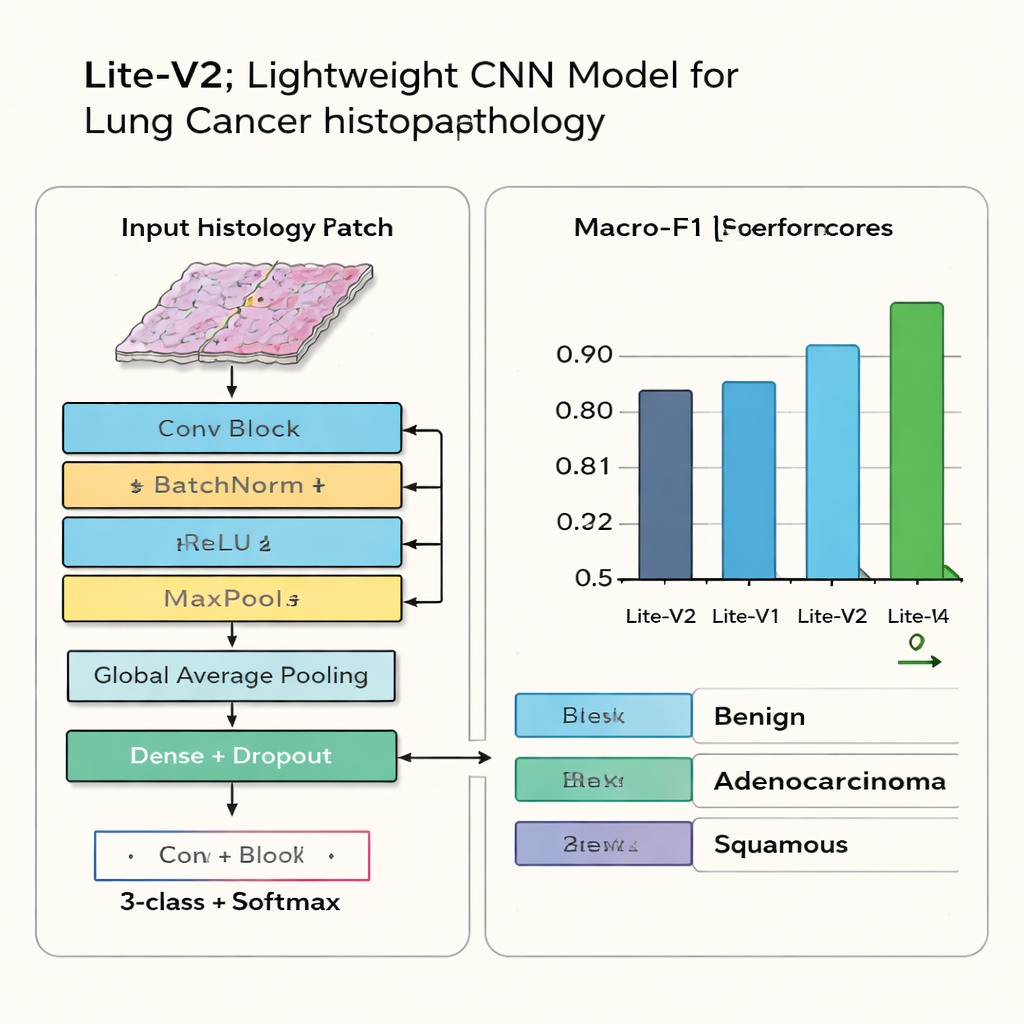
From research code to real-world support
Beyond performance numbers, the study emphasizes practical deployment. Because Lite-V2 and its siblings are compact, they can run on modest hospital hardware or even edge devices without offloading sensitive images to the cloud. The authors publish a reproducible framework that logs every experimental detail, from data processing to training curves and error patterns, so other teams can verify or extend the work. For patients and clinicians, the key takeaway is that thoughtfully designed lightweight AI can help bring reliable lung cancer classification closer to everyday pathology practice, supporting faster, more consistent decisions—even in clinics that lack cutting-edge computing power.
Citation: Raza, A., Hanif, F. & Mohammed, H.A. Clinical validation of lightweight CNN architectures for reliable multi-class classification of lung cancer using histopathological imaging techniques. Sci Rep 16, 6512 (2026). https://doi.org/10.1038/s41598-026-36652-6
Keywords: lung cancer, histopathology, convolutional neural networks, medical imaging AI, computer-aided diagnosis