Clear Sky Science · en
Diagnosis of Alzheimer’s disease with high accuracy via Petri net modeling of signaling pathways
Why early brain disease detection matters to all of us
Alzheimer’s disease slowly erodes memory and independence, often long before clear symptoms appear. Today’s diagnostic tools, like brain scans and cognitive tests, can be expensive, hard to access, and not very sensitive in the earliest stages. This study introduces a new way to read the body’s molecular “wiring” using blood or brain samples and a mathematical framework called a Petri net. The goal is simple but profound: to tell, with very high accuracy, whether someone’s gene activity pattern looks more like that of a healthy person or a person with Alzheimer’s disease—ideally early enough to change the course of the illness.
Reading disease in the body’s signaling “circuits”
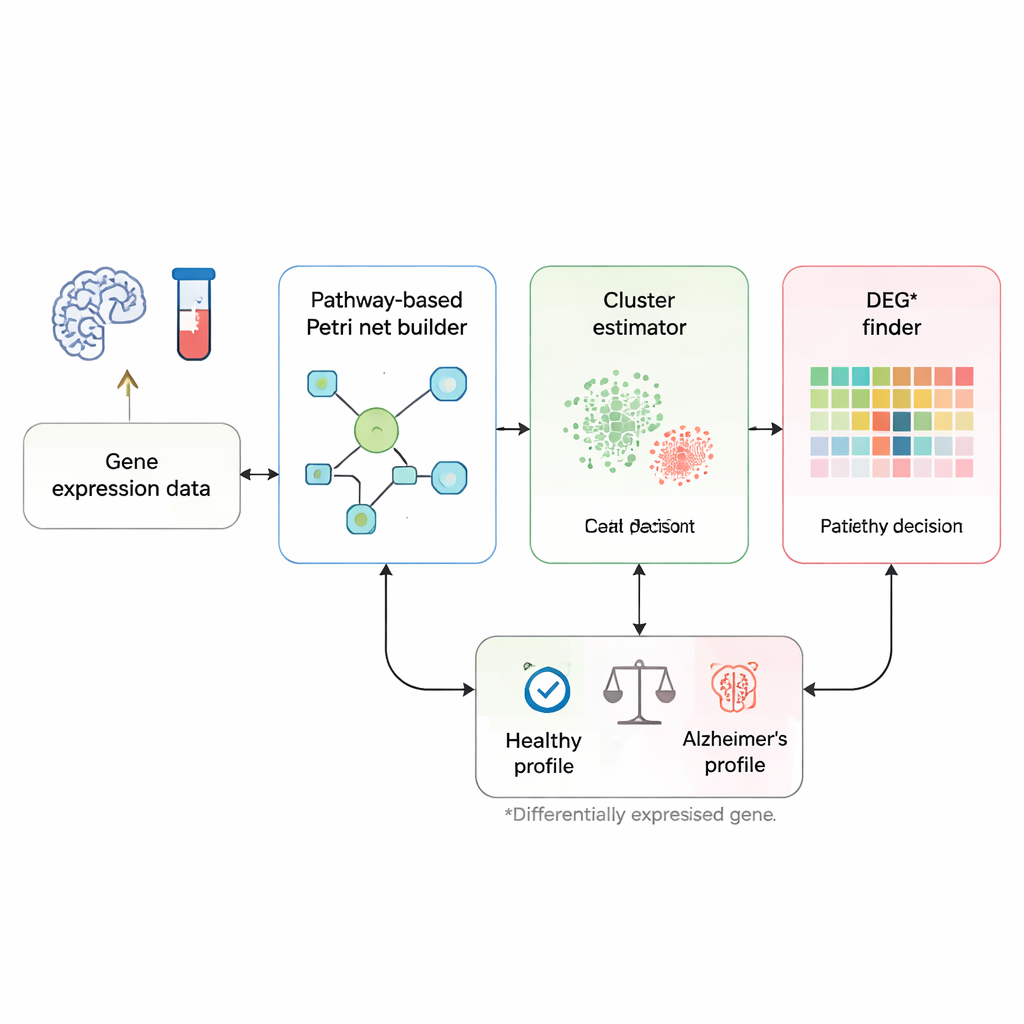
Instead of treating each gene as an isolated marker, the researchers focus on how genes talk to each other along known biological signaling pathways linked to Alzheimer’s disease. They start from a curated pathway map from the KEGG database, which lists genes and the activating or blocking relationships between them. This pathway is turned into a Petri net—a kind of flow diagram where circles stand for genes, rectangles for interactions, and small tokens for signals moving through the network. When a token travels from one end of the pathway to special “disease” nodes, it represents a chain of gene events that could lead to brain cell death and dementia.
From a blood sample to a yes-or-no answer
The method begins with gene expression profiles: measurements that indicate how active thousands of genes are in a person’s blood or brain tissue. First, a custom “cluster estimator” compares a new sample with previously labeled healthy and patient samples, focusing on genes already known to differ between these groups. For each such gene, the tool checks which training samples have the most similar expression levels and uses a kind of neighborhood voting to give the new sample an initial label as likely healthy or likely diseased. This step forms two working groups, or clusters, of samples: one representing typical healthy patterns and one representing typical Alzheimer’s patterns.
Finding the most telling genes for each person
Next comes a more personalized step. For a given person, the method looks gene by gene to see where their expression levels truly stand out. For each gene, it compares that person’s value to the range seen in both the healthy cluster and the patient cluster, using robust statistics that downplay the influence of noisy outliers. If a gene’s expression clearly sits inside the “diseased” range and outside the healthy range, it is flagged as a DEG*, a particularly informative gene for that individual. These DEG* genes receive initial tokens in the Petri net, marking where abnormal activity first appears in the signaling circuit.
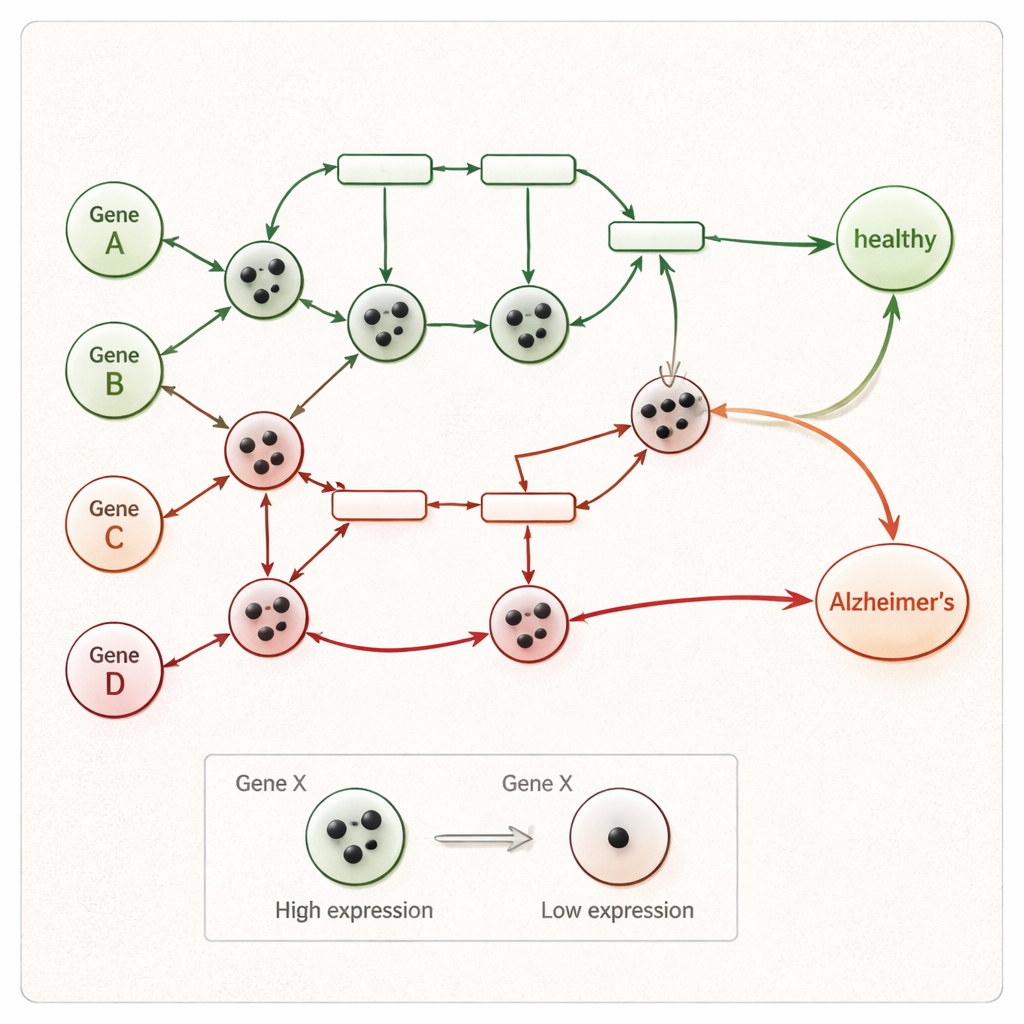
Letting the network play out to a diagnosis
Once the tokens are placed, the Petri net runs step by step. At each step, every interaction whose input genes have enough tokens “fires,” moving tokens to downstream genes while respecting activation and inhibition rules. This continues until no further moves are possible. If, at the end, at least one token reaches the network’s final disease node—representing processes like neuron death or neurodegeneration—the sample is classified as Alzheimer’s; if not, it is labeled healthy. Because every firing step is recorded, researchers can trace a clear path from a person’s unusual gene activity all the way to a healthy or diseased outcome, rather than relying on an opaque statistical score.
How accurate is this approach in practice?
The team tested their framework on several large, public gene expression datasets from both blood and brain tissue. On two widely used blood datasets, their method correctly distinguished Alzheimer’s patients from healthy individuals in about 98–99% of cases, far surpassing earlier machine learning approaches that often reached only 65–81% accuracy. They also showed strong performance across additional brain and blood datasets, even when the number of samples was small. Importantly, unlike many competitors, their method did not discard noisy or hard-to-interpret samples; instead, it uses its internal design to handle variability while keeping all data in the analysis.
What this means for future Alzheimer’s testing
To a layperson, the main message is that the study turns complex genetics and pathway diagrams into a clear, stepwise decision process that can read early signs of Alzheimer’s disease from a simple sample, potentially even just blood. By modeling how abnormal genes work together, not just how they behave alone, the Petri net framework provides both high diagnostic accuracy and an understandable “story” of how disease may unfold in each person. While this work is still research and not yet a clinical test, it points toward future tools that could detect Alzheimer’s earlier, guide treatment choices, and eventually be adapted to other brain diseases and cancers using the same underlying idea.
Citation: Ebrahimian, H., Asadzadeh, F., Rahgozar, M. et al. Diagnosis of Alzheimer’s disease with high accuracy via Petri net modeling of signaling pathways. Sci Rep 16, 6457 (2026). https://doi.org/10.1038/s41598-026-36585-0
Keywords: Alzheimer’s diagnosis, gene expression, signaling pathways, Petri net modeling, blood biomarkers