Clear Sky Science · en
Four novel Planctomicrobium species isolated from sewage sludge or leakage water of a compost heap in Northern Germany
Strange bacteria in everyday waste
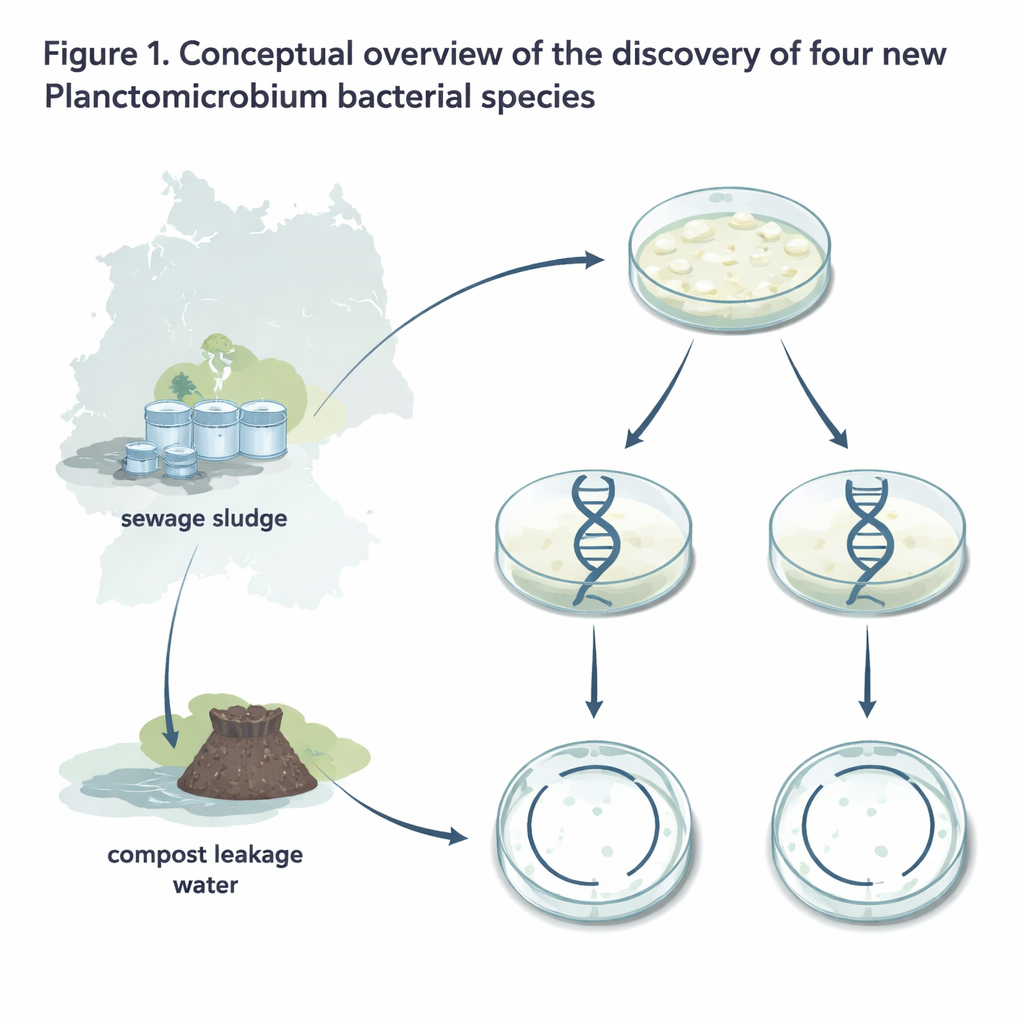
Household sewage and steaming compost heaps might not sound like frontiers of discovery, yet they are teeming with microscopic life that quietly recycles our waste. In this study, researchers dug into old bacterial samples from sewage sludge and compost leakage water in Northern Germany and uncovered four previously unknown species of unusual bacteria. These tiny organisms, part of a group called Planctomicrobium, help break down complex plant materials and could one day inspire new biotechnologies and medicines.
A quirky branch of the bacterial tree
The new species belong to a larger phylum of bacteria known as Planctomycetota. Members of this group are oddities in the microbial world: they have intricate internal membranes, divide by budding rather than splitting in two, and lack some of the standard cell-division machinery found in most bacteria. Planctomycetota live in oceans, lakes, soils, and on the surfaces of algae, seagrasses, and sponges, where they help drive key cycles of carbon and nitrogen by breaking down complex carbohydrates. Scientists have only begun to explore their diversity, and many more lineages likely remain to be described.
Four new neighbours in the waste stream
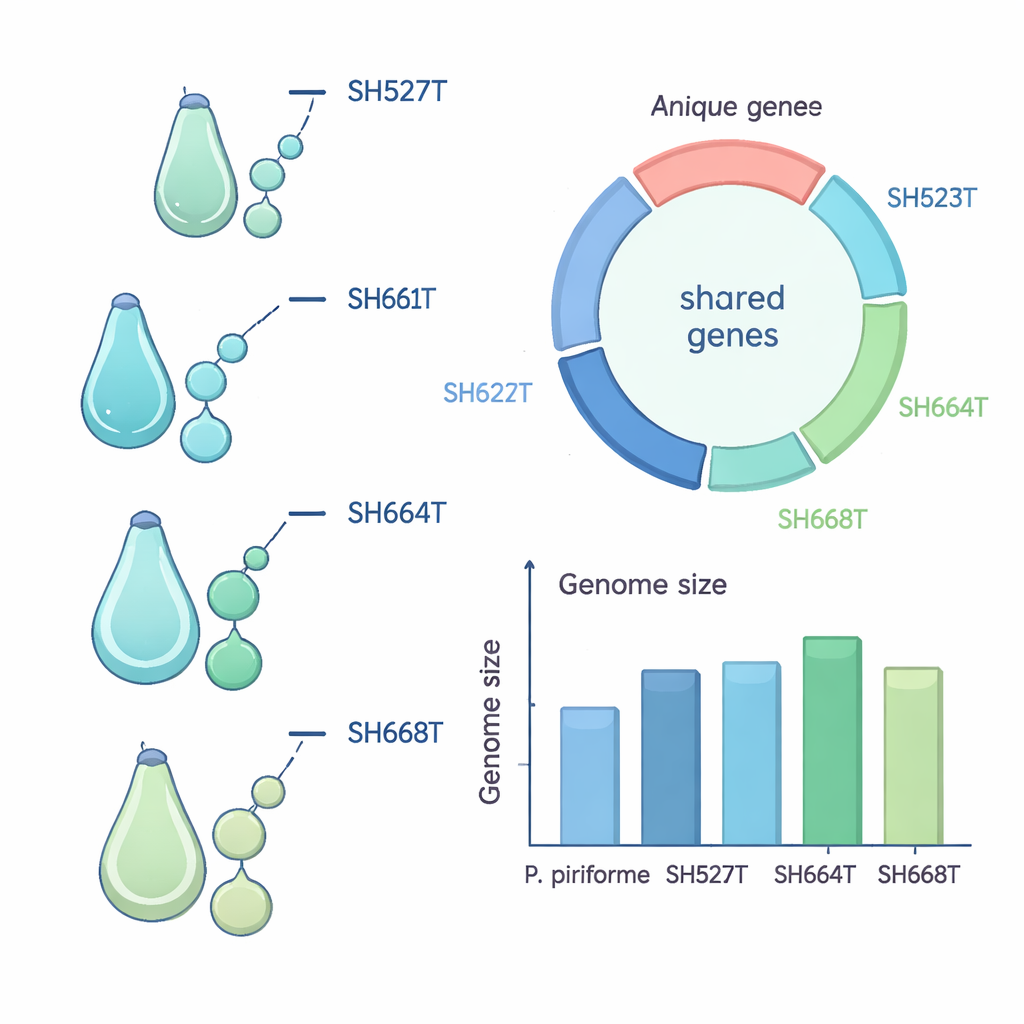
The four strains described here—named Planctomicrobium limosum, P. stercoris, P. aquicomposti, and P. mucosum—were originally isolated decades ago by microbiologist Heinz Schlesner and later revived for modern analysis. All four were collected from either sewage treatment sludge or the watery runoff from industrial compost heaps in Northern Germany. On laboratory plates, they form off‑white, ivory-colored colonies; two of the species make especially large, slimy patches. Under the microscope, their cells are oval to pear-shaped and reproduce by growing a small “bud” on one end that eventually pinches off as a motile daughter cell. They grow best at room-like temperatures and near‑neutral pH, and they all require oxygen and organic nutrients, which fits their lifestyle in oxygenated, nutrient-rich waste environments.
Reading and comparing their genomes
To understand how these bacteria relate to known species, the team sequenced and carefully assembled their entire genomes, each packaged as a single circular chromosome without extra plasmids. They then compared several genetic markers, including the standard 16S rRNA gene and whole-genome similarity measures, against the only previously described Planctomicrobium species, P. piriforme from a Russian peat bog. The new strains were clearly close relatives at the genus level but fell below accepted cutoffs used to define a single species, meaning each represents its own species. The genomes are also noticeably more compact: one, P. mucosum, has the smallest genome yet reported for this family, making it a useful candidate for future studies that want to home in on essential genes.
What these microbes are built to eat
By scanning the genomes for specific enzyme families, the researchers inferred what kinds of food these bacteria are best equipped to use. All five Planctomicrobium strains carry many genes for carbohydrate-active enzymes that degrade complex sugars, reinforcing the idea that they specialize in chewing up polysaccharides found in decaying plant matter and biofilms. In contrast, they largely lack the enzyme sets needed to dismantle tougher aromatic building blocks from lignin, the woody component of plants. The genomes also harbor several biosynthetic gene clusters predicted to make terpenes, polyketides, and small peptide-based molecules—types of compounds that in other bacteria often turn out to be antibiotics or signaling chemicals—highlighting their potential as a source of new natural products.
Why naming new bacteria matters
By combining careful microscopy with detailed genome comparisons, the authors show that these four strains are distinct from each other and from the previously known P. piriforme, justifying their recognition as four new species within the genus Planctomicrobium. Beyond expanding the bacterial family tree, this work sharpens our picture of how specialized microbes in sewage and compost help recycle complex sugars while ignoring other plant components. It also enriches a growing collection of Planctomycetota strains whose unusual biology and hidden chemical talents may eventually be harnessed for environmental cleanup, green chemistry, or new drugs.
Citation: Kallscheuer, N., Kumar, G., Hammer, J. et al. Four novel Planctomicrobium species isolated from sewage sludge or leakage water of a compost heap in Northern Germany. Sci Rep 16, 4347 (2026). https://doi.org/10.1038/s41598-026-36544-9
Keywords: Planctomicrobium, wastewater bacteria, compost microbiome, bacterial genomics, polysaccharide degradation