Clear Sky Science · en
Full-Length transcriptome assembly and SSR marker development for Spinibarbus hollandi using PacBio SMRT sequencing
Why a humble river fish matters
Along the rivers of southern China swims Spinibarbus hollandi, a carp-like fish prized both for the dinner table and as an ornamental species. Farmers would like to raise it more efficiently, but the fish grows slowly, matures late, and produces relatively few eggs. The study described here uses cutting‑edge DNA and RNA sequencing tools to build a detailed genetic “parts list” for this species. That genetic information can ultimately help breeders select hardier, faster‑growing fish and support conservation of wild populations.
Turning many tissues into a genetic map
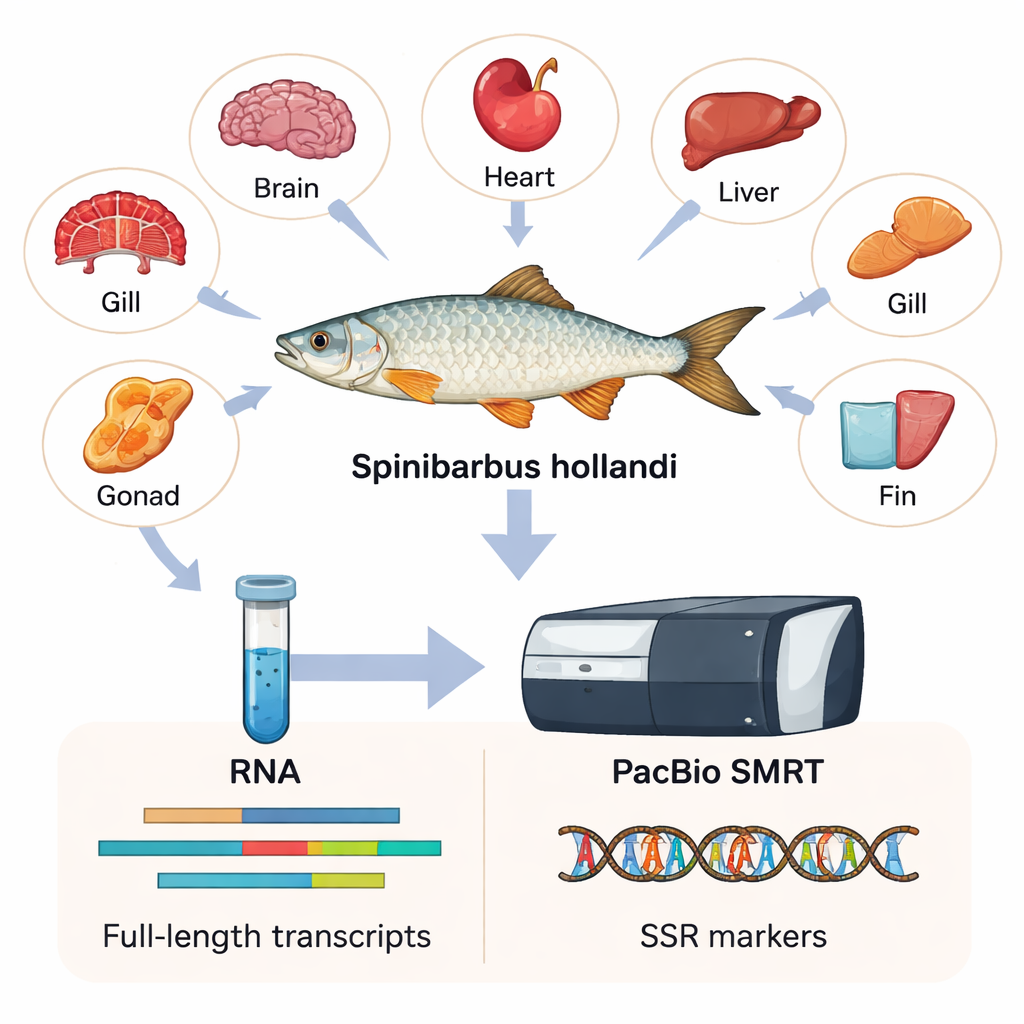
To capture as much genetic information as possible, the researchers collected six different tissues—heart, gill, brain, fin, liver, and gonad—from six adult males and females. From these tissues they extracted RNA, the molecule that carries copies of active genes inside cells. Using a technology called PacBio SMRT, which reads very long stretches of RNA copies, they assembled the first full-length transcriptome for S. hollandi. In simple terms, they built a high‑resolution catalog of 15,197 distinct genes and 23,403 transcript sequences, most of them much longer and more complete than what older, short‑read methods could produce.
What the genes reveal about the fish
Next, the team asked what these genes actually do. They compared each sequence with large public databases that group genes by function. More than 95% of the genes could be matched to known entries, a very high success rate that signals good data quality. Many genes fell into categories related to basic cell activities such as metabolism, signal transmission, and development, and most closely resembled genes from other carp family fishes. This confirms that the new transcriptome is biologically sound and provides a foundation for finding genes tied to traits like growth rate, stress tolerance, and reproduction in S. hollandi.
Hidden layers of control in RNA
Beyond identifying genes, the scientists also looked at how those genes are controlled. They found 373 cases of alternative splicing, where the same gene is cut and pasted in different ways to make distinct RNA messages. The most common pattern kept pieces of RNA that are often removed in other species, hinting at a special way this fish fine‑tunes its proteins. They also discovered 2,397 long non‑coding RNAs—RNA molecules that do not make proteins but can regulate when and where other genes turn on. Together, these findings show that S. hollandi uses a rich set of RNA‑level controls that may influence growth, sexual maturation, and adaptation to local environments.
Building DNA landmarks for breeding and conservation
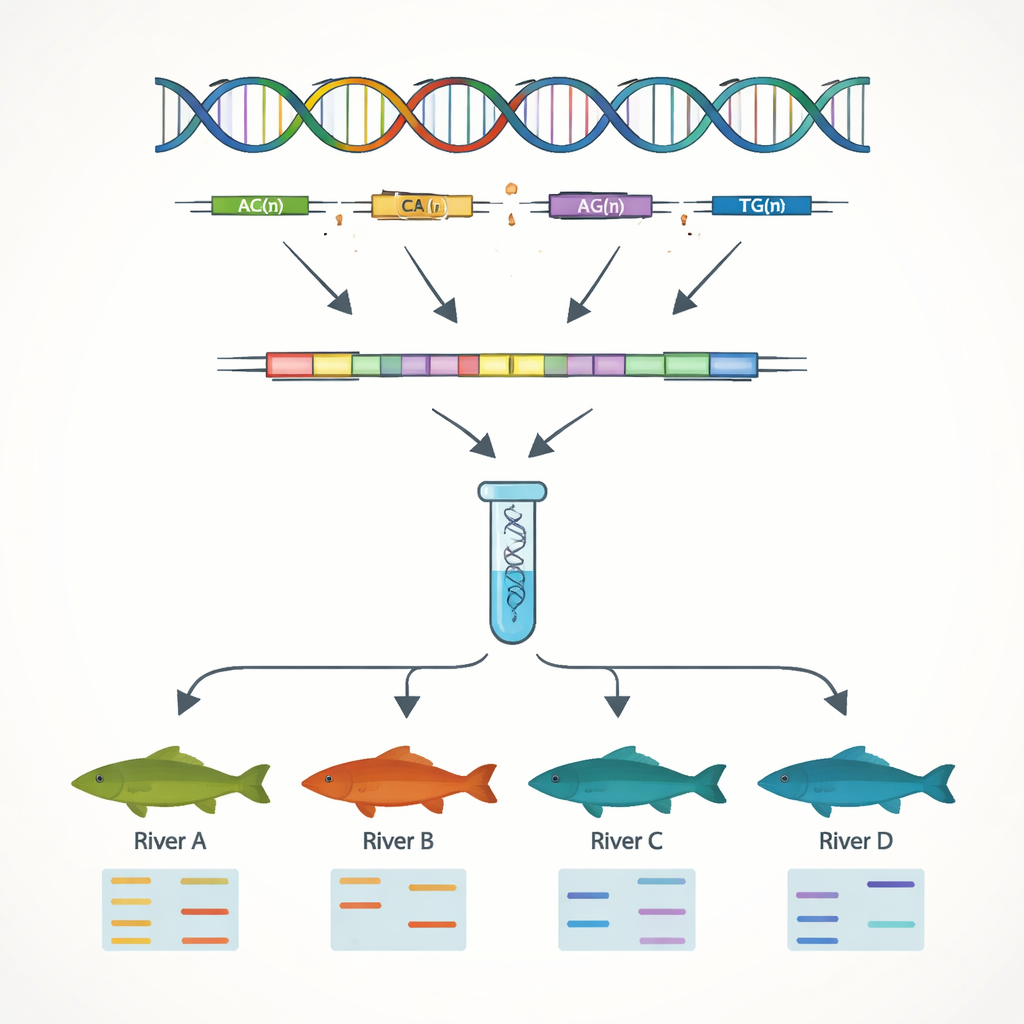
A major goal of the work was to develop simple DNA markers that can tell individual fish and populations apart. Focusing on the gene sequences, the authors searched for simple sequence repeats, or SSRs—short DNA motifs like “AC” or “AAT” repeated many times in a row. These stretches tend to vary between individuals, making them useful as genetic barcodes. They uncovered 7,449 such SSR sites and designed dozens of short DNA primers to amplify them in the lab. Thirteen of these markers turned out to be highly variable and reliable when tested in 51 fish from four river systems. Using just these 13 markers, the team could clearly detect genetic differences between populations from the Yangtze River and those from the Pearl River, reflecting the mountain ranges that limit natural mixing between basins.
What this means for fish farmers and rivers
For non‑specialists, the key takeaway is that the authors have created a detailed, high‑quality genetic reference for S. hollandi and a set of practical DNA markers. This toolkit will help researchers pinpoint genes behind slow growth or low fertility, assist breeders in selecting broodstock with desirable traits, and allow conservationists to track genetic diversity across river systems. While more work is needed to link specific genes to performance in ponds or in the wild, this study lays the molecular groundwork for turning a traditional river fish into a modern, sustainably managed aquaculture species.
Citation: Li, S., Lai, J., Wu, M. et al. Full-Length transcriptome assembly and SSR marker development for Spinibarbus hollandi using PacBio SMRT sequencing. Sci Rep 16, 5629 (2026). https://doi.org/10.1038/s41598-026-36468-4
Keywords: Spinibarbus hollandi, fish transcriptome, PacBio sequencing, microsatellite markers, aquaculture genetics