Clear Sky Science · en
Artificial neural network modeling and optimization of an electrochemical biosensor for plasma miR-155-based breast cancer detection
Why this matters for early cancer checks
Doctors and researchers are searching for simple blood tests that can spot cancer long before symptoms appear. One promising sign in the blood is a tiny genetic fragment called miR-155, which is linked to breast cancer and several immune and inflammatory conditions. Building a sensor that can reliably measure such a faint signal is tricky and usually demands months of trial-and-error in the lab. This study shows how pairing smart computer models with an electrochemical biosensor can make that process faster, cheaper, and more effective, bringing practical early-detection tests a step closer.
Turning a blood drop into a warning signal
The work centers on an electrochemical biosensor that reads out levels of miR-155 in plasma, the liquid part of blood. The heart of the device is a tiny gold electrode whose surface is carefully prepared in several stages. First, short DNA strands designed to recognize miR-155 are attached to the metal. Then a small molecule fills the gaps to keep stray substances from sticking. When a patient sample is added, any miR-155 present latches onto the DNA strands, and a dye-like molecule called Oracet Blue wedges into these paired structures. Finally, an electrical scan measures how much charge is transferred, producing a current whose size reflects how much miR-155 is in the sample. 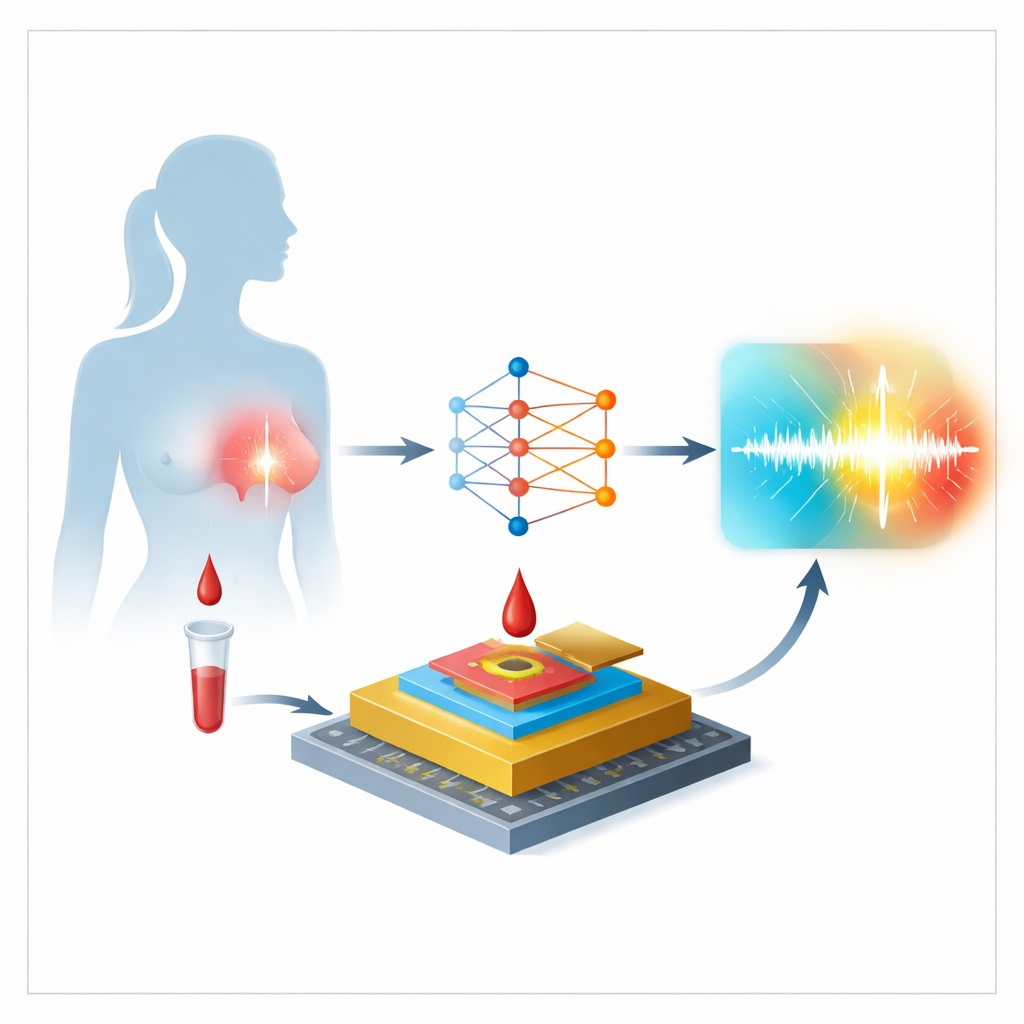
Why traditional tuning is slow and wasteful
Although the basic sensing idea is straightforward, the performance of the biosensor strongly depends on how it is built. Six key steps matter: how concentrated the DNA probe is, how long it is allowed to attach, how long the blocking molecule sits, how long the target miRNA is given to bind, and both the amount and the waiting time for Oracet Blue. In earlier work, researchers would tweak one or two of these at a time, run fresh experiments, and slowly inch toward a better design. That approach consumes expensive materials, takes a great deal of researcher time, and can miss the best combinations, especially when the underlying behavior is highly nonlinear—small changes in timing or concentration can suddenly double or halve the signal.
Letting computers learn the best build recipe
To break this bottleneck, the authors turned to machine learning, using two modeling styles: artificial neural networks (ANN) and an adaptive neuro-fuzzy system (ANFIS). They compiled data from 51 different ways of fabricating the same miR-155 sensor, each defined by a unique mix of the six key steps, and recorded the resulting electrical current. The ANN learned to map fabrication choices directly to sensor output using a compact network with one hidden layer and 13 internal units. The ANFIS approach combined fuzzy rules (such as "low" or "high" levels) with neural-style training and also used a statistical trick to reduce overlap among inputs. After careful training and testing, the ANN turned out to be more accurate and easier to handle, capturing the complex interactions between steps better than the more rule-heavy fuzzy model.
Searching the design space with digital evolution
Once the ANN could faithfully mimic the biosensor, the team linked it to a genetic algorithm, a search method inspired by evolution. This digital optimizer started with many random recipes for building the sensor and repeatedly "bred" and mutated them, keeping those that the ANN predicted would give stronger signals. Within this virtual environment, no new lab work was needed: the computer could explore countless possibilities in silico. The winning recipe was striking. Compared with the best conditions the researchers had actually tested at the bench, the optimized combination used less DNA probe and less Oracet Blue overall, shortened several incubation steps, and yet was predicted to more than double the sensor’s output current—from 98 to 223 nanoamperes—making the signal stronger and easier to distinguish from background noise. 
What this means for future blood tests
For a non-specialist, the key message is that the authors have turned biosensor tuning from guesswork into a guided, data-driven process. By letting a neural network learn how fabrication steps influence the final electrical signal, and then letting an evolutionary algorithm hunt for the best recipe, they found conditions that should deliver a brighter, more reliable readout of miR-155 while wasting less time and material. Although the optimized settings still need full experimental confirmation and the study focuses on a single type of breast cancer marker, the same strategy can be applied to many other biosensors. In the long run, this kind of smart design could help bring fast, inexpensive blood tests for early cancer and other diseases into routine clinical use.
Citation: Imani, A., Hosseinpour, S., Azimzadeh, M. et al. Artificial neural network modeling and optimization of an electrochemical biosensor for plasma miR-155-based breast cancer detection. Sci Rep 16, 7893 (2026). https://doi.org/10.1038/s41598-026-36466-6
Keywords: breast cancer biosensor, microRNA-155 detection, electrochemical sensor, artificial neural networks, genetic algorithm optimization