Clear Sky Science · en
An integrative approach to identify novel miRNA-mRNA interaction networks in LMNA-cardiomyopathy
Why tiny heart regulators matter
Dilated cardiomyopathy is a serious condition in which the heart becomes enlarged and weak, often leading to heart failure, dangerous heart rhythms, and even sudden death. For many families, this disease is inherited, and changes in a gene called LMNA are a frequent culprit. This study asks a deceptively simple question with big consequences: in hearts damaged by LMNA mutations, which genes are switched on or off, and which tiny RNA molecules are pulling those switches? By mapping these molecular conversations in detail, the work points toward new ways to predict, monitor, and perhaps one day treat this aggressive form of heart disease.
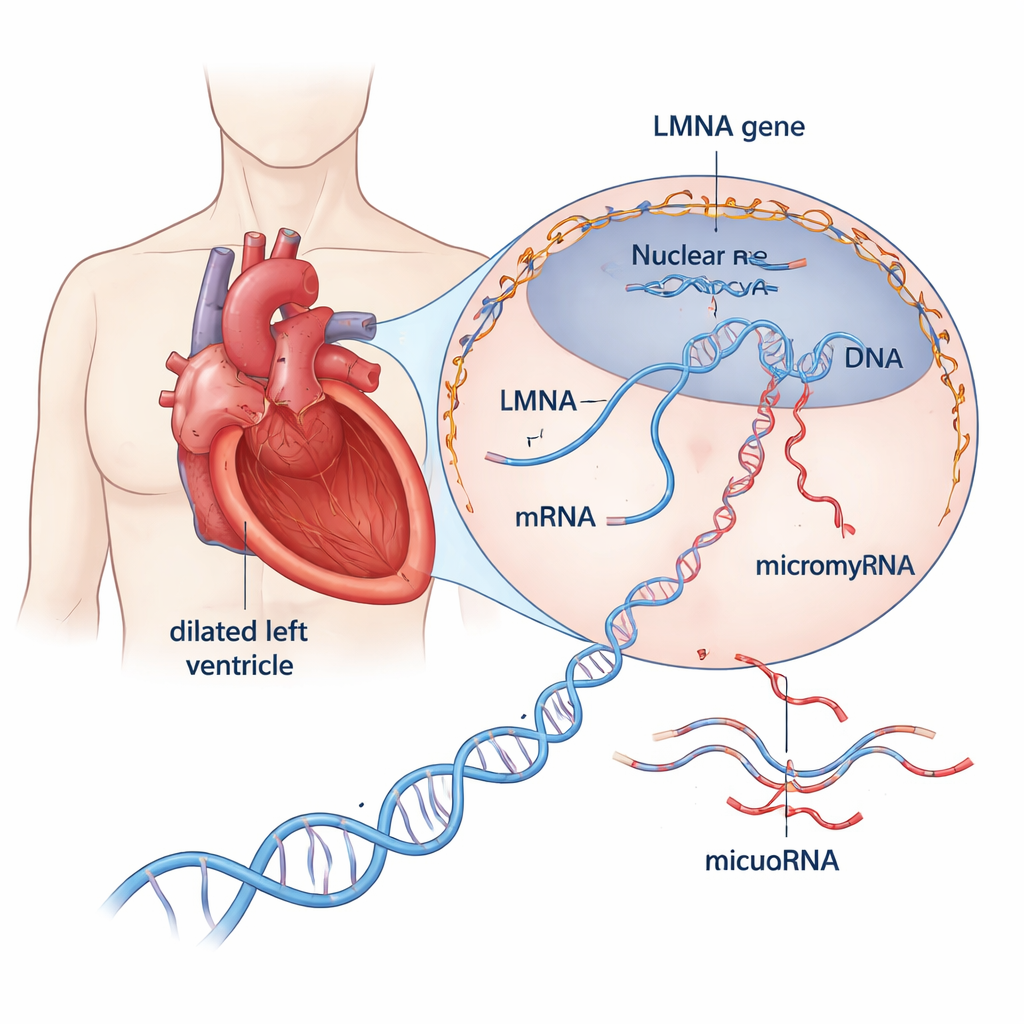
A closer look at a dangerous heart disease
Dilated cardiomyopathy stretches and thins the main pumping chamber of the heart, weakening its ability to push blood through the body. About one third to one half of cases once called “idiopathic” now turn out to have a genetic basis. Among the more than 30 genes involved, LMNA stands out: harmful variants in this gene account for roughly 10% of inherited cases and are linked to rapid disease progression, early rhythm problems, and a high risk of sudden cardiac death. LMNA makes structural proteins that form a scaffold around the cell’s DNA, helping maintain nuclear shape and influencing which genes are active. Yet how faulty LMNA proteins drive hearts toward failure has remained poorly understood.
Following the messages inside mouse hearts
To explore this, the researchers used a mouse model carrying a well-known LMNA mutation (R249W) that develops a form of dilated cardiomyopathy similar to the human disease. At 50 weeks of age—when the mice show enlarged ventricles, reduced pumping function, and fibrotic scarring—the team extracted heart tissue and examined two types of RNA: messenger RNA (mRNA), which carries instructions for making proteins, and microRNA (miRNA), short non-coding strands that fine-tune gene activity by blocking or degrading specific mRNAs. Using high-throughput sequencing and rigorous statistical filters, they identified 2,148 genes whose activity changed in mutant hearts and 53 miRNAs that were either increased or decreased compared with healthy mice.
What changes inside diseased heart cells
When the researchers grouped the altered genes into biological pathways, several themes emerged. Many of the changed genes were linked to how heart muscle cells contract, how they stick to each other and to their surrounding support matrix, how electrical signals travel through the heart, and how cells process fats to generate energy. These findings fit well with the clinical picture of LMNA-related disease: stiff, fibrotic tissue; disturbed electrical conduction that favors arrhythmias; and impaired energy balance. Pathway analyses using established databases confirmed enrichment in extracellular matrix remodeling, voltage-gated ion channels, synaptic-like communication in the heart, and fatty acid metabolism—suggesting that the LMNA mutation disrupts multiple, interconnected systems rather than a single faulty step.
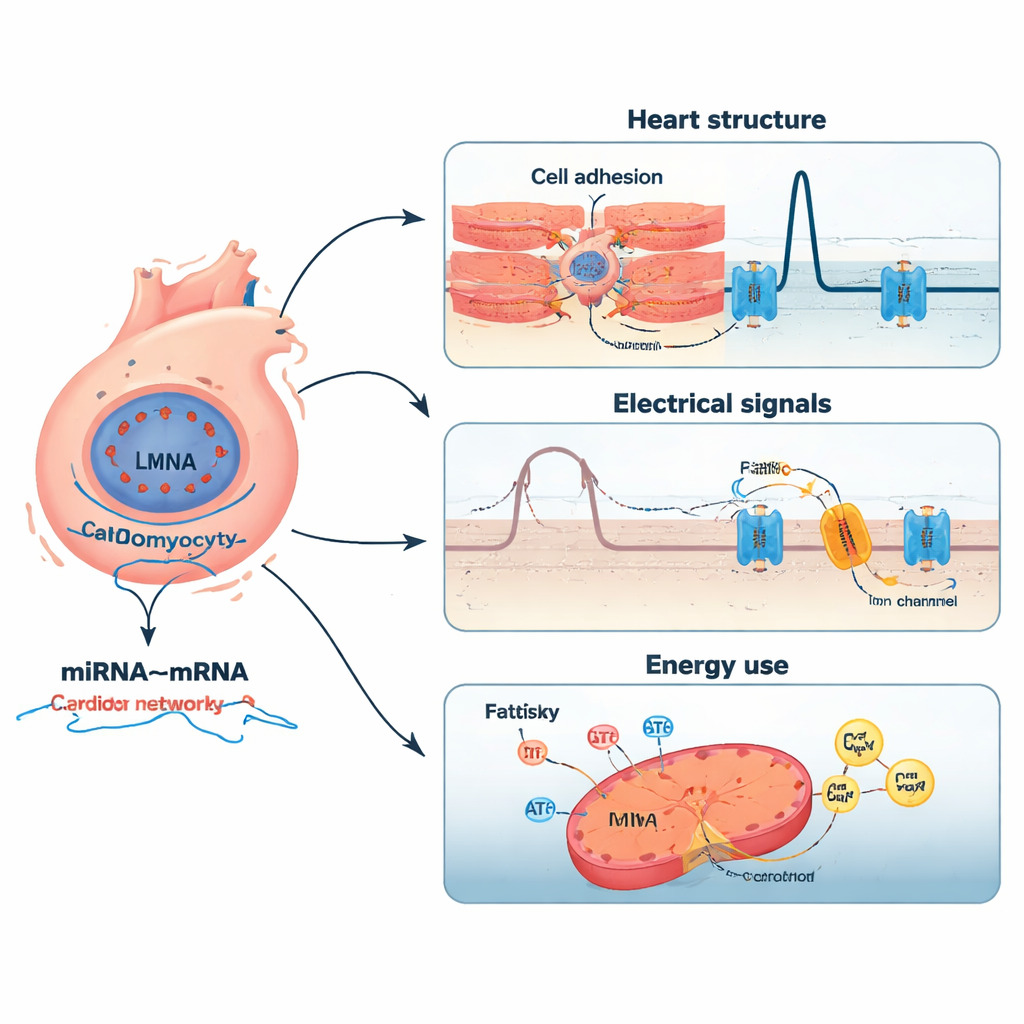
Networks of tiny switches and their targets
The study went further by pairing miRNAs with their likely mRNA targets. Using correlation analyses and cross-checking against large databases of experimentally validated interactions, the authors assembled a high-confidence network of 2,197 miRNA–mRNA pairs involving 12 key miRNAs. Some miRNAs were more active and appeared to dampen genes critical for heart development, calcium handling, and responses to oxidative stress. Others were less active, potentially releasing brakes on genes involved in cell adhesion, inflammation, and remodeling of the heart’s structural scaffold. For example, one miRNA (miR-183-5p) was linked to a receptor in the Wnt signaling pathway, which influences cell-to-cell communication, while another (miR-3473a) connected to a calcium-release channel central to heartbeat rhythm. Together, these networks outline how subtle shifts in tiny RNA regulators can amplify the impact of an LMNA mutation.
From molecular maps to future therapies
For non-specialists, the bottom line is that this study offers a detailed map of how an LMNA mutation reshapes the heart’s internal wiring at the level of gene control. Instead of a single “bad gene,” the disease reflects a cascade of altered messages—many orchestrated by miRNAs—that affect heart structure, electrical stability, and energy use. While the work was done in mice and relies heavily on computational analysis, it highlights specific miRNA–gene pairs as promising candidates for future blood-based biomarkers or targeted therapies. In the long run, tuning these tiny RNA switches could help doctors better predict who is at highest risk, monitor disease progression more precisely, and design more personalized treatments for families affected by LMNA-related cardiomyopathy.
Citation: Córdoba-Caballero, J., Martínez, F.B., Campuzano, O. et al. An integrative approach to identify novel miRNA-mRNA interaction networks in LMNA-cardiomyopathy. Sci Rep 16, 6110 (2026). https://doi.org/10.1038/s41598-026-36439-9
Keywords: dilated cardiomyopathy, LMNA gene, microRNA, gene regulation, heart failure