Clear Sky Science · en
QAL333’s antitumor activity and predictive modeling: integrated transcriptomic-bioinformatic analysis reveals selective cytotoxicity and sensitivity determinants
Why this new drug candidate matters
Cancer drugs often work well for some patients but fail for others, largely because tumors differ in their inner wiring. This study introduces a newly designed compound, QAL333, that appears to strike certain colorectal cancers much harder than others while sparing many normal cells. Just as important, the researchers show how patterns of gene activity in tumor cells could help predict who is most likely to benefit, hinting at more precise, personalized treatment strategies.
A focused strike on specific tumor types
The team began by testing QAL333, a synthetic molecule related to a family of compounds already known to damage cancer cells, on a broad panel of 15 human cancer cell lines and three non-cancerous lines. They found that colorectal cancer cells, especially a line called SW620, were highly sensitive, with the drug killing them at relatively low doses. In contrast, triple-negative breast cancer cells were far more resistant and required doses more than ten times higher that were difficult to achieve safely. Interestingly, not all non-cancerous cells reacted the same way: some were sensitive, but others, such as a widely used kidney cell line, were surprisingly resistant. This pattern showed that QAL333’s effects depend more on a cell’s internal biology than on whether it is labeled “cancer” or “normal.”
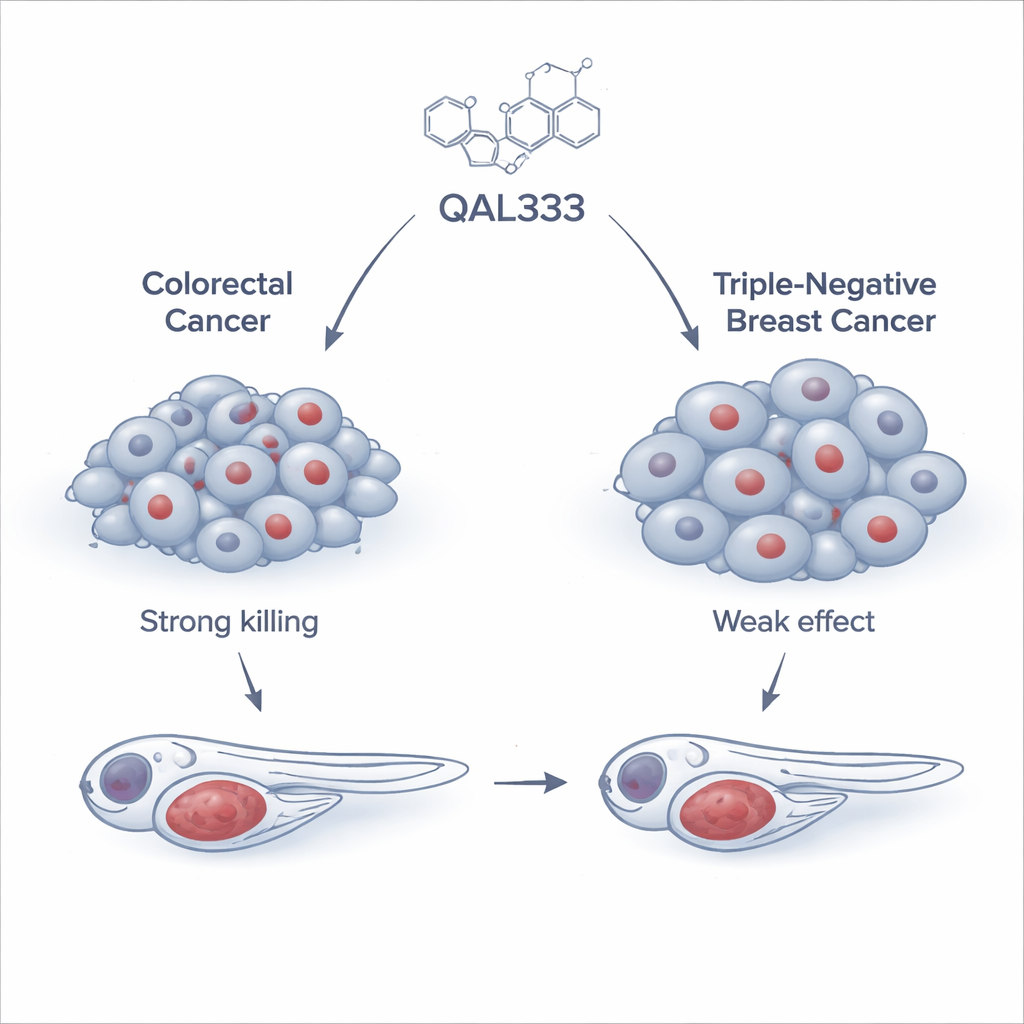
Testing the drug in living organisms
To see whether the promising cell-culture results would hold up in a living body, the researchers turned to zebrafish, a small transparent fish widely used to model human disease. They implanted fluorescently labeled human cancer cells into zebrafish embryos and treated them with QAL333. In fish carrying colorectal cancer cells, the drug clearly slowed tumor growth, approaching the effect of a standard chemotherapy combination used in the clinic. In contrast, tumors formed from the triple-negative breast cancer cells barely responded, even though a common breast cancer drug, paclitaxel, worked well in the same model. These in vivo tests reinforced that QAL333’s strength lies in a subset of tumors rather than across all cancers.
Peering inside cells to see how QAL333 works
To understand why some cells die and others resist, the scientists measured how thousands of genes changed their activity levels after QAL333 treatment. In sensitive colorectal cancer cells, key genes that drive the cell’s division cycle and growth were turned down, and major survival pathways were dampened. Signals linked to oxidative stress and a form of cell death distinct from classic programmed cell death were also altered, suggesting that QAL333 overloads already fragile systems that manage damage inside these cells. Resistant breast cancer cells, by contrast, switched on stress and inflammation programs controlled by factors like NF-κB, which are known to help cells cope with harmful conditions. This side-by-side comparison painted a picture of QAL333 overwhelming weak defenses in some cells while others mount an effective counter-response.
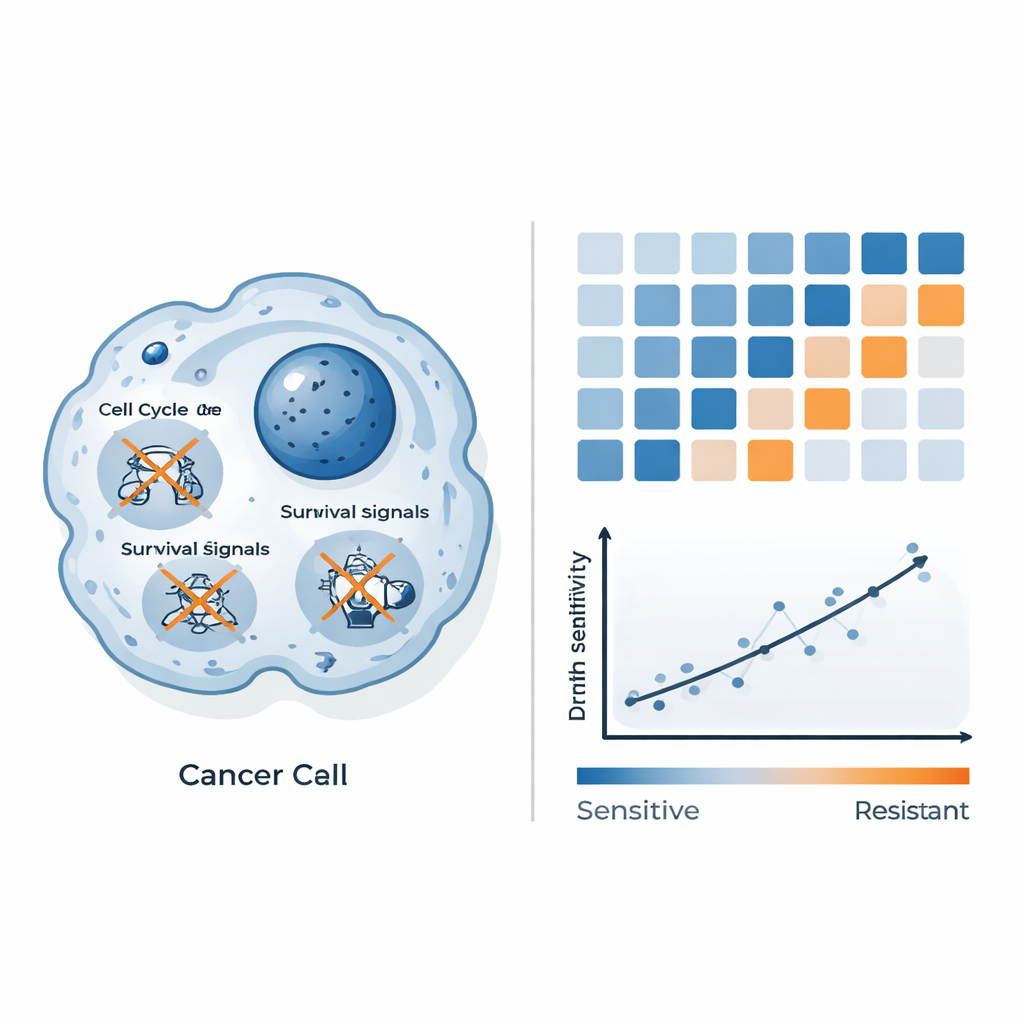
Using gene patterns to predict who will respond
Beyond cataloging gene changes, the researchers asked whether these patterns could be turned into a practical prediction tool. They combined baseline gene activity data from the panel of cell lines with their measured drug sensitivity and used machine learning to search for the most informative signals. Rather than relying on single genes, they grouped genes into biological pathways and created a combined “pathway score” built from three systems: how cells process energy and chemicals (metabolic pathways), how they organize their surroundings (proteoglycans and related structures in cancer), and how their internal skeleton is arranged. This composite score strongly tracked with how much QAL333 was needed to kill each cell line: lower scores aligned with higher sensitivity, and higher scores with resistance. In other words, the way these networks are tuned inside a tumor may forecast how well QAL333 will work.
Unexpected hints of broader biological effects
While the main focus was cancer, QAL333 also showed an ability to disrupt “quorum sensing,” a communication system bacteria use to coordinate behaviors like forming biofilms. At tiny doses, the compound reduced biofilm formation by a gum-disease bacterium and dampened certain inflammation signals in immune cells. The authors treat these as early, exploratory findings, but they suggest that QAL333 or related molecules might one day have uses beyond cancer therapy, for example in infections where bacterial communities are hard to eradicate.
What this could mean for future patients
Overall, the study positions QAL333 as a selective drug candidate that hits vulnerable colorectal tumors while sparing many other cell types, and it couples this with a roadmap for predicting which tumors are most likely to respond. If confirmed in more advanced animal models and eventually in patients, a simple gene-expression test could help doctors decide when QAL333 is a good match, aligning with the broader goal of tailoring cancer treatment to each tumor’s wiring. At the same time, the work underscores that even promising drugs have limits: breast cancers and other resistant tumors may require smart combinations or entirely different strategies.
Citation: Kim, H.H., Im, S., Kim, J. et al. QAL333’s antitumor activity and predictive modeling: integrated transcriptomic-bioinformatic analysis reveals selective cytotoxicity and sensitivity determinants. Sci Rep 16, 5648 (2026). https://doi.org/10.1038/s41598-026-36430-4
Keywords: colorectal cancer, targeted therapy, gene expression, drug sensitivity modeling, zebrafish xenograft