Clear Sky Science · en
From in-silico QSAR modeling to in-vitro MTT assay: experimental validation of novel uPAR leads for triple-negative breast cancer (TNBC) and skin cancer
Why this research matters
Cancers like triple‑negative breast cancer and certain skin cancers are notoriously hard to treat because they spread quickly and often resist standard therapies. This study explores a promising new target on the surface of cancer cells—called uPAR—and shows how combining computer‑based drug design with lab experiments can uncover new drug candidates that may slow or stop the spread of these aggressive tumors.
A doorway that helps cancer spread
For a tumor to turn deadly, its cells must break free from their original site, chew through surrounding tissue, enter the bloodstream, and colonize distant organs. uPAR is a key helper in this process. It sits on the cancer cell surface and works with partner proteins to activate enzymes that dissolve the surrounding support structure, making it easier for cells to move and invade. uPAR is found at especially high levels in triple‑negative breast cancer cells and in certain skin cancer cells, and its abundance is linked with faster growth, greater invasiveness, and poorer patient outcomes. Because of this, uPAR is an attractive—but so far underused—target for anti‑cancer drugs.
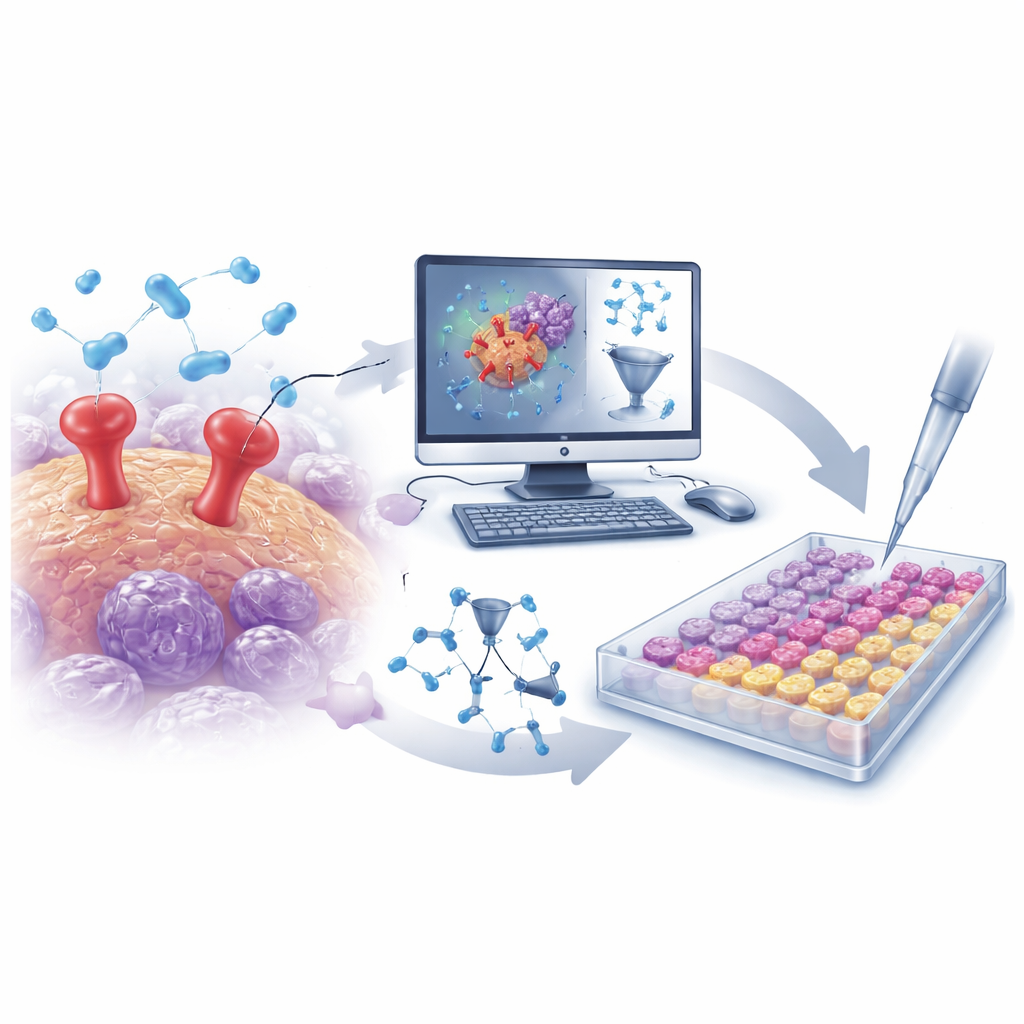
Mining big chemical data with smart algorithms
The researchers began by assembling a large collection of more than 500 known uPAR‑blocking molecules from a public database. Each molecule’s structure was translated into thousands of numerical descriptors that capture features such as size, shape, charge distribution, and flexibility. Using these data, the team built a quantitative structure–activity relationship (QSAR) model—a type of mathematical tool that learns which structural features tend to make a molecule a stronger uPAR inhibitor. After careful validation to avoid overfitting, the model showed high predictive power on unseen compounds, highlighting several key features that either strengthened or weakened uPAR blocking. These insights guided the search for new, better‑designed molecules.
From virtual screens to real test tubes
Armed with the trained QSAR model, the team virtually screened a focused library of about 30,000 protease‑targeting molecules. The model rapidly narrowed this vast pool to a small set of likely uPAR inhibitors. In parallel, computer docking simulations were used to see how well each candidate might fit into the uPAR binding site, and molecular dynamics simulations tested whether these docked complexes would remain stable over time. From this combined in‑silico campaign, two standout molecules emerged: D685‑0061, flagged by the QSAR model as highly potent, and C878‑1660, highlighted by docking and simulation as forming especially stable interactions with uPAR.
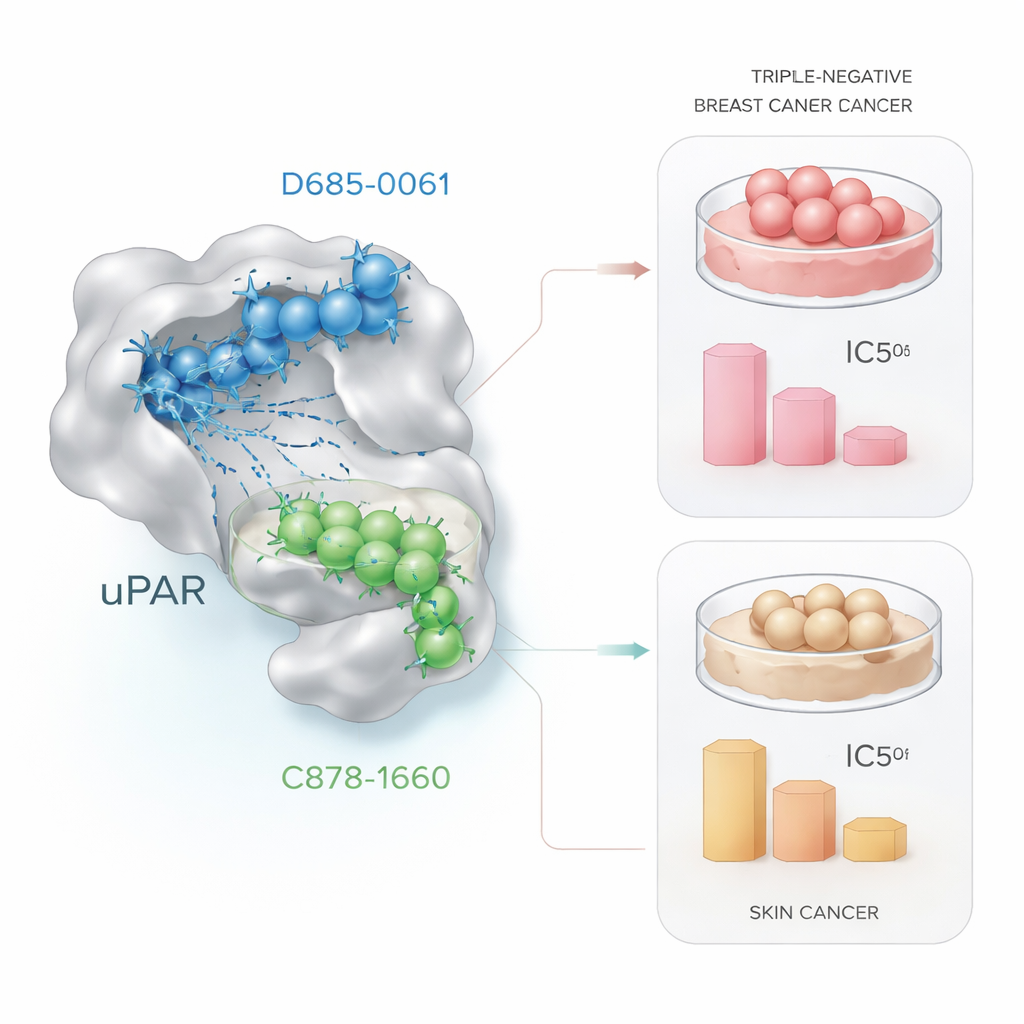
Putting the candidates to the test in cancer cells
Computational promise is only meaningful if it translates into real biological effects. To test this, the researchers exposed live cancer cells to increasing doses of the two lead molecules and measured cell survival using a standard laboratory method called the MTT assay. In triple‑negative breast cancer cells (MDA‑MB‑231), D685‑0061 killed cells more efficiently, cutting viability in half at around 21 micromoles, while C878‑1660 required roughly four times that amount. Under the microscope, cells treated with D685‑0061 showed clear signs of programmed cell death, such as rounding up, detaching from the surface, and breaking into fragments. In contrast, in skin cancer cells (A431), C878‑1660 was the stronger killer, achieving 50% cell death at about 19 micromoles compared with roughly 28 micromoles for D685‑0061, again with visible dose‑dependent damage.
What this means for future cancer treatments
This work does not deliver an immediate new drug, but it does deliver two promising starting points and, just as importantly, a powerful blueprint for finding more. By linking big‑data modeling, 3D simulations, and careful cell‑based experiments, the study shows that uPAR can be selectively targeted with small molecules that damage aggressive breast and skin cancer cells. D685‑0061 and C878‑1660 now serve as lead compounds that chemists can refine to boost potency, improve safety, and enhance how they move through the body. In the longer term, drugs built on this approach could help block the spread of uPAR‑driven cancers, turning some of today’s most dangerous tumors into more manageable diseases.
Citation: Badukle, H., Jawarkar, R.D., Shah, U. et al. From in-silico QSAR modeling to in-vitro MTT assay: experimental validation of novel uPAR leads for triple-negative breast cancer (TNBC) and skin cancer. Sci Rep 16, 5786 (2026). https://doi.org/10.1038/s41598-026-36406-4
Keywords: uPAR inhibitors, triple-negative breast cancer, skin cancer, computer-aided drug design, MTT cell viability assay