Clear Sky Science · en
An interactive cascaded deep learning framework with expert refinement for accurate striatal subregion segmentation
Why mapping tiny brain regions matters
The striatum, a small structure buried deep in the brain, plays a central role in movement and motivation and is heavily affected in disorders such as Parkinson’s disease. Doctors often use PET and MRI scans to measure chemical changes in this area, but the striatum is divided into many tiny subregions that are hard to see and outline precisely. This paper introduces a new computer-assisted method, called StriaSeg‑iARM, that helps specialists draw these boundaries more accurately and efficiently, even in people whose brains are altered by aging or disease.
The clinical problem in plain sight
When neurologists evaluate conditions like Parkinson’s disease, they rely on PET scans to track how dopamine-related signals vary across different parts of the striatum. To do this correctly, they must first define small three‑dimensional “regions of interest” on MRI, which serve as measuring cups for the PET data. Manually outlining these regions is slow, tedious work and can differ from one expert to another. Existing software can automatically locate only the larger structures and often struggles in patients whose brains are shrunken or distorted, a common situation in advanced neurodegenerative disease. These shortcomings can blur the true pattern of damage and weaken the power of imaging as a diagnostic tool.
A smarter two-step assistant
The researchers designed StriaSeg‑iARM as a two‑stage deep learning system that works directly in each person’s own brain space rather than forcing the scan into a standard template. In the first stage, the program finds the overall striatum on a three‑dimensional MRI. In the second stage, it divides this structure into 12 anatomically defined subregions on both sides of the brain. Between these two stages lies the key innovation: an interactive step in which an expert can quickly check and, if needed, correct the coarse outline produced by the first stage before the finer subdivision is run. This approach blends the speed of automation with the judgement of a trained human observer.
Training and testing in real-world brains
To teach the system what to look for, the team assembled hundreds of MRI scans from multiple hospitals and scanner types, and created careful manual outlines of the 12 striatal subregions. They then tested the model on two outside datasets that included both MRI and dopamine‑related PET scans: one from relatively early Parkinson’s patients and healthy volunteers, and another from an older, more severely affected clinical group with marked brain shrinkage and mixed diagnoses. Across these challenging cases, StriaSeg‑iARM consistently matched manual expert drawings better than traditional atlas‑based methods and a simpler one‑stage deep learning model, with higher overlap, more accurate boundaries, and stable volume measurements.
Sharper measurements for brain chemistry scans
Because the final goal is to quantify PET signals, the authors also examined how different segmentation methods affected PET readouts. Using a standard measure of tracer binding in each subregion, they compared automatic results with those based on manual outlines. The atlas method tended to underestimate binding, especially in patients with more severe changes, while the basic deep learning model still showed noticeable bias in some regions. In contrast, the cascaded models—especially when the interactive correction step was used—produced PET values that closely tracked the expert reference, with minimal bias and scatter. The team further showed that a simple uncertainty score from the first stage can flag about one quarter of the most difficult cases where expert refinement is truly needed, keeping the rest fully automatic.
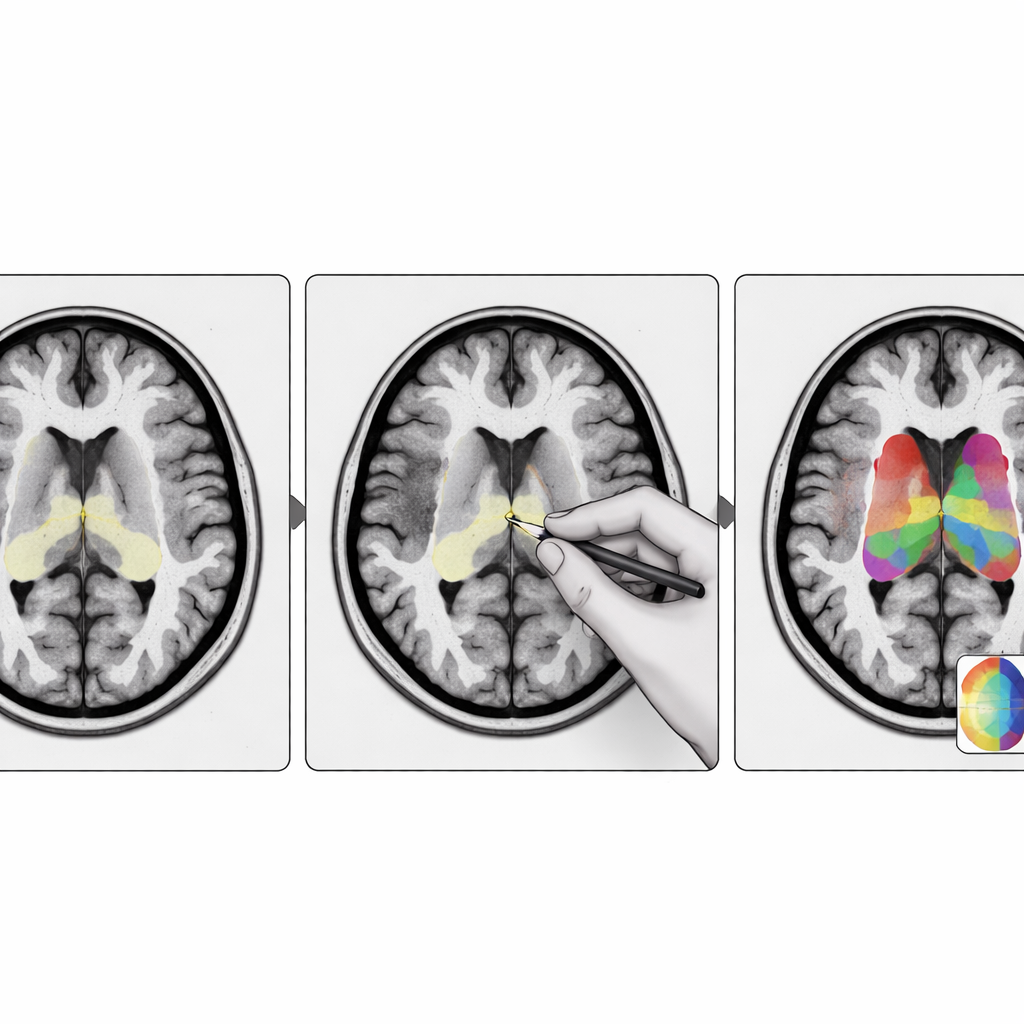
What this means for patients and clinicians
For non‑specialists, the take‑home message is that better digital maps of small brain regions can make chemical brain scans more trustworthy. StriaSeg‑iARM provides a flexible framework in which a deep learning model does most of the work, but specialists can still step in to correct tricky cases without retraining the system. This combination of automation and expert oversight improves the precision of measurements used to diagnose and track disorders like Parkinson’s disease, and it could be adapted to other organs and diseases where small, hard‑to‑see structures hold vital clues.
Citation: Kim, J., Kim, D., Kim, S. et al. An interactive cascaded deep learning framework with expert refinement for accurate striatal subregion segmentation. Sci Rep 16, 6550 (2026). https://doi.org/10.1038/s41598-026-36399-0
Keywords: Parkinson’s disease imaging, striatal segmentation, deep learning in radiology, brain PET MRI, neurodegenerative biomarkers