Clear Sky Science · en
Deploying the high-throughput virtual screening (HTVS) approach for the identification of new lactate dehydrogenase (LDH) inhibitors with anticancer assets
Why this research matters for cancer treatment
Cancer cells often rewire how they make energy, and this shift can be exploited to design smarter, more selective drugs. This study explores a key metabolic enzyme called lactate dehydrogenase (LDH), which helps tumors thrive in harsh conditions and resist treatment. Using advanced computer-based screening and lab tests, the researchers identified two new small molecules that block LDH and show promising anticancer effects in prostate cancer cell lines.
The strange metabolism of cancer cells
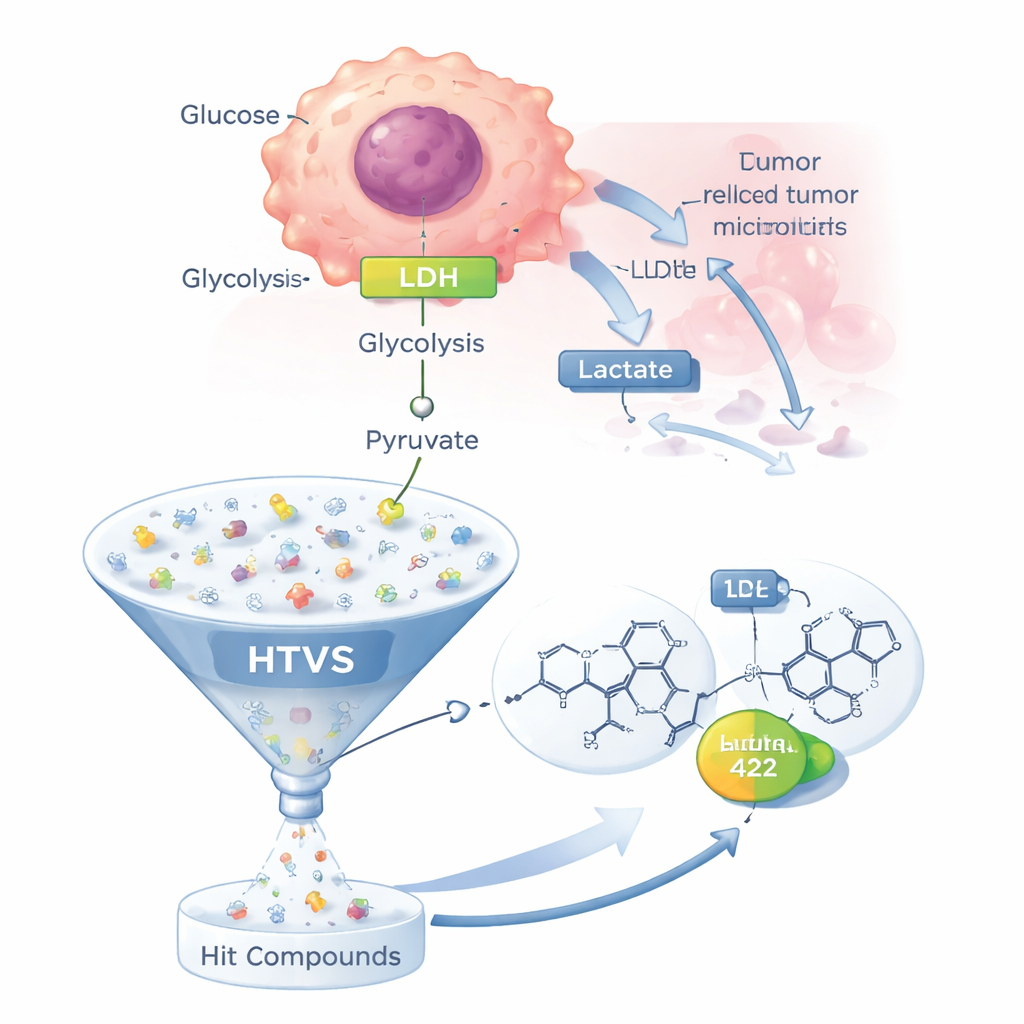
Most healthy cells use oxygen efficiently in their mitochondria to generate energy. Many cancer cells, however, rely heavily on a less efficient pathway called glycolysis, even when oxygen is plentiful—a phenomenon known as the Warburg effect. In this shortcut route, glucose is broken down to pyruvate, which is then converted into lactate by LDH instead of being fed into the mitochondria. The lactate is pumped out of the cell, acidifying the tumor’s surroundings, helping cancer cells evade cell death, weaken immune attack, and even blunt the effects of many chemotherapy drugs. Because LDH sits at the heart of this altered metabolism and no LDH-targeting drug has yet been approved, it has become a high-priority target for new anticancer therapies.
Using computers to sift through half a million molecules
Rather than testing potential drugs one by one in the lab, the team used a high-throughput virtual screening (HTVS) strategy. They started with 28 LDH-blocking molecules already reported in the scientific literature and distilled from them a "pharmacophore"—an abstract 3D pattern of features that any good LDH inhibitor is likely to share. They then applied this model to a commercial library of about 500,000 drug-like molecules, asking which candidates matched the essential features. Roughly 110,000 molecules passed this first filter, and further selection based on standard "drug-likeness" rules shrank the list to 2,337 more realistic candidates for docking to the LDH enzyme.
Zooming in on the best binders
The next step was to estimate how tightly these candidates could fit into the active site of LDH. Using several levels of computational docking, the researchers calculated how each molecule might interact with specific amino acids inside the enzyme pocket. This multi-stage process progressively narrowed the pool from thousands of candidates to just 59 promising molecules, and then to five clear "hits" with especially strong predicted binding. Two of these, labeled compounds 15 and 422, stood out because computer-based molecular dynamics simulations suggested that they form unusually stable complexes with LDH over time, maintaining key contacts with critical residues in the active site while preserving a reasonable overall protein structure.
From the screen to real cancer cells
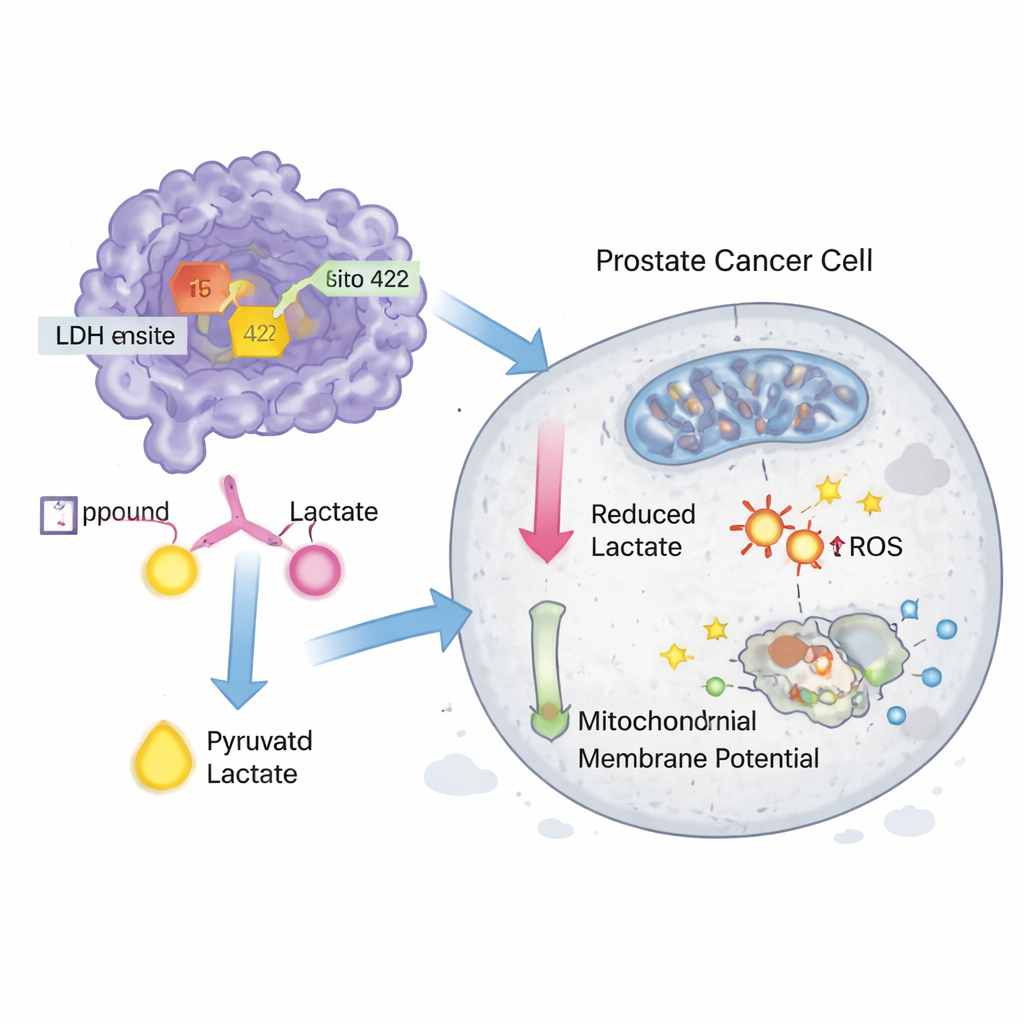
To see whether these hits mattered in living systems, the scientists purchased compounds 15 and 422 and tested them in prostate cancer cell lines DU-145 and PC-3. Both molecules inhibited LDH activity in cells at nanomolar concentrations and also directly slowed the activity of a purified LDH enzyme, although they were somewhat less potent than a known experimental inhibitor called GNE‑140. In growth assays, the compounds reduced cancer cell survival at low micromolar doses, again approaching the performance of GNE‑140. Additional experiments showed that the treatments slightly increased oxidative stress, disturbed mitochondrial membrane potential—a sign of impaired energy factories—and triggered programmed cell death (apoptosis) in DU‑145 cells, with compound 15 being the more active of the two.
What this could mean for future cancer drugs
While these molecules are not ready-made medicines, they provide valuable starting points for drug design. Both compounds 15 and 422 have chemical properties that suggest better solubility, permeability, and overall "drug-likeness" than some existing LDH inhibitors, making them attractive leads for further optimization. The study demonstrates how large-scale computer screening, guided by known structural features, can uncover new LDH inhibitors that weaken cancer cells by starving their altered energy supply. With future work to refine their structures, test direct binding more rigorously, and study their behavior in animals, these compounds could contribute to a new class of anticancer agents that attack tumors through their unique metabolic vulnerabilities.
Citation: Huang, Y., Benni, S., Yadav, U.P. et al. Deploying the high-throughput virtual screening (HTVS) approach for the identification of new lactate dehydrogenase (LDH) inhibitors with anticancer assets. Sci Rep 16, 5921 (2026). https://doi.org/10.1038/s41598-026-36385-6
Keywords: lactate dehydrogenase inhibitors, cancer metabolism, virtual screening, prostate cancer, anticancer drug discovery