Clear Sky Science · en
Translational evaluation of BUB1B as a precision medicine biomarker for hepatocellular carcinoma
Why a cell-division gene matters for liver cancer
Most people think of cancer in terms of tumors seen on scans, but deep inside those growths are microscopic control systems that decide how and when cells divide. This study focuses on one such control switch, a gene called BUB1B, and shows how it could help doctors predict which liver cancer patients will do poorly, who might respond to certain drugs, and why some tumors outsmart the immune system. By turning a basic cell-cycle regulator into a "dashboard" for liver cancer behavior, the work points toward more precise and individualized treatment.
A common liver cancer needs better guides
Hepatocellular carcinoma, the most frequent form of primary liver cancer, is often discovered late, when symptoms finally appear and treatment options are limited. Existing blood tests and tissue markers, such as alpha-fetoprotein or Ki-67, are imperfect: they do not reliably catch early disease or guide tailored therapies. The authors therefore searched for a different kind of marker—one that reflects the inner wiring of cancer cells rather than just their presence. They focused on BUB1B, a protein that helps ensure chromosomes are properly divided when cells split. When this system goes awry, genetic errors accumulate, which can fuel aggressive tumors.
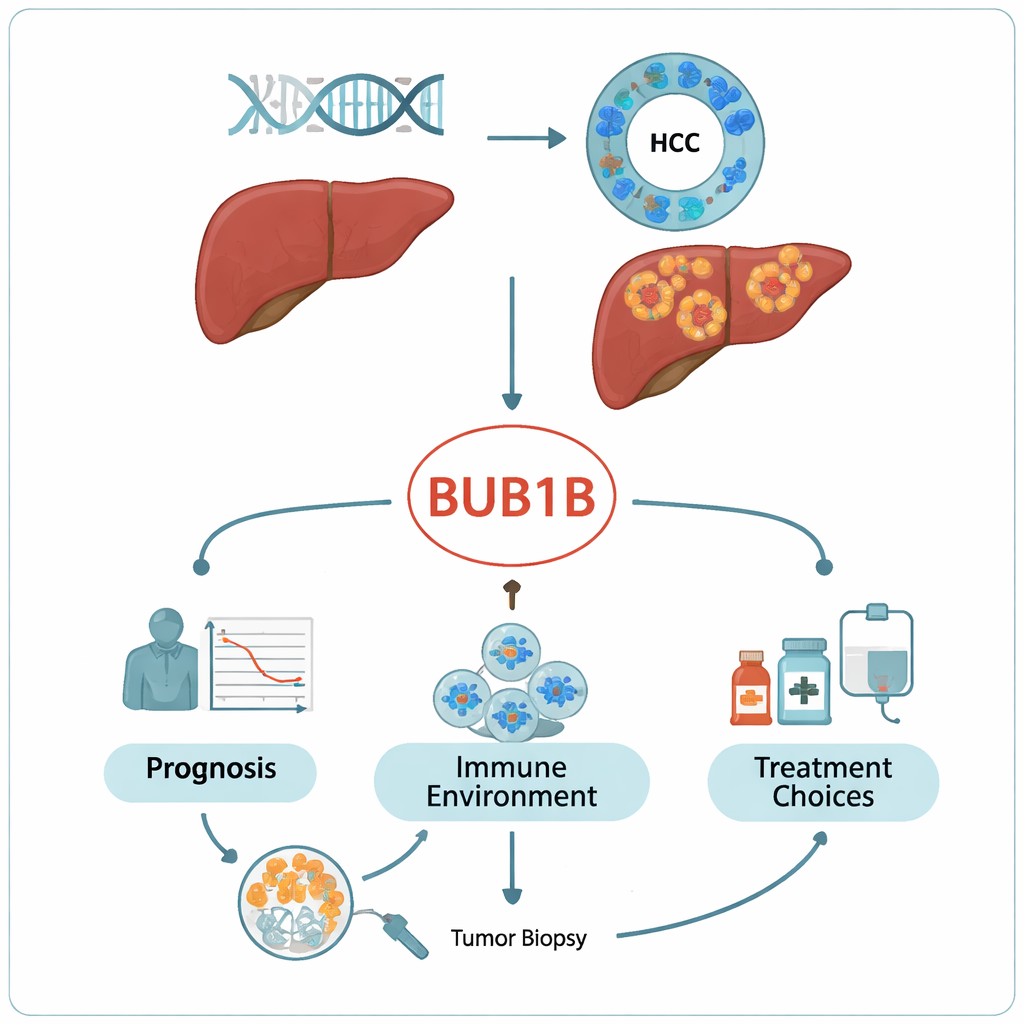
Tracking BUB1B across cancers and real patients
Using large public cancer databases, including The Cancer Genome Atlas, the researchers first looked at BUB1B activity across many tumor types. They found it was switched on at higher levels in 19 different cancers, with particularly strong overactivity in liver tumors compared with nearby healthy tissue. Patients whose liver cancers had more BUB1B tended to live for a shorter time, even after accounting for how advanced their cancer was, making BUB1B an independent warning sign. An in-depth review of 50 liver cancer patients treated at their own hospital confirmed the pattern: tumors showed intense BUB1B staining under the microscope, especially in the cell cytoplasm, and high levels went hand in hand with larger tumors, more advanced stage, and worse survival.
Genetic chaos, drug response, and a confused immune system
High BUB1B did not just signal that a tumor was dangerous; it was tied to deeper instability in the cancer’s DNA. Tumors with more BUB1B showed more signs of defective DNA repair and subtle forms of microsatellite instability, hallmarks of genomic chaos that can both drive rapid evolution and shape how tumors respond to treatment. Computer models linking gene patterns to drug tests suggested that BUB1B-high liver cancers might actually be more vulnerable to certain medicines, including the targeted drug sorafenib and classic chemotherapies such as paclitaxel and doxorubicin, while being relatively resistant to drugs focusing on the EGFR pathway. At the same time, these tumors contained many immune cells, especially T cells, yet still scored high on measures of immune dysfunction and exclusion—meaning the immune system was present but effectively disarmed, and less likely to respond to common immunotherapies that block PD-1 or PD-L1.
How BUB1B fuels growth signals inside cancer cells
To move beyond statistical links, the team tested what happens when BUB1B is chemically blocked in liver cancer cells grown in the lab. Inhibiting this gene dialed down a whole chain of growth signals commonly grouped under the MAPK pathways. Key players at several levels—including RAS, certain RAF family members, ERK, and multiple p38 and other related kinases—were sharply reduced. Because these pathways act like a relay that tells cancer cells to keep dividing and resist stress, cutting them back suggests that BUB1B is not just a bystander marker but an active driver that helps maintain the cancer’s internal wiring. Some components showed unexpected rebounds, hinting at built-in feedback loops, but the overall effect was a broad weakening of pro-growth signaling.
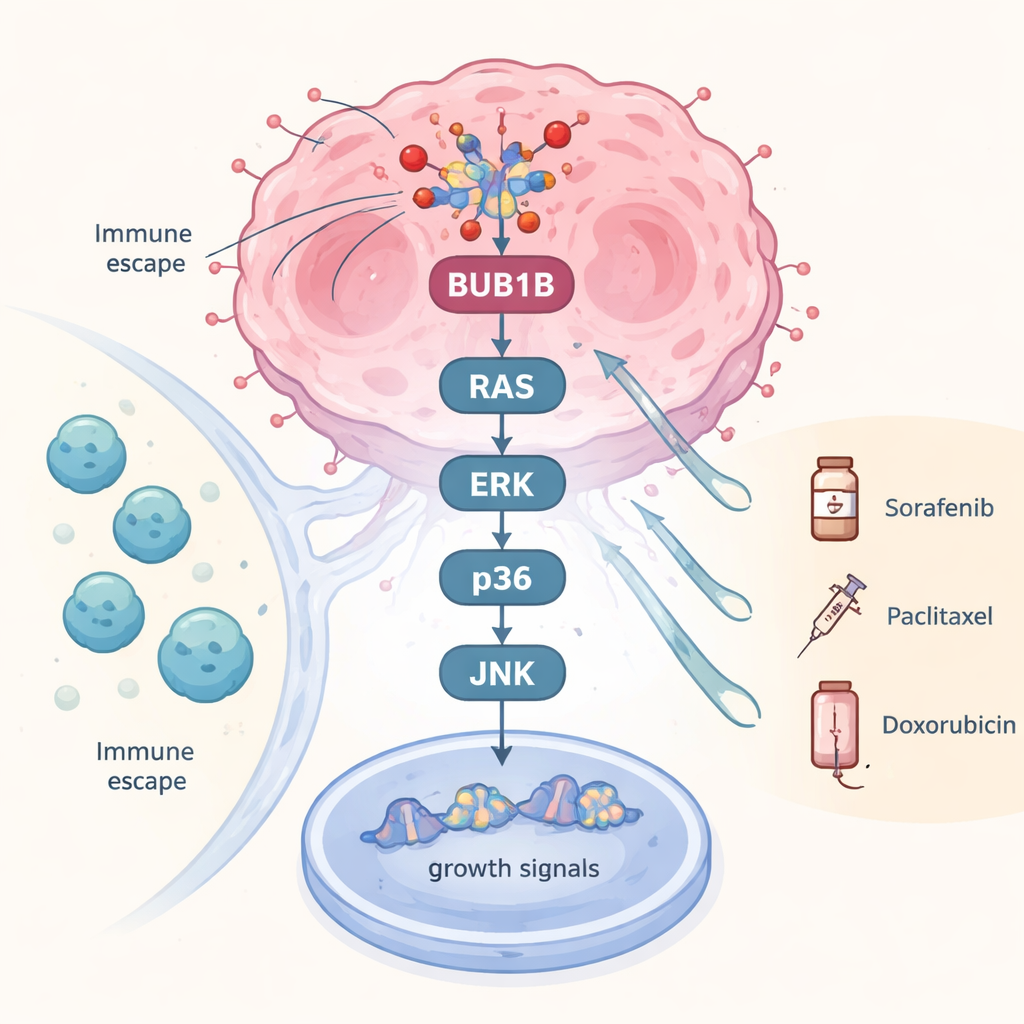
What this means for patients and future care
For non-specialists, the takeaway is that a single cell-division gene, BUB1B, appears to sit at the crossroads of three crucial features of liver cancer: how unstable the tumor’s DNA is, how it talks to the immune system, and how it responds to several key drugs. Measuring BUB1B in tumor samples could help doctors sort patients into risk groups, estimate survival more accurately, and choose treatments that exploit the cancer’s weaknesses—while recognizing when standard immunotherapy is unlikely to work. Because liver tumors rely on BUB1B to keep their growth and stress-response signals running, drugs that target this protein, alone or in combination with existing therapies, may open new paths toward more precise and effective treatment of hepatocellular carcinoma.
Citation: Sun, Cy., Yu, X., Deng, Lq. et al. Translational evaluation of BUB1B as a precision medicine biomarker for hepatocellular carcinoma. Sci Rep 16, 5301 (2026). https://doi.org/10.1038/s41598-026-36364-x
Keywords: hepatocellular carcinoma, BUB1B, biomarker, MAPK signaling, precision oncology