Clear Sky Science · en
Bacterial genome reconstruction and community profiling in Neotropical Drosophila
Why tiny flies and their germs matter
Fruit flies may seem like kitchen pests, but for scientists they are powerful tools for understanding how microbes shape animal health, including our own. This study looks at the microscopic life living in and on wild fruit flies from Ecuador, using modern DNA sequencing to rebuild the genomes of their bacteria and map out who lives where and how they interact. By examining dozens of fly species from the tropical Andes, the researchers ask a simple but far-reaching question: are a fly’s microbes mainly a product of its family tree, or of the food and environment it experiences?
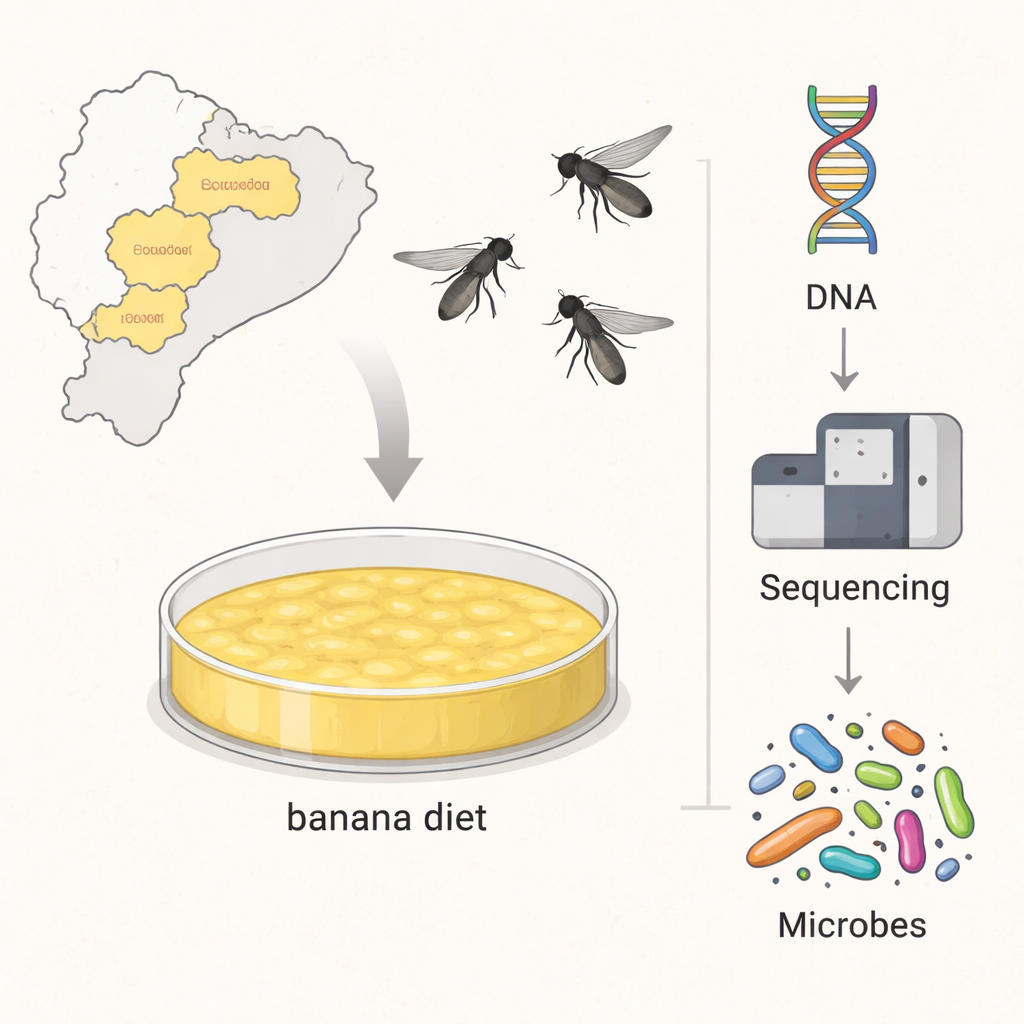
Flies from the Andes under the microscope
The team collected 24 species of Neotropical Drosophila from nine provinces across Ecuador and then kept them on the same banana-based diet under controlled laboratory conditions. Instead of targeting just a few genes, they used shotgun metagenomics, reading all the DNA present in each sample. This allowed them not only to list the microbes present but also to piece together whole bacterial genomes from the mixed genetic soup. After stripping away the flies’ own DNA and any human contamination, the remaining sequences revealed a rich community of bacteria and fungi associated with these tropical flies.
Who lives in the fly gut?
Across species, the microbial cast of characters was surprisingly consistent. Yeasts from the group Saccharomycetales were the most common non-bacterial residents, while the main bacterial players were acetic acid bacteria (such as Acetobacter and Gluconobacter), lactic acid bacteria, and members of the Enterobacterales, along with the reproductive parasite Wolbachia in some flies. However, these groups did not appear in fixed proportions. Samples tended to fall into two broad patterns: some were dominated by acetic acid bacteria and yeasts, while others had more Enterobacterales and lactic acid bacteria instead. This suggested that the community can reorganize into different stable mixtures rather than being locked into a single arrangement.
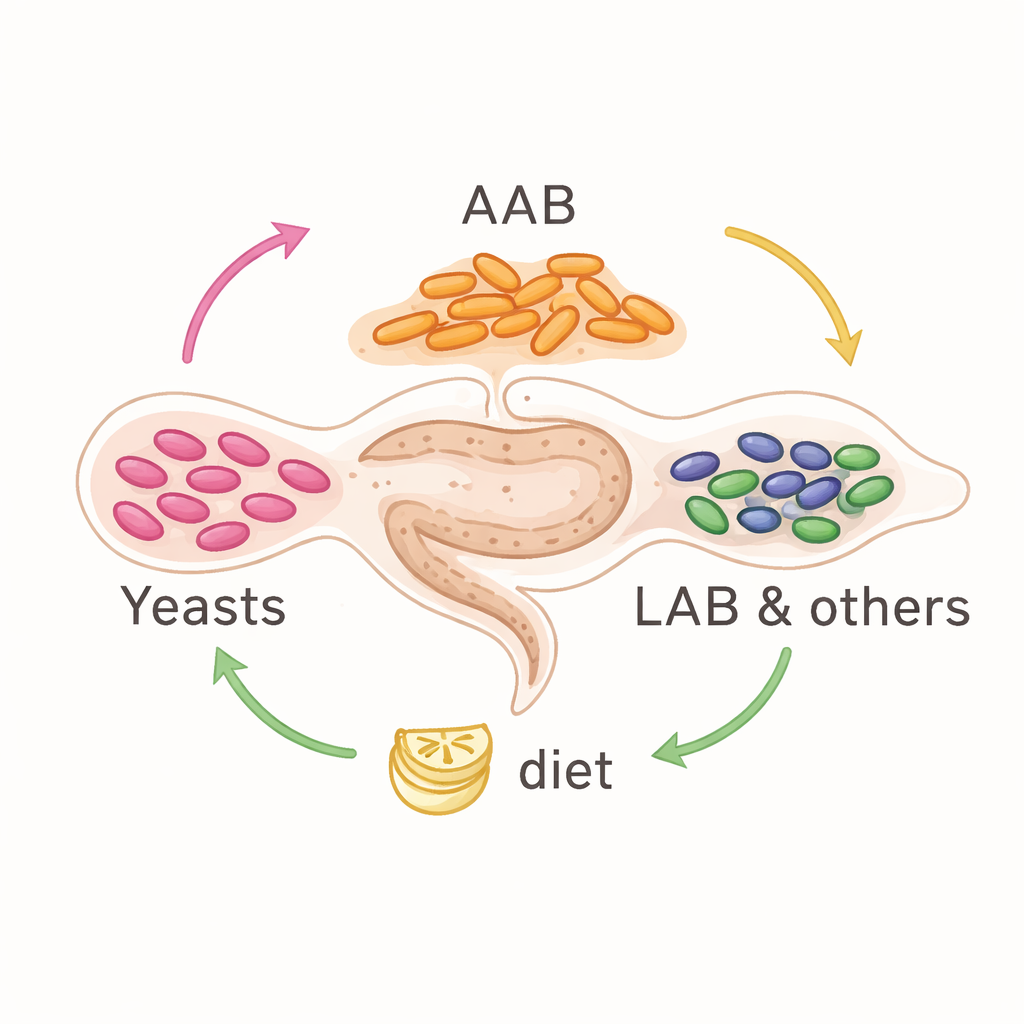
Rebuilding bacterial “blueprints”
Using the metagenomic data, the researchers reconstructed 64 high-quality bacterial genomes, mostly from acetic acid and Enterobacterales lineages. These genomes included repeated appearances of species such as Acetobacter thailandicus and Gluconobacter kondonii, many of which have also been found in the common lab fly Drosophila melanogaster. Detailed comparisons showed that the recovered genomes matched known reference strains closely and carried genes suited to life in a sugar-rich, fermenting environment. Many were equipped to break down sugars via specialized pathways, handle fermentation products like lactate and acetate, and synthesize vitamins and amino acids that could, in principle, benefit their fly hosts.
Environment over ancestry
A central question was whether closely related fly species host more similar microbial communities, an idea known as phylosymbiosis. To test this, the team compared evolutionary trees built from fly genomes with diagrams summarizing the similarities and differences among their microbiota. They also focused on one widespread bacterial species, Acetobacter thailandicus, and compared its evolutionary tree with that of its fly hosts. In both cases, the match between host relatedness and microbial similarity was weak. Instead, factors like the balance between acetic acid bacteria and Enterobacterales, and the abundance of yeasts, explained much more of the variation, pointing toward diet and microbial interactions as major drivers.
A changing community shaped by food and cooperation
Putting the pieces together, the authors propose that the microbes in these Neotropical fruit flies form a flexible community molded less by the flies’ family histories and more by shared food sources and microbe–microbe cooperation. In a banana-based, fermenting environment, yeasts and bacteria trade nutrients and by-products, with some groups paving the way for others in a kind of ecological relay. The newly reconstructed bacterial genomes, many from species repeatedly seen in different flies, provide a rich resource for exploring how these tiny partners help their hosts grow, cope with stress, and adapt to changing diets—insights that ultimately inform our broader understanding of how environment and ecology, not just ancestry, shape the invisible worlds inside animals.
Citation: Ulloa, M.A., Serrano, A.V., Camelo, L.C. et al. Bacterial genome reconstruction and community profiling in Neotropical Drosophila. Sci Rep 16, 6601 (2026). https://doi.org/10.1038/s41598-026-36282-y
Keywords: fruit fly microbiome, Neotropical Drosophila, gut bacteria, metagenomics, host microbe interactions