Clear Sky Science · en
Weakly supervised colorectal gland segmentation through self-supervised learning and attention-based pseudo-labeling
Why this matters for cancer diagnosis
When a pathologist looks at a colon biopsy under the microscope, one of the most important clues to cancer severity is the shape and organization of tiny tube-like structures called glands. Carefully outlining every gland by hand is slow, expensive, and hard to standardize across hospitals. This study shows how artificial intelligence can learn to trace these glands almost as well as expert humans while using far less detailed human labeling, potentially speeding and sharpening colorectal cancer diagnosis.
The challenge of drawing every tiny outline
Colorectal cancer is among the most common and deadly cancers worldwide, and grading its severity depends heavily on gland appearance. In healthy or early-stage tissue, glands look like neat, round tubes; in aggressive tumors they become jagged, fused, or barely recognizable. Computers can be trained to segment, or "color in," each gland to allow automatic measurements, but traditional deep-learning systems require laborious pixel-by-pixel outlines drawn by expert pathologists. In real clinics, what is far easier to obtain are simple image-level labels, such as whether a tile of tissue does or does not contain glands, or whether it is benign or malignant.
Teaching an AI from unlabeled and weakly labeled slides
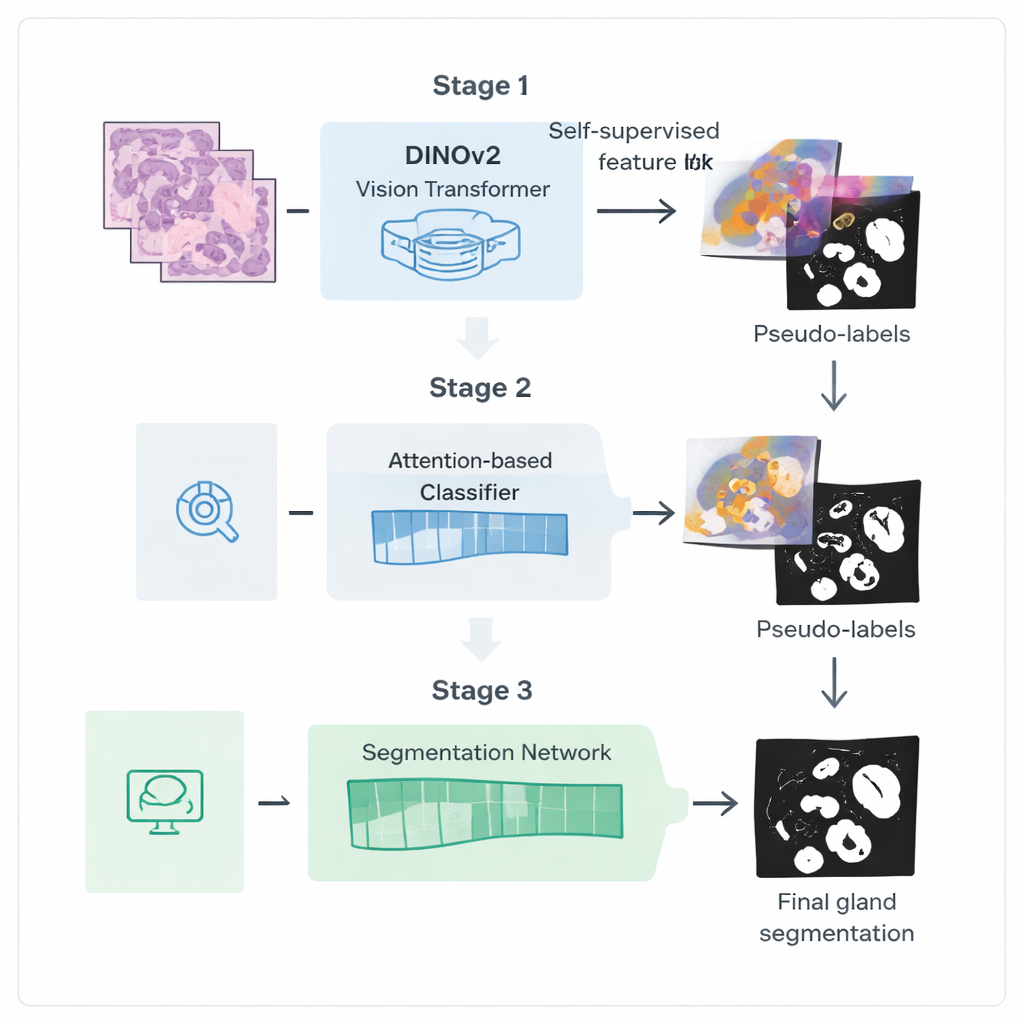
The authors introduce a three-step training pipeline designed to squeeze more value out of these weaker labels. First, they start from a powerful vision model called DINOv2, originally trained on natural photographs, and expose it to thousands of unlabeled colorectal biopsy images. By asking the model to match different views of the same tissue patch to each other, it learns visual features that are tuned to the colors and textures of histology slides without needing any annotations. This step creates a specialized "encoder" that transforms raw images into rich internal representations capturing gland-like structures.
Letting the AI show where it is looking
In the second stage, this encoder is plugged into a classification network that only needs image-level labels, such as whether glands are present. An attention mechanism inside the network learns to assign higher weights to the image regions that matter most for its decision. These attention maps effectively highlight where the network "believes" glands are located. The researchers turn these soft heatmaps into rough binary masks using blending and thresholding, then clean them further with a probabilistic smoothing technique called a Conditional Random Field. The result is a set of refined pseudo-labels: computer-generated gland outlines that are not perfect, but good enough to guide a more specialized segmentation model.
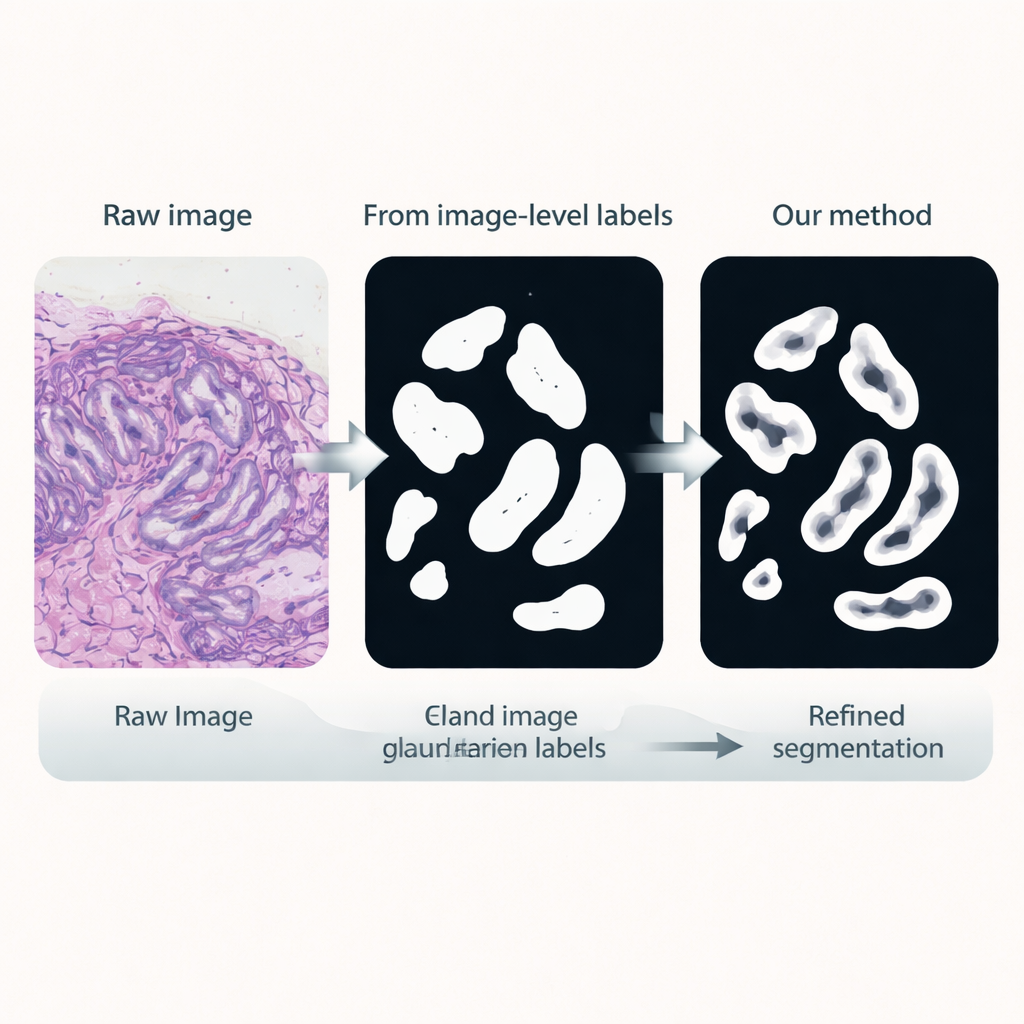
Sharpening gland boundaries
In the third stage, a dedicated segmentation network is trained using these pseudo-labels as stand-ins for manual annotations. It reuses the fine-tuned encoder but adds a light-weight decoder head that converts features back into a detailed gland mask. Crucially, the loss function used during training pays extra attention to boundaries: mistakes that distort the gland edges are penalized more than small errors in the interior. This boundary-aware training encourages crisp, anatomically realistic outlines, which are essential for accurately measuring gland shape and separation.
How well does it work in practice?
The team tested their method on two standard benchmarks of colorectal tissue. On the GlaS dataset, their weakly supervised approach not only beat other methods that also use limited labels, but in several measures approached or exceeded classic fully supervised systems that relied on full pixel-level annotations. On a tougher dataset called CRAG, packed with highly irregular, malignant glands, performance dropped for all methods, yet the new framework still outperformed other weak-label competitors and narrowed the gap with fully supervised models. Ablation studies showed that each component—self-supervised fine-tuning, attention-based pseudo-labeling with post-processing, and boundary-aware loss—contributed meaningfully to the gains.
What this means for future pathology tools
For a lay reader, the key takeaway is that this work points toward AI systems that can deliver high-quality, boundary-precise maps of microscopic gland structures while relying mainly on simple slide-level labels that are already common in hospital archives. By reducing dependence on painstaking manual outlining, the approach could make advanced image-based grading and quantitative analysis more feasible across many centers, helping pathologists diagnose colorectal cancer more consistently and efficiently, and potentially extending to other tissue types and structures in the future.
Citation: Wen, H., Wu, Y., Huang, D. et al. Weakly supervised colorectal gland segmentation through self-supervised learning and attention-based pseudo-labeling. Sci Rep 16, 5771 (2026). https://doi.org/10.1038/s41598-026-36256-0
Keywords: colorectal cancer, digital pathology, gland segmentation, weakly supervised learning, self-supervised vision