Clear Sky Science · en
Hybrid deep learning framework for accurate classification of high dimensional genomic data
Making Sense of the Genome Data Deluge
Modern DNA technologies can measure tens of thousands of genes in a single experiment, promising earlier disease detection and more precise treatments. Yet this wealth of data is so large, noisy, and complex that even powerful computer models often struggle to find clear, trustworthy patterns. This paper introduces a new kind of artificial intelligence (AI) system that is designed specifically to handle such overwhelming genomic data, aiming to make predictions more accurate while also explaining how those predictions were made.
Why Genomic Data Is So Hard to Use
Genomic studies routinely produce far more measurements than there are patients or samples. Many of these measurements are irrelevant, redundant, or distorted by technical noise. Traditional machine-learning methods either require human experts to manually pick which genes might matter, or they attempt to use everything and risk overfitting—that is, performing well on training data but failing on new cases. Deep learning, which has transformed fields like image recognition, can automatically learn patterns from raw data. However, in genomics it often behaves like a black box: it may give accurate answers, but offers little insight into why, limiting its acceptance in medicine where transparency is essential.
A Hybrid AI Blueprint for Gene-Based Decisions
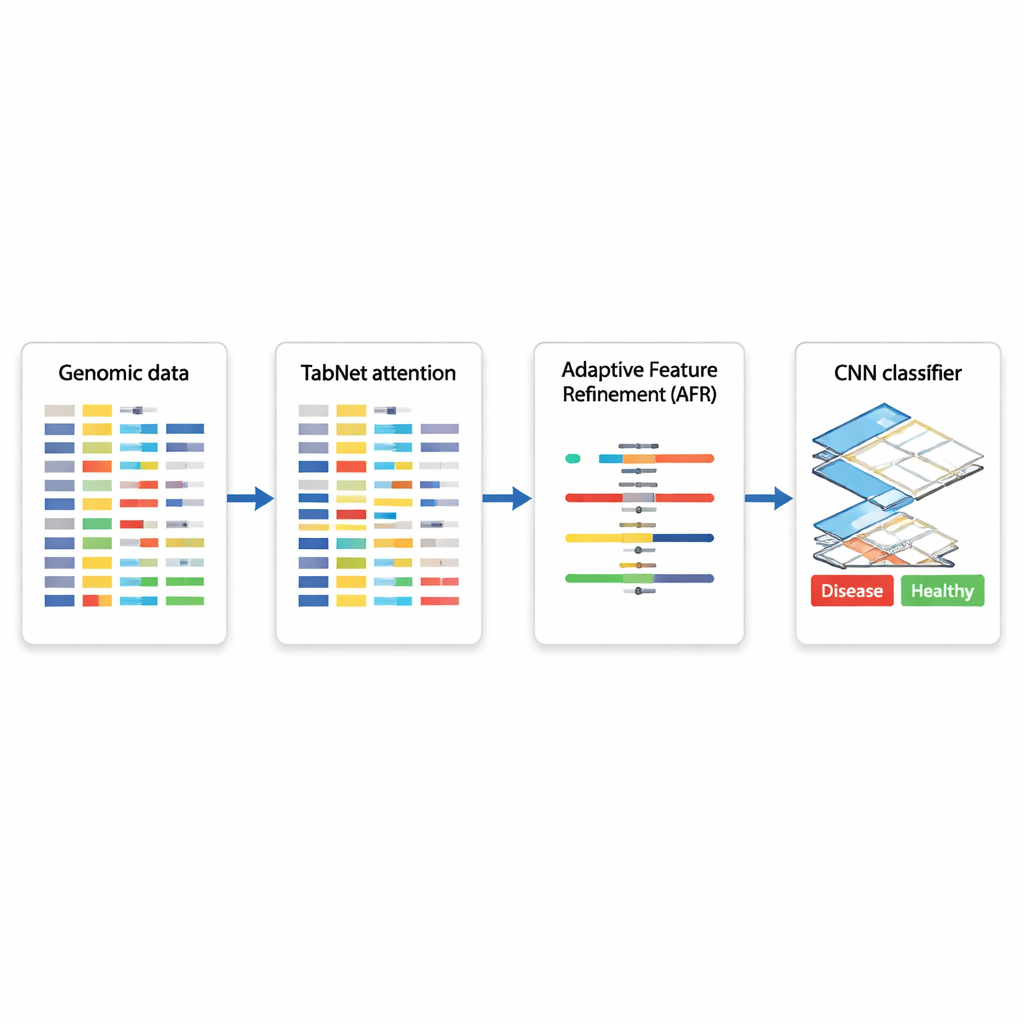
The authors propose a hybrid deep learning architecture that chains together three specialized modules. First, a component called TabNet acts like a spotlight, scanning all available genomic measurements and learning which features are most informative for a given task—for example, distinguishing cancerous from non-cancerous tissue. Instead of treating every gene equally, TabNet focuses attention on a sparse subset that appears most relevant. Next, an Adaptive Feature Refinement (AFR) layer takes these selected signals and reweights them, strengthening consistent, meaningful patterns while further dampening noise. Finally, a convolutional neural network (CNN), commonly used in image analysis, examines how the refined features interact locally, capturing subtle relationships among groups of genes that might signal a particular disease subtype or biological state.
Putting the Model to the Test
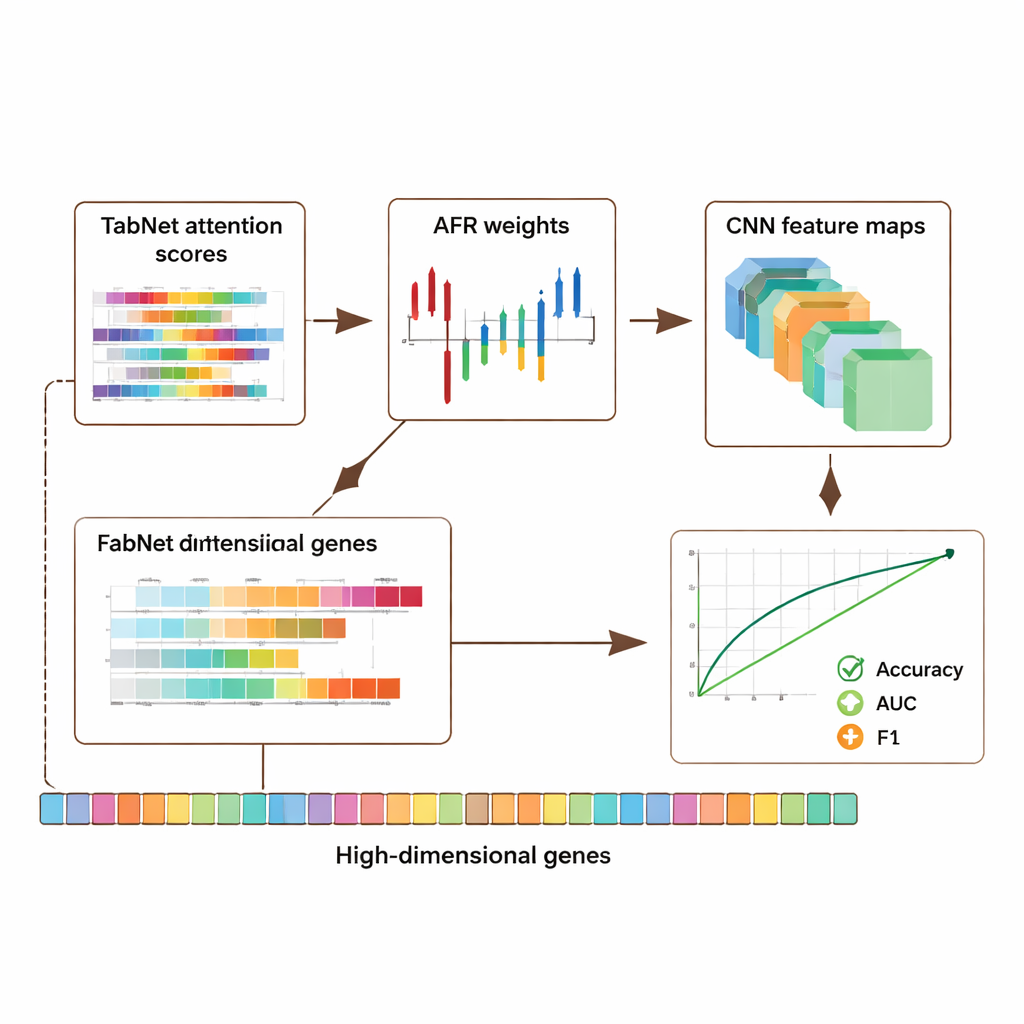
The framework was evaluated on three major public resources: a breast cancer dataset from The Cancer Genome Atlas, a single-cell melanoma dataset from the Gene Expression Omnibus, and an epigenomic dataset from the ENCODE project. Together, these collections include thousands of samples and tens of thousands of features per sample, covering gene activity and chemical marks on DNA. Across all datasets, the hybrid model outperformed several state-of-the-art approaches, improving accuracy and key measures of classification quality such as the area under the receiver operating characteristic curve (AUC) and F1-score by roughly 5–8 percentage points. Importantly, these gains did not come at the cost of transparency: the model produces attention maps from TabNet and activation maps from the CNN that highlight which genes and regions were most influential in each prediction.
Balancing Accuracy, Privacy, and Trust
Because genomic data are deeply personal, the authors also studied how to protect privacy while retaining useful signal. They introduced an adaptive privacy mechanism that adds more noise to highly sensitive features and less to others, combined with masking of selected inputs. Tests showed that even when moderate noise was introduced, the model maintained strong accuracy and discrimination, with performance degrading gracefully as protection was tightened. At the same time, the interpretable attention and activation patterns often pointed toward genes already known to play roles in cancer and immune regulation, suggesting that the system is not merely memorizing data but capturing biologically meaningful signals. An ablation study—systematically removing parts of the architecture—confirmed that each module, especially the AFR layer, made a measurable contribution to performance.
What This Means for Future Medicine
In plain terms, this work offers a smarter way to sift through enormous genomic spreadsheets to find patterns linked to disease, while also showing which entries in the spreadsheet mattered most. By combining targeted feature selection, careful refinement, and pattern recognition, the hybrid model improves prediction accuracy, remains computationally manageable, and provides visual clues that clinicians and biologists can interpret. Although more testing is needed on broader and more diverse patient groups, such frameworks could help identify new biomarkers, refine disease subtypes, and support clinical decision tools in precision medicine—bringing AI analysis of DNA one step closer to real-world use.
Citation: Swain, M.K., Kamila, N.K., Jena, L. et al. Hybrid deep learning framework for accurate classification of high dimensional genomic data. Sci Rep 16, 5919 (2026). https://doi.org/10.1038/s41598-026-36128-7
Keywords: genomic deep learning, cancer biomarker discovery, interpretable AI, precision medicine, privacy-preserving genomics