Clear Sky Science · en
ONCOPLEX: an oncology-inspired hypergraph model integrating diverse biological knowledge for cancer driver gene prediction
Why this research matters
Cancer is driven by a small number of powerful genetic changes hidden among thousands of harmless ones. Finding those truly dangerous “driver” genes is essential for better diagnosis and targeted treatments, but it is like spotting a few ringleaders in a vast, noisy crowd. This study introduces ONCOPLEX, a new artificial-intelligence framework that looks at genes not one by one, but in the context of the biological pathways they work in together, offering a sharper way to pinpoint the genes that truly fuel tumors.
Seeing cancer genes in their biological neighborhoods
Most current methods scan cancer genomes for mutations that appear unusually often or that stand out in simple gene networks. These approaches help, but biology is rarely that simple. Genes usually act in groups inside pathways that control cell growth, DNA repair, and many other processes. ONCOPLEX embraces this complexity by representing genes as dots and pathways as larger overlapping groups that can contain many genes at once. This kind of structure, known as a hypergraph, lets the model consider multi-gene relationships directly instead of breaking them into many separate pairs.
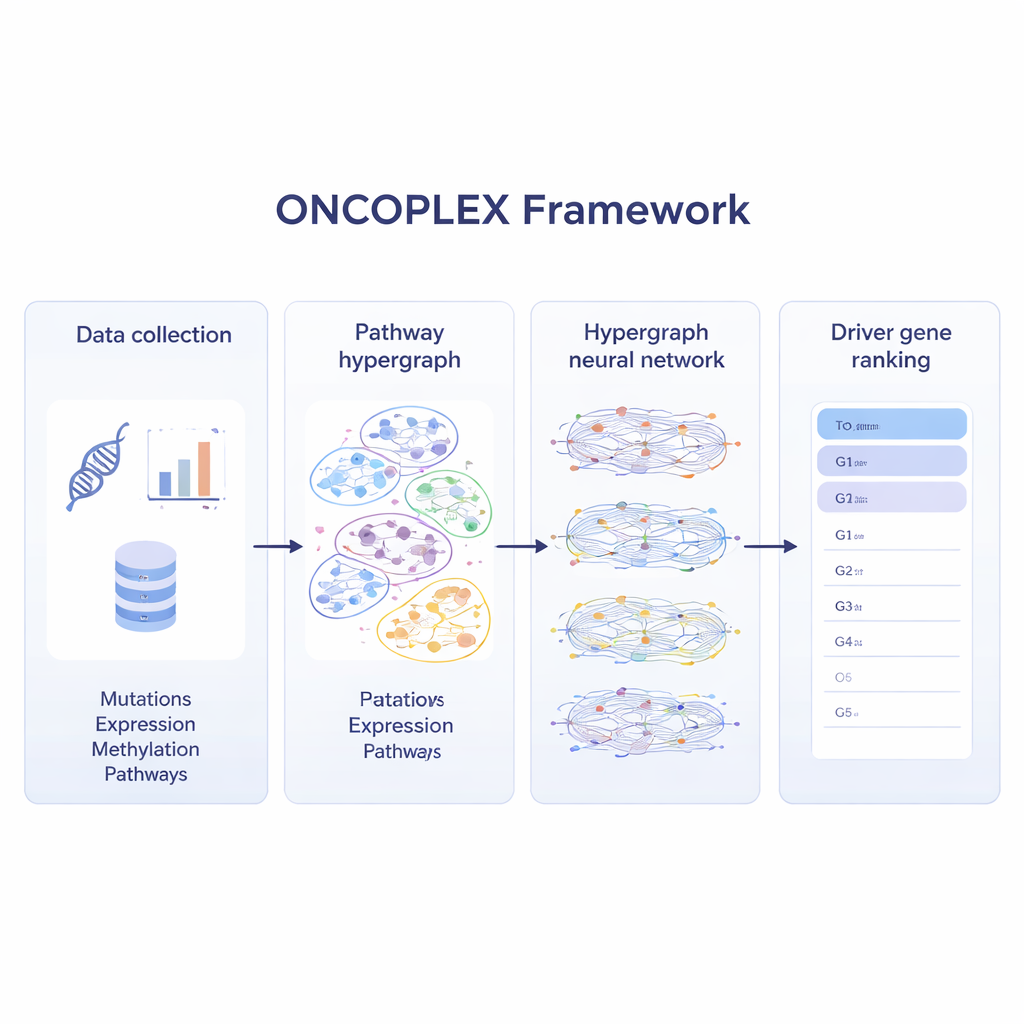
Blending many layers of cancer data
To make the most of modern cancer datasets, ONCOPLEX combines several kinds of information about each gene. It uses mutation frequencies, changes in gene activity, chemical tags on DNA (methylation), and a rich set of biological features such as evolutionary conservation and functional annotations. These features are attached to each gene in the hypergraph. A specialized neural network then passes information through the pathways, allowing each gene’s representation to be shaped both by its own data and by the behavior of the genes it works with. The model is trained using genes already known to be cancer drivers, while also learning from many unlabeled genes that might be important but are not yet recognized.
Outperforming existing tools across many cancers
The researchers tested ONCOPLEX on data from The Cancer Genome Atlas, both by pooling many tumor types together and by examining 11 individual cancers, including breast, lung, liver, bladder, and head and neck cancers. They compared it with several leading graph- and hypergraph-based methods. Across the board, ONCOPLEX was better at distinguishing known driver genes from the far more common non-drivers and at ranking likely drivers near the top of its lists. Its advantage was especially clear when looking at the highest-ranked genes, where accurate identification is most valuable for follow-up experiments and clinical translation.
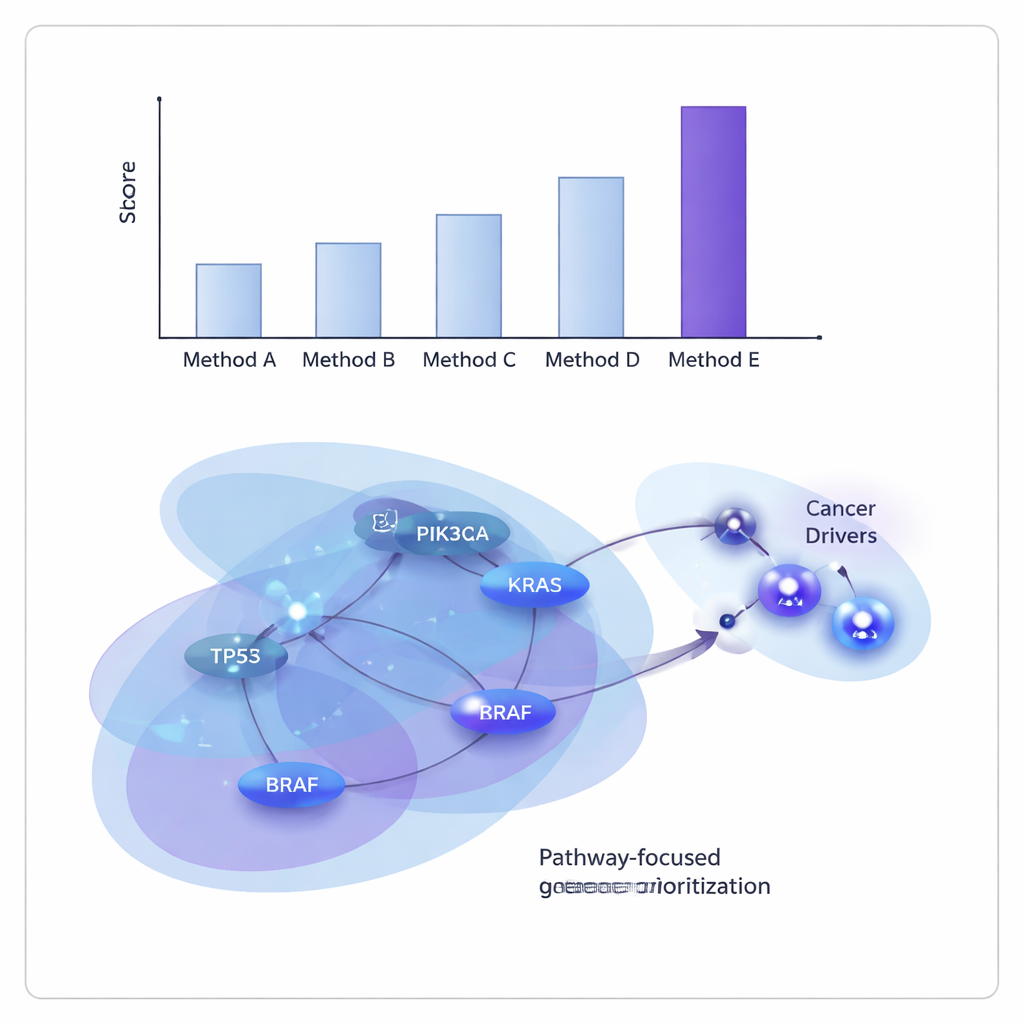
Revealing shared and cancer-specific culprits
Beyond raw performance numbers, ONCOPLEX’s ranked gene lists recovered many familiar cancer genes, such as KRAS, BRAF, and members of the PI3K–AKT signaling pathway, confirming that the model captures well-established biology. It also highlighted promising candidates that are not yet firmly recognized as drivers in certain cancer types, including genes like GRB2 and MAPK3 in breast cancer and SHC1 in stomach cancer. When the team examined the top-ranked genes using pathway enrichment analysis, they found strong signatures of well-known cancer pathways, including ErbB signaling and PI3K–AKT–mTOR, as well as immune-related pathways, suggesting that ONCOPLEX is zeroing in on networks that matter clinically.
Strengths, limits, and what comes next
By showing that richer biological features steadily improve its predictions, ONCOPLEX demonstrates the value of fusing many data sources within a pathway-centered framework. At the same time, the study uncovers a limitation: because many cancers share a large number of pathways, the model sometimes favors widely acting “pan-cancer” genes over those that are truly specific to one tumor type. The authors suggest that future work should refine how pathway information is used so that common and cancer-specific signals can be teased apart more clearly.
What this means for patients and clinicians
For non-specialists, the key takeaway is that ONCOPLEX offers a more biologically realistic way to search for the genes that drive cancer. By looking at genes in the company they keep—within pathways rather than in isolation—it improves our ability to spot both well-known and previously overlooked drivers, even in cancers where little is currently known. This kind of tool can help researchers prioritize which genes to study in the lab, guide the hunt for new drug targets, and ultimately support more precise, pathway-aware treatment strategies in oncology.
Citation: Alotaibi, E.M., Alkhnbashi, O.S. & Tran, V.D. ONCOPLEX: an oncology-inspired hypergraph model integrating diverse biological knowledge for cancer driver gene prediction. Sci Rep 16, 5164 (2026). https://doi.org/10.1038/s41598-026-36127-8
Keywords: cancer driver genes, hypergraph neural networks, multi-omics integration, pathway analysis, precision oncology