Clear Sky Science · en
Biochemical and epigenomic dissection of TFIIE function reveals gene-selective requirement in human transcription
Rethinking a “universal” gene switch
Every cell in your body relies on a finely tuned system for turning genes on and off. Textbooks often describe this system as using a fixed toolkit of “general” factors that all genes need. This study revisits that idea for one such factor, TFIIE, and finds that it is not a one‑size‑fits‑all component. Instead, TFIIE appears to be especially important for certain groups of genes, many of which help organize our DNA and maintain genome stability. Understanding when TFIIE is required, and when it can be bypassed, offers a more nuanced picture of how our cells control gene activity and respond to stress or disease.
How cells normally start reading genes
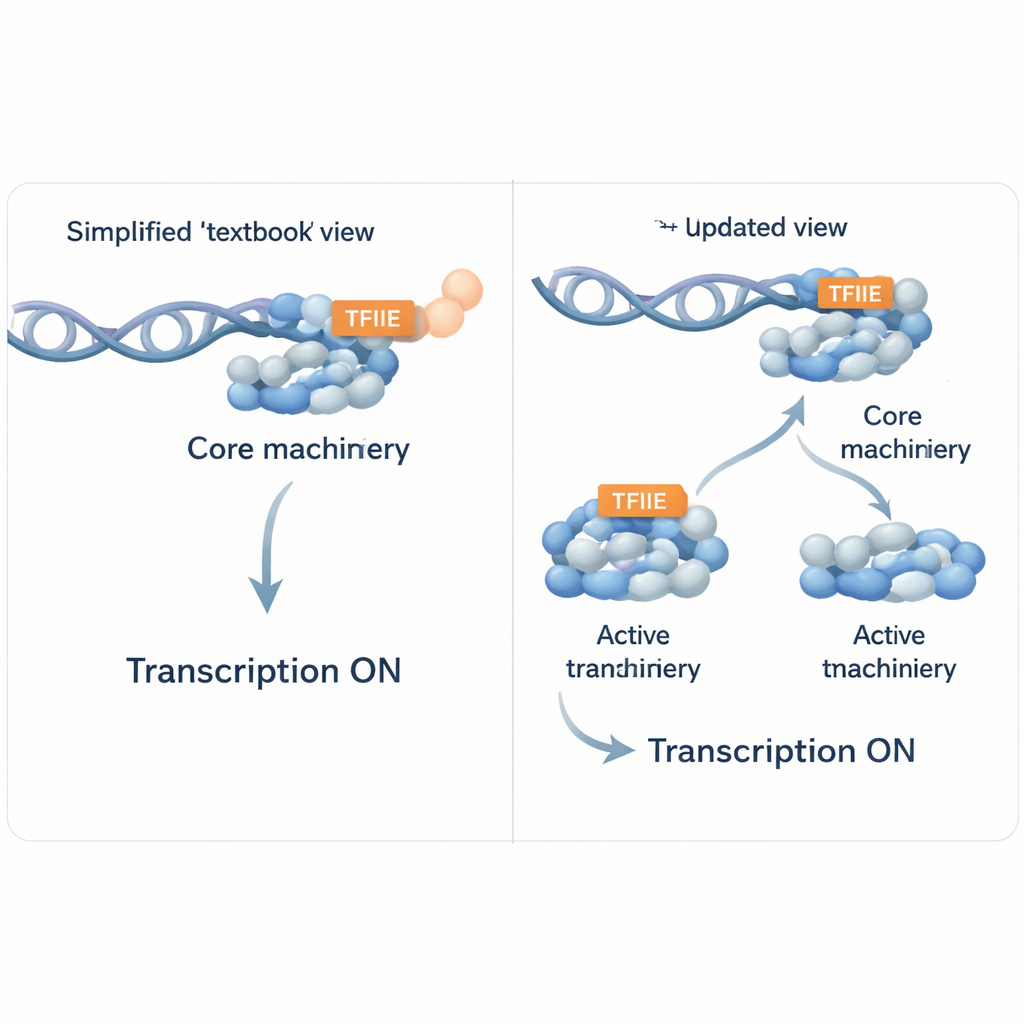
To switch on a gene, cells build a large assembly of proteins at the gene’s “start line,” called the promoter. This assembly, known as the pre‑initiation complex, includes RNA polymerase II (the enzyme that reads DNA into RNA) and several helper proteins traditionally called general transcription factors. TFIIE is one of these helpers and has long been thought to be essential at nearly every active promoter. It helps recruit another factor, TFIIH, which opens up the DNA and lets polymerase II begin copying. Under simplified laboratory conditions that only include purified components, TFIIE truly does look indispensable: when it is removed, accurate transcription barely occurs.
When the textbook rule breaks down
The authors asked what happens in a more realistic setting that still allows careful control. They used nuclear extracts from human cells—complex mixtures that contain thousands of proteins—and selectively removed a large co‑activator complex called Mediator, which normally helps bridge distant regulatory elements to promoters. In these Mediator‑depleted extracts, the team added back different purified versions of the Mediator complex and monitored which proteins bound to a model promoter and whether RNA was produced. Surprisingly, when a complete core Mediator was restored, RNA polymerase II could bind and produce transcripts even though TFIIE was barely detectable at the promoter. In other words, in a crowded nuclear environment containing many additional factors, some genes could still be transcribed without clear TFIIE recruitment, challenging the notion that TFIIE is always required.
Zooming in on specific gene neighborhoods
To see how these biochemical findings play out across the genome, the researchers examined public datasets that map where TFIIE and other key factors sit on DNA in human blood cancer cells. Using a technique called ChIP‑seq, they focused on regions around transcription start sites and asked which promoters were occupied by TFIIE together with TBP (a core DNA‑binding “landing pad”), a Mediator subunit (MED1), or a component of the ATAC co‑activator complex (ZZZ3). They found that TFIIE binds not everywhere, but to distinct subsets of promoters. Some promoters carried TFIIE along with TBP and MED1, others with TBP and ZZZ3, and a third group appeared to be enriched primarily for TFIIE itself, with little sign of canonical Mediator. Careful inspection of the raw signal suggested that “TFIIE‑only” promoters often have weak but real TBP presence, indicating a spectrum of occupancy rather than strict all‑or‑nothing categories.
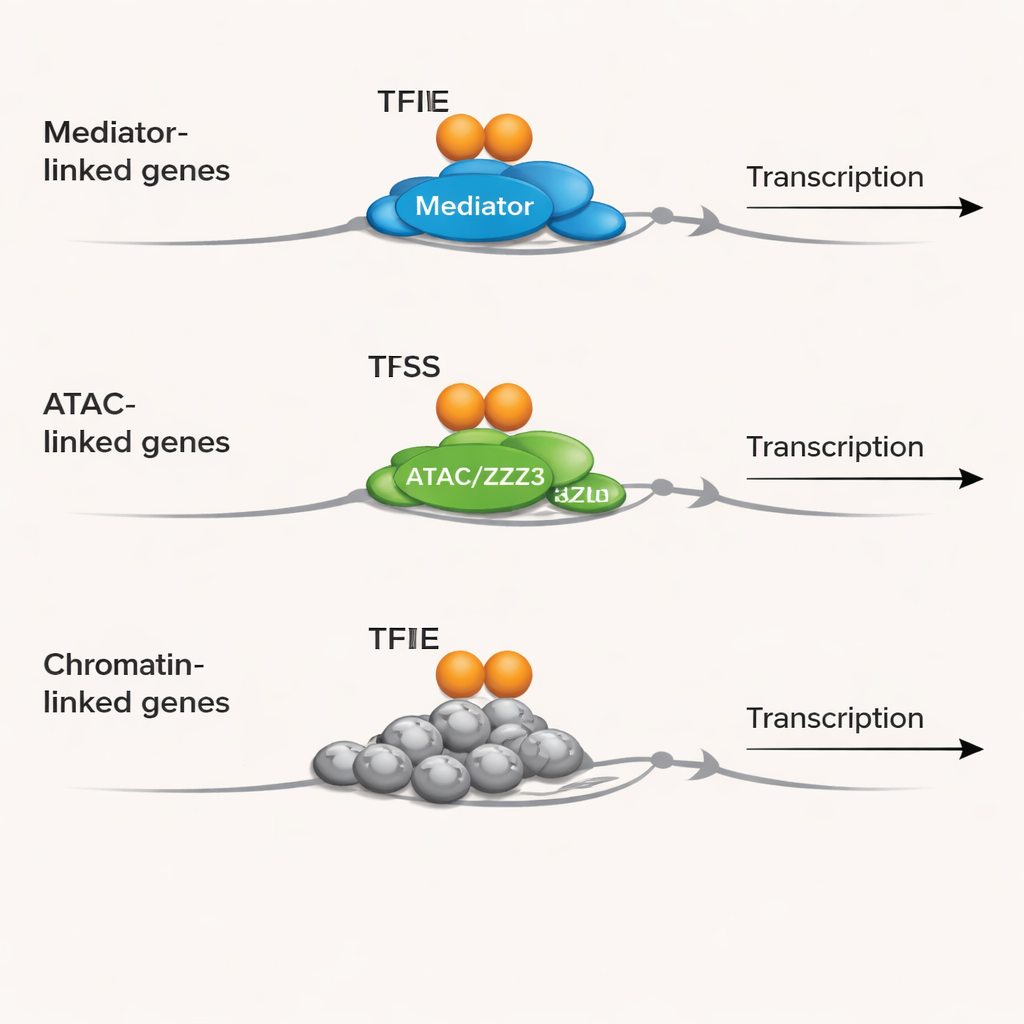
Different partner sets, different jobs
By linking these promoter groups to known gene functions, the team discovered that TFIIE seems to specialize. Promoters where TFIIE partners with TBP and MED1 are strongly associated with genes involved in RNA splicing, RNA processing, and control of protein production—steps that refine and interpret the messages made from DNA. Promoters where TFIIE works with TBP and ZZZ3 are enriched for genes that build ribosomes, assemble protein–RNA complexes, and repair DNA, tying TFIIE to basic cell growth and genome maintenance. Meanwhile, promoters marked mainly by TFIIE itself are packed with genes that shape chromatin—the way DNA is wrapped around proteins to form nucleosomes—and that influence epigenetic regulation. This pattern suggests that TFIIE helps coordinate when and how chromatin structure and RNA handling are coupled to the start of transcription.
What this means for our view of gene control
Overall, the study argues that TFIIE is not a rigid, universal requirement but a flexible player whose importance depends on the gene’s local environment and its co‑factors. In stripped‑down systems, removing TFIIE stalls transcription, but in the rich environment of the nucleus, other proteins can sometimes compensate so that certain genes still turn on. At the same time, genome‑wide maps reveal that TFIIE tends to concentrate at promoters controlling RNA processing, ribosome production, DNA repair, and chromatin organization. For a lay reader, the takeaway is that one of the supposed “basic” parts of the gene‑reading machine actually helps tune specialized programs that keep our DNA organized and our cells’ information flow in balance—insights that may eventually inform how we think about cancer, developmental disorders, and other conditions where gene regulation goes awry.
Citation: Cevher, M.A., Wijerathne, P.N., Yozgat, Y. et al. Biochemical and epigenomic dissection of TFIIE function reveals gene-selective requirement in human transcription. Sci Rep 16, 5797 (2026). https://doi.org/10.1038/s41598-026-36090-4
Keywords: gene regulation, transcription initiation, TFIIE, chromatin organization, Mediator complex