Clear Sky Science · en
Haplotype-level analysis of environmental DNA metabarcoding revealed the biogeography and phylogeography of freshwater fishes in Korean Peninsula
Reading Rivers Through Invisible Clues
Rivers and streams may look clear to the naked eye, but they are full of microscopic traces of life. This study shows how scientists can read that invisible genetic “ink” in water to discover which fish live where, how their populations are related across the Korean Peninsula, and even whether some fish have been moved by people from one region to another. The work is a glimpse of how conservationists may soon monitor biodiversity and protect rare species without casting a single net.
Fishing with DNA, Not Nets
Instead of capturing fish, the researchers collected simple one-liter bottles of water from three small streams—Seom, Gilan, and Oshipcheon—each located in a different biogeographical region of South Korea. In these samples, they searched for environmental DNA (eDNA): tiny fragments of genetic material shed by fish through skin, waste, and mucus. Using a technique called metabarcoding, they amplified and sequenced a short piece of fish DNA, then compared millions of reads to reference databases to identify which species—and even which genetic variants, or haplotypes—were present in each stream.
Who Lives Where in Korea’s Streams?
From just 54 water samples taken at nine sites, the team detected 107 distinct DNA variants representing 76 fish species—around one-third of all freshwater fish known from Korea. The Seom stream, a lower-altitude river linked to the Yellow Sea, hosted the richest fish community, with 49 species. Gilan and Oshipcheon, higher and steeper tributaries flowing toward different coasts, each held 28 species. Many fish were shared among streams, but nearly 30 species were found to be endemic to Korea’s freshwater habitats, and each stream also had its own unique set of species. Overall, the patterns matched long-standing knowledge that Korea’s freshwater life is split into three main regions by mountain ranges and geological history.
Genetic Fingerprints and Hidden Transfers
Because the scientists worked at the haplotype level—the fine-grained “flavors” of DNA within a species—they could do more than list species; they could compare populations. Several common native fish showed clear genetic differences between regions, indicating long-term separation and limited natural mixing across the peninsula. At the same time, the DNA data exposed likely cases of human-assisted movement. For species such as Pungtungia herzi, Coreoleuciscus splendidus, and Nipponocypris koreanus, the haplotype patterns in the eastern Oshipcheon stream closely matched those from western or southern rivers, pointing to past stocking or translocation. By virtually reconstructing “family trees” of these DNA variants, the team could infer probable source regions for moved fish—offering a kind of genetic forensics for river management.
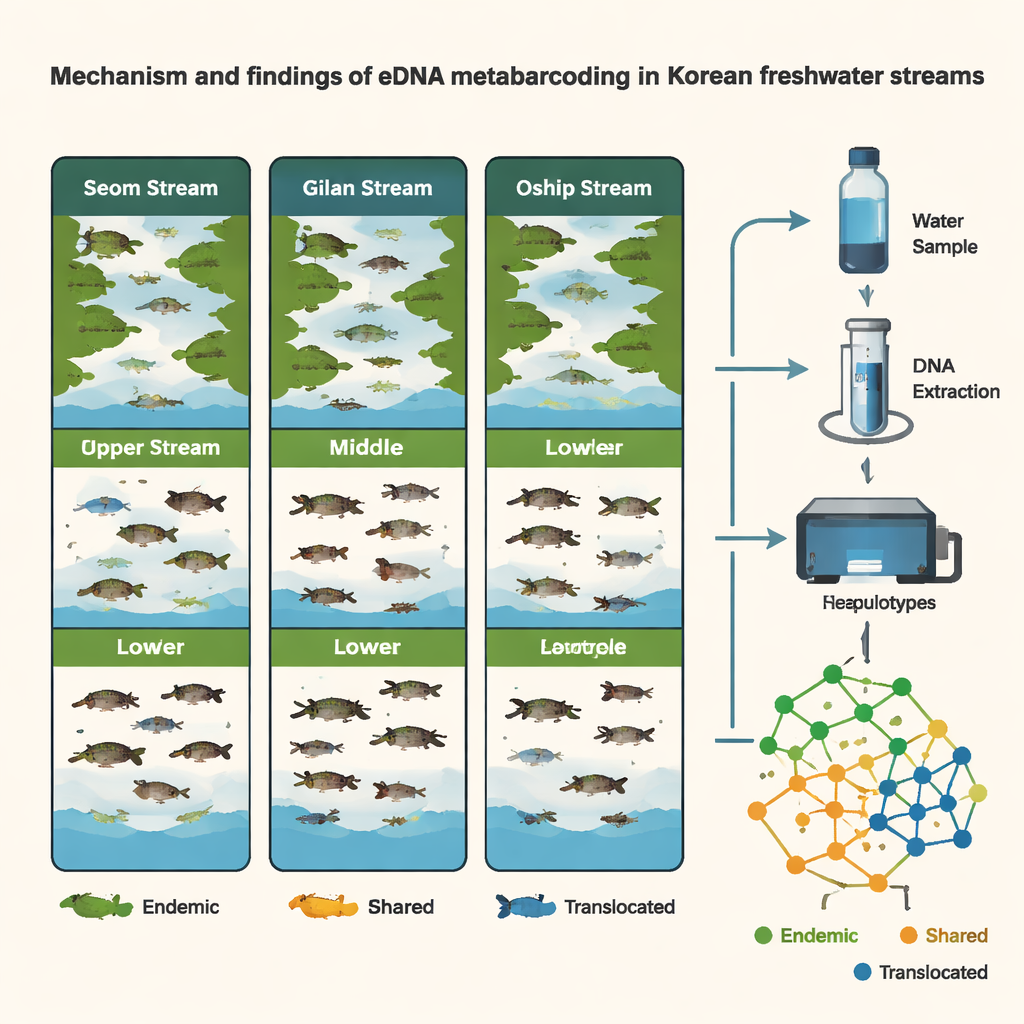
Seasons, Habitats, and Moving Targets
The study also tracked how fish communities changed over time and space. Comparing samples collected in late winter (March) and summer (August), the researchers found higher species richness in summer and clear seasonal shifts, especially in the Seom stream. In Oshipcheon, eDNA from migratory species—such as salmon and other fish that move between river and sea—appeared and disappeared in ways that matched their known life cycles. Surprisingly, within each site, riffles, runs, and pools did not show strong differences in fish DNA, likely because water flow mixes eDNA laterally across these small channels. Instead, the main contrasts came among different stream sections (upper, middle, lower) and among the three streams themselves, underscoring the importance of where along a river samples are taken.
What This Means for Protecting Freshwater Life
In plain terms, this research demonstrates that a bottle of river water can reveal a detailed snapshot of who is living in the stream, how local fish populations are related across large regions, and whether people have altered those patterns by moving fish around. For conservation, that means managers can rapidly monitor rare and endemic species, spot invasive and translocated fish, and track seasonal shifts in migratory populations without intensive fieldwork or harming animals. While eDNA cannot yet replace traditional genetic studies, it provides a powerful, low-impact tool for watching over freshwater biodiversity in a warming, heavily managed world—and for ensuring that Korea’s unique river fish continue to thrive in their native homes.
Citation: Amin, M.H.F., Kim, A.R., Jang, J.E. et al. Haplotype-level analysis of environmental DNA metabarcoding revealed the biogeography and phylogeography of freshwater fishes in Korean Peninsula. Sci Rep 16, 6955 (2026). https://doi.org/10.1038/s41598-026-36043-x
Keywords: environmental DNA, freshwater fish, biodiversity, Korean rivers, conservation genetics